- *Corresponding Author:
- M. M. Mahroop Raja
Research and Development Centre, Bharathiar University, Coimbatore-641 046
E-mail: mahroop_raja07@yahoo.co.in
| Date of Submission | 05 August 2016 |
| Date of Revision | 02 January 2017 |
| Date of Acceptance | 12 April 2017 |
| Indian J Pharm Sci 2017;79(3):369-376 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
Multidrug resistance is an emerging major clinical problem rapidly enhancing throughout the world, which require novel effective drugs to control this problem. The present study was aimed at investigation of silver nanoparticles synthesized by Micromonospora species for antibacterial activity against hospitalacquired uropathogens. Marine actinomycetes were collected from Kayalpatnam, located at Tuticorin, Tamil Nadu, India. Totally, six actinomycetes were isolated and identified based on their spore formation and biochemical studies. Six isolates were belonged to the genera of Micromonospora species. Among the six, Micromonospora species, KPMS10 showed potent antibacterial activity against Gram-negative and Gram-positive uropathogens. Silver nanoparticles synthesized by Micromonospora species KPMS10 were isolated, characterized using ultra violet-visible spectroscopy, Fourier transform infrared spectroscopy and scanning electron microscopy. The shape of nanoparticle was found to be spherical with an average size of 80 nm. The antibacterial activities of silver ion against uropathogens were found to be superior to third generation cephalosporins. These silver nanoparticles were found to be least haemolytic at 20 µg/ml. Biosynthesis of silver in combination with actinomycetes would be effective against clinical multidrug-resistant pathogens.
Keywords
Multidrug, hospital infection, silver, haemolytic
Hospital-acquired infections are emerging over the past few decades, which are the cause of high morbidity and mortality of hospitalised patients. These infections has become a global health problem as mechanism for resistance has been reported for all currently available antimicrobial agents [1]. Most hospital deaths are directly or indirectly linked to hospital-acquired infections and estimated as 80% [2]. In India, 10-30% patients are admitted in hospitals or nursing homes are associated with hospital-acquired infection. This optimisms regarding the new chemical compound was soon shattered as report on drug resistant bacteria was arisen shortly in associated with hospital infections and development of drug resistant organisms and the rate is 3.4% [3]. As new antibiotics were discovered and taken in to use, the bacteria responded by developing various resistant forms.
The use of antimicrobials has been increased and the resistant mechanism by pathogenic bacteria also increased in complexity [4]. Microbes can pose a threat in hospital environments where patients with weakened immunity are at increased risk for contracting hospitalacquired infections [5]. Arrest of the spread of drug resistant bacteria with effective treatment is yet to be established. Thus, endeavours for the discovery and application of nanotechnology are the need of the hour. Biosynthetic silver nanoparticles (AgNPs) using bacteria, fungi and plants have already been well documented [6-8]. Indeed over the past several years, AgNPs were synthesized by a potential new antibiotic developer, actinomycetes from marine sediments [9]. Actinomycetes are autochthonous free living Grampositive saprophytic marine micro flora and can be isolated from marine sediments. The obligate marine actinomycetes represent a major source for production of numerous natural biological metabolites especially antibiotics [10]. AgNPs play an important role in controlling bacterial infections [11,12]. Silver is the metal of interest as it provides the most durable antimicrobial protection against microorganism. The mechanism of silver resistance offered by bacteria using a silver binding protein is well documented [13].
In addition to nanoparticle synthesis, the broad spectrum antimicrobial activity of AgNPs was studied against clinical pathogens [14]. AgNPs have been the most promising source to kill microbes very effectively because they act both intra and extracellularly. AgNPs act as a strong bactericidal agent and depict activity against both multidrug-resistant Gram-positive and negative bacteria [15,16]. Marine actinomycetes would constitute a valuable resource for novel metabolites of pharmaceutical and medicinal interest, including antimicrobial agents [17]. The objective of this work was to synthesize, characterize and evaluate the efficacy of biosynthesized AgNPs against hospital-acquired uropathogens.
Materials and Methods
Sample collection
Systematic screening of actinomycetes from marine sediments was done by random sample collection at 200 m distance and at a depth of 5 m using core samples from Gulf of Mannar, East coastal region, Kayalpatnam, Tuticorin district, Tamilnadu, India. The central portion of the marine sediments were aseptically transferred to the sterile bottles during June 2013 and brought to the laboratory covered in an ice bag. The sediment sample was blackish brown colour and of a sandy texture.
Isolation of actinomycetes
The marine sediments were air-dried to minimize bacterial contaminants. One gram of sediment was serially diluted up to 10-6 dilution. About 1 ml of the diluted sample was permitted in to the Petri plate followed by actinomycetes isolation agar (AIA) medium supplemented with cyclohexamide 50 μg/ml and nystatin 50 μg/ml. After solidification, all the plates were incubated at 28° for 7-15 d until the colonies were developed. All the isolates were identified by slide culture method. A drop of 0.1% trypan blue stain was placed over the glass slide and then cover slip was placed over the stain gently. The slide was examined under bright field microscope and the spore was noted.
Biochemical characterization of actinomycetes
After preliminary studies, the isolates were identified by biochemical analysis i.e. indole production, methyl red and Voges-Proskauer test, citrate utilization test, catalase test, oxidase test, starch hydrolysis, casein hydrolysis and urea hydrolysis were studied.
Analyses of multiple antibiotic resistance (MAR) index
According to the guidelines of the Clinical and Laboratory Standards Institute, multidrug resistance was detected by using disc diffusion test, which was performed on Muller-Hinton agar (MHA) medium. The test bacteria were inoculated using the swab method. After 10-15 min, selected antibiotic discs were placed over the bacterial growth at equal distance. The plates were incubated at 37° for 24 h. After incubation, the zone of inhibition was considered as an indicator of drug resistance to the bacterial growth. MAR index was calculated by using following Eqn., MAR index = number of antibiotics to which the isolate was resistant/ total number of antibiotics tested.
Antibacterial activity of secondary metabolites
Antimicrobial activity of Micromonospora species KPMS10 culture filtrate was analysed by agar well diffusion method. The 24 h nutrient broth culture of multidrug-resistant pathogens such as Escherichia coli, Klebsiella pneumoniae, Enterococcus sp., Staphylococcus aureus, Pseudomonas aeruginosa, Proteus mirablis and P. vulgaris were isolated from hospital-acquired infection and bioassayed against standard antibiotics. About 100 μl of culture filtrate was used as a test sample against clinical isolates. Inoculated plates were incubated at 37° for 24 h. After incubation, all plates were examined for the presence of zone of inhibition around the wells.
Biosynthesis and characterization of AgNPs
The isolated Micromonospora species KPMS10 strain was inoculated in to International Streptomyces Project medium IV (ISP IV) and incubated at 28° for 7-15 d. The culture broth medium was centrifuged at 5000 rpm for 10-20 min to remove cell debris. About 10 ml of cultured filtrate was permitted in to 100 ml of 1 mM AgNO3 solution and incubated at room temperature in a shaker under dark condition. The bioreduction of pure AgNO3 solution was monitored at regular intervals using a UV/Vis spectrophotometer between 190-1100 nm. Synthesized AgNPs were collected by subsequent high speed centrifugation at 15 000 rpm and the functional group was detected by Fourier transform infrared spectroscopy (FTIR) between 4000- 400 cm-1. Morphology and size of AgNPs was studied under scanning electron microscope (SEM).
Antibacterial activity of AgNPs
Antimicrobial activity of the AgNPs was evaluated using agar well diffusion method. Multidrugresistant pathogens such as E. coli, K. pneumoniae, Enterococcus sp., S. aureus, P. aeruginosa, P. mirablis and P. vulgaris were isolated from hospitalacquired infection and inoculated into the nutrient broth and incubated at 37° for 24 h. After incubation, test pathogens were inoculated on MHA using sterilized cotton swabs. In each of these plates, wells were cut using a sterile gel borer and 50 μl of AgNPs were placed against clinical isolates. Inoculated plates were incubated at 37° for 24 h. After incubation, all plates were examined for the presence of zone of inhibition around the wells.
Haemolysis assay of AgNPs
The cytotoxic effect of AgNPs was studied using the haemolytic test [18]. A pellet of RBC was collected and washed three times with phosphate-buffered saline (PBS) at 10 000 rpm for 3 min. The obtained RBC was diluted with PBS at 1:10 ratio. About 400 μl of RBC suspensions were treated with 100 μl of nanoparticles in Tyrode at 1, 5, 10, 25 and 50 μg/ml and Triton X-100 as positive control. The suspension was incubated at room temperature by shaking the plate for 24 h. The percent haemolysis was calculated by following Eqn., H (%) = (sample OD–Tyrode OD)/(1% Triton X-100 OD–Tyrode OD)100.
Phylogenetic analysis
The 16S rRNA sequences of Micromonospora species KPMS10 was blasted and deposited in the Genbank of National Centre for Biotechnology Information (NCBI).
Results and Discussion
The present investigation involved the synthesis of AgNPs by the marine actionomycets. Based on cultural characterization, the actinomycetes used were identified as Micromonospora species. All isolated Micromonospora strains were Gram-positive but differed morphologically by producing different mycelia and spores. Micromonospora species showed monospore on aerial mycelium. The colors of the substrate mycelium were yellowish white to dark ash after sporulation. The morphological characteristics of these isolates were consistent with their classification in the genus Micromonospora [19]. The Micromonospora species KPMS10 was strictly found to be positive in indole, Voges-Proskauer, catalase, oxidase, urease and casein hydrolysis. Identification of actinomycetes genera and species based on the morphology and biochemical properties indicated potential strain of marine actinomycetes [20].
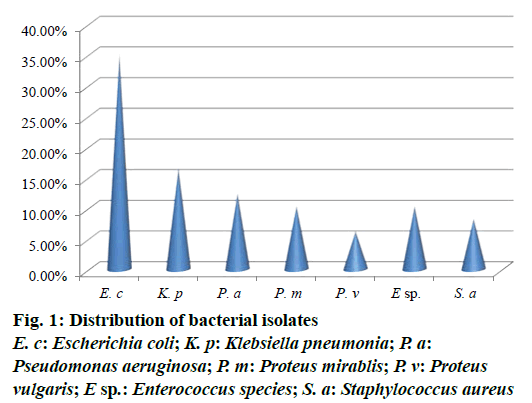
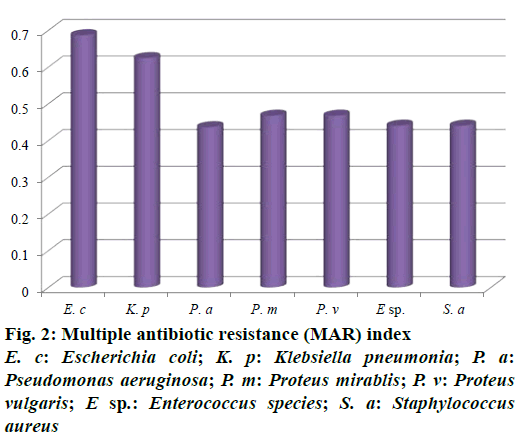
Out of 73 hospital isolates, 48 showed Gram-positive types of infection and 25 found to be Gram-negative. Totally 48 isolates were recorded and identified as E. coli, K. pneumoniae, P. aeruginosa, P. mirablis, P. vulgaris, Enterococcus species and S. aureus. The highest number of patients with hospital infections was of the age range of 40-50 y followed by the age range 50-60 y. The frequency of pathogens in our study was E. coli (35.41%) and K. pneumonia (16.66%), P. aeruginosa (12.5%), P. mirablis (10.41%), P. vulgaris (6.25%), Enterococcus species (10.41%) and S. aureus (8.33%) as shown in Figure 1. The most common hospital-acquired infection was caused by E. coli, the predominant bacteria found in hospitalized patients [21]. All tested pathogens were analysed by MAR index using standard antibiotics. The antimicrobial sensitivity of isolated clinical pathogens revealed that 65.6% of isolates were found to be MDR. Among the isolates tested, E. coli was found in 0.687, K. pneumoniae as 0.625 and lowest MAR index found in Gram-positive isolates Enterococcus species and S. aureus as 0.441, respectively (Figure 2). The highest multiple antibiotic resistant indices (MARI) for E. coli were 0.687. The overall rate of resistance against E. coli was worldwide reported, which was similar with this study [22]. MAR index is a tool that reveals the spread of bacteria resistance in a given population [23]. MAR index greater than 0.20 implies that the strains of such bacteria originate from an environment where several antibiotics are used. The MAR indices obtained in this study is a possible indication that a very large proportion of the bacterial isolates have been exposed to several antibiotics. These bacteria are common environmental organisms, which act as opportunistic pathogen in clinical cases where the defence system of the patient was compromised to broad spectrum antibiotics particularly to penicillin and cephalosporins [24].
About 32 broad spectrum antibiotics were tested against hospital pathogens. Antibiotic sensitivity revealed that the high degree of resistance (50-100%) was reported against cephalosporin, monobactam and ampicillin. Ampicillin resistance of 100% was significantly reduced to 28% by clavalunate. Significant resistant (≤50%) was observed against fluoroquinolones derivatives. Among the tested antibiotics, 7 antibiotics belonged to amikacin, carbapenems, tazobactam and cepahatoxime clavulnate showed potent antibacterial activity against tested pathogens. No resistance was observed against these antibiotics. It has been reported that meropenem is the most effective antibiotic against Gram-negative and Gram-positive uropathogens in the rate of 93-100% [25].
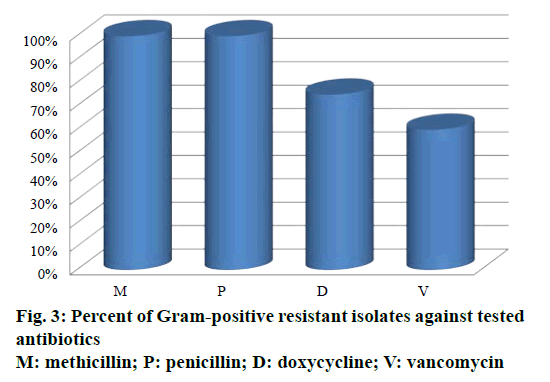
Out of 34 antibiotics tested against Gram-positive S. aureus and Enterococcus species, 100% of resistance was developed against penicillin and methicillin (Figure 3). S. aureus showed 75% resistance against doxycycline and no resistance against vancomycin. Enterococcus species showed 60% resistance against vancomycin and no resistance against doxycycline. As described previously, methicillin resistance was associated with resistant to other antibiotics [26]. In the present study, the fact that high prevalence of MRSA infections have showed sensitivity to vancomycin encourages the usage of vancomycin rather than doxycycline [27].
The antimicrobial activity of Micromonospora species KPMS10 was given in Table 1. Potent antibacterial effect was recorded as 18 mm against K. pneumoniae and E. coli, moderately active against P. aeruginosa (15 mm), P. vulgaris (16 mm), Enterococcus species (16 mm) S. aureus (14 mm) but least activity against P. mirablis (13 mm). It was found that meropenem was more effective than actinomycetes. Micromonospora species are the common inhabitants that have proven to be a constant source of novel bioactive compounds with structural and functional diversity giving rise to antibacterial, antifungal, antiviral, anticancer activity [28]. Marine actinomycetes has the unique distinction that various secondary metabolites can be produced by a single strain [29].
| Strain code | E. coli | K. pneumoniae | P. vulgaris | P. aeruginosa | P. mirablis | S. aureus | Enterococcussp. |
|---|---|---|---|---|---|---|---|
| KPMS10 | 18±0.04 | 18±0.036 | 15±0.02 | 16±0.04 | 13±0.4 | 14±0.04 | 16±0.02 |
| Meropenem | 18±0.02 | 18±0.02 | 18±0.1 | 18±0.04 | 20±0.1 | 20±0.2 | 20±0.04 |
Zone of inhibition mm in diameter, mean±SD, n=3. Among the six tested strains, KPMS10 showed potent antibacterial activity
Table 1: Antibacterial activity of marine actinomycetes against multidrug resistant pathogens
In this study, AgNPs were biosynthesized with the help of culture filtrate of Micromonospora species KPMS10. It showed pale yellow colour before the addition of silver ions, which changed to brownish colour. No colour change was observed without the addition of silver nitrate in culture supernatant. Surprisingly, the intensity of brown colour increased periodically and was maintained throughout the experiments. The appearance of a pale yellow colour to brownish colour in the solution containing biomass became an indicator of the formation of AgNPs [30].
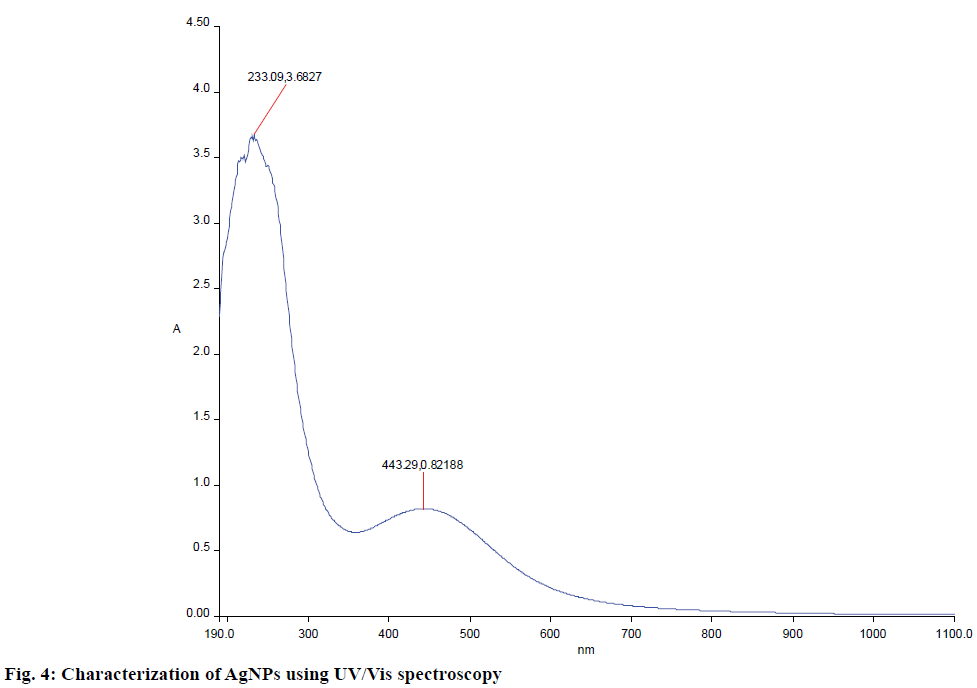
Biosynthesized AgNPs were further confirmed using UV/Vis spectrophotometer. A strong and broad plasmon absorption peak was observed at 450 nm in UV/Vis surface (Figure 4). This might occur due to the reduction of metal ions by secondary metabolites present in the cell. The absorption peaks indicated the presence of biosynthesized AgNPs. Homogenous AgNPs have been known to produce a surface plasmon resonance band at the range of 420-450 nm [31]. An absorption peak at 450 nm by S. glaucus was previously reported [32].
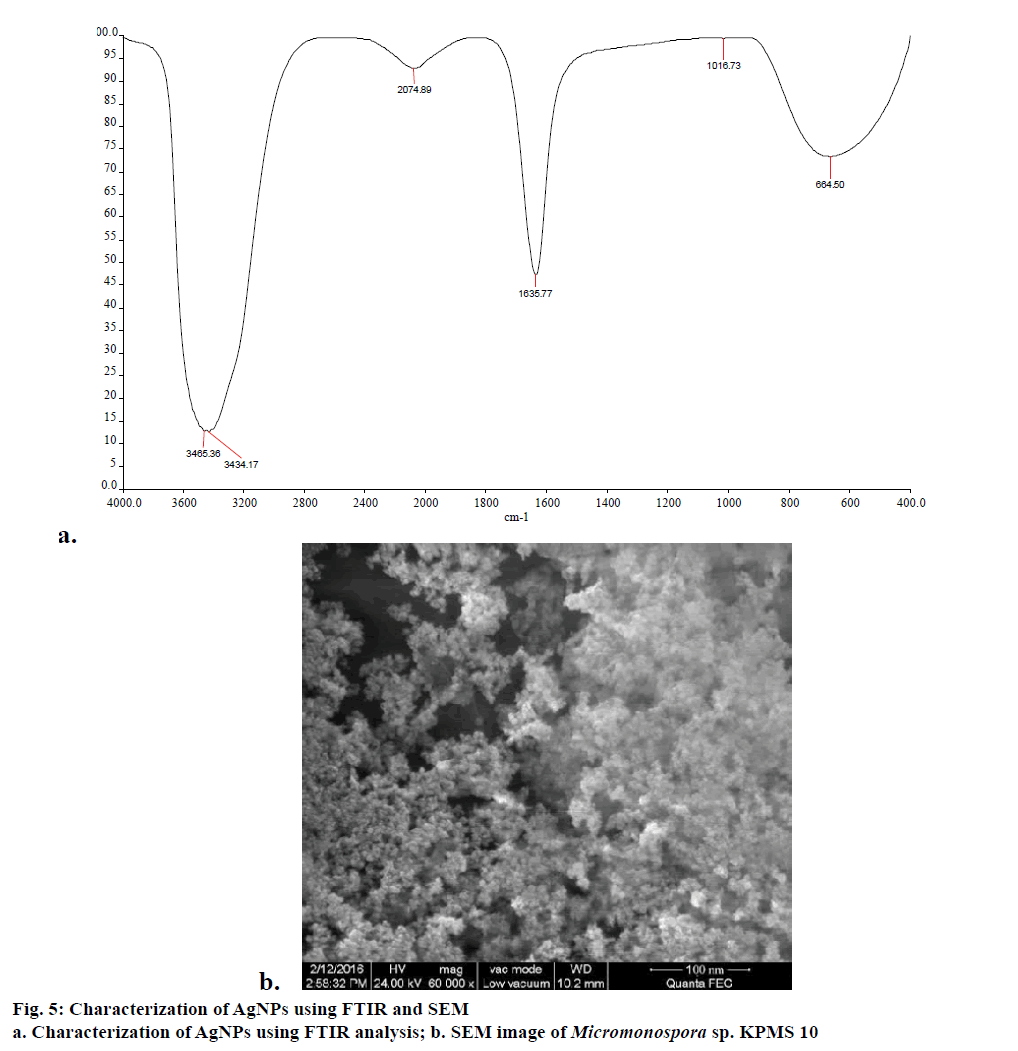
FTIR measurements were carried out to identify possible interaction between silver and protein molecules, which could be responsible for synthesis, stabilization of well dispersed AgNPs in the reaction mixture [33]. FTIR revealed a spectrum at 3465.36 and 3434.17 cm-1, which is assigned to the hydrogen bonded OH stretch, peak as 2074.39 cm-1 corresponding to carboxylic acid group (Figure 5a). Similarly, peak 1635.77 is assigned to the primary or secondary amines (N-H). The FTIR studies confirmed the fact that COOH derivative have strong ability to bind silver and to stabilize the synthesis of nanoparticles [34].
Smaller size nanoparticles effectively penetrated to cell due to its larger surface availability for interaction and interfere with metabolism of cell. In this study, SEM analysis shows that individual silver particles as well as a number of aggregates. The SEM analysis reveals that size of nanoparticles was found to be 80 nm (Figure 5b). The biomolecules present in the surface of nanoparticles leads to agglomeration structure. Similar results were observed by Bacillus licheniformis mediated AgNPs [35].
The present study showed antibacterial activity of silver ions against all tested pathogens. The maximum zone of inhibition of 24 mm was found with E. coli and K. pneumoniae with 75% of relative zone of inhibition (RIZD). About 22 mm zone of inhibition was observed against P. aeruginosa and Enterococcus species with 55 and 66% RIZD (Table 2). These results suggested that the AgNPs produced were more potent than meropenem. Antimicrobial activity was reported to be due to the penetration of AgNPs into the drug resistant bacteria resulting in disruption of cell membrane and release of cell contents [36]. Antimicrobial property of AgNPS depended on the size of nanoprticles [37]. It was found that smaller AgNPs synthesized by microorganisms had a greater antibacterial activity.
| Components | E. coli | K. pneumoniae | P. vulgaris | P. aeruginosa | P. mirablis | S. aureus | Enterococcussp. |
|---|---|---|---|---|---|---|---|
| AgNPs B | 24 | 24 | 20 | 22 | 20 | 18 | 22 |
| Cephotaxime | 16 | 16 | 16 | 18 | 16 | 16 | 18 |
| Negative control | 12 | 12 | 12 | 12 | 12 | 12 | 10 |
| RIZD (%) | 75 | 75 | 44 | 55 | 44 | 37 | 66 |
Zone of inhibition mm in diameter
Table 2: Antimicrobial activity of biosynthesized agnps against multidrug resitant pathogens
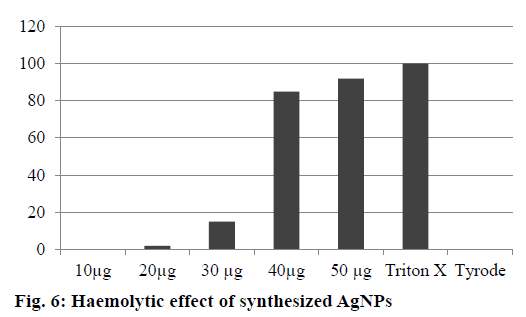
The AgNPs showed no haemolytic activity at 10 and 20 μg/ml, slight haemolysis at 30 μg and significant haemolytic activity at 50 μg/ml after 12 h incubation. Ninety percent haemolysis was observed at 50 μg/ml after 12 h incubation. The lower haemolytic activity observed at 25 μg could lead to adverse effects. Based on the results, the safe range of AgNPs was 10 to 20 μg/ml. Negative control (Tyrode) and positive control (Triton X-100 1%) induced 0 and 100% haemolysis, respectively (Figure 6). The determination of haemolysis is based on haemoglobin absorbance at 550 nm, with subtraction of the interference of AgNPs. Various studies have reported haemolytic activity of AgNPs, and our results were in agreement with those reported in previous studies about the prohaemolytic properties of AgNPs [38].
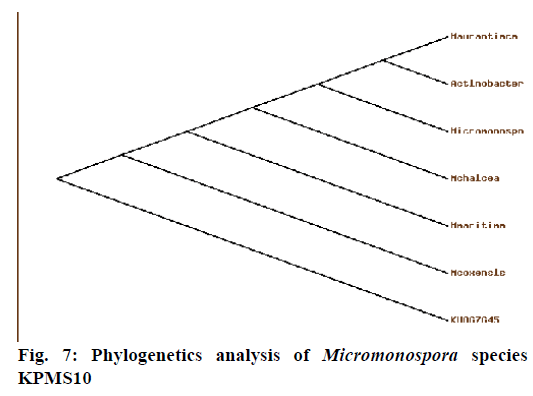
Phylogenetic analysis (Figure 7) of 16S rRNA sequence analysis showed that the Micromonospora species KPMS10 was closely related to Micromonospora coxensis (98%) and the Genbank accession number was KU867645. Currently, taxon actinomycetes are accommodates spore forming Gram-positive bacteria that from extensive branching substrates on aerial mycelia. Based on these studies, the actinomycetes strains were recorded [39].
In conclusion, application of isolated marine Micromonospora species in nanotechnology could meet the challenges in pharmaceutical industry for developing new antibacterial compounds to treat existing infectious diseases. Marine derivative could provide a promising source of structurally diverse secondary metabolites from actinomycetes with potential activity against multidrug-resistant pathogens. Further studies are needed to identify the active lead molecule.
Acknowledgements
Authors thank the Department of Microbiology, Jamal Mohamed College (Autonomous), Tiruchirappalli, for supporting this work.
Conflicts of interest
There are no conflicts of interest.
Financial support and sponsorship
Nil.
References
- Harbottle H, Thakur S, Zhao S, White DG. Genetics of antimicrobial resistance. Anim Biotechnol 2006;17:111-24.
- Hughes AJ, Ariffin N, Huat TL, Abdul Molok H, Hashim S, Sarijo J. Prevalence of nosocomial infection and antibiotic use at a university medical center in Malaysia. Infect Control Hosp Epidemiol 2005;26:100-4.
- Khoiee EM, Vaziri MH, Alizadegan S, Motevallian SA, Razzaghikashani OM, Goushegir SA. Developing the moral distress scale in the population of Iranian nurses. Iran J Psychiatry 2008;3:55-8.
- Tenover FC. Mechanisms of antimicrobial resistance in bacteria. Am J Infect Control 2006;34:S3-10.
- Hidron AI, Edwards JR, Patel J, Horan TC, Sievert DM, Pollock DA. NHSN annual update: antimicrobial-resistant pathogens associated with healthcare-associated infections: annual summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2006-2007. Infect Control Hosp Epidemiol 2008;29:996-1011.
- He W, Wu L, Gao Q, Du Y, Wang Y. Identification of AHBA biosynthetic genes related to geldanamycin biosynthesis in S. hygroscopicus 17997. Curr Microbial 2006 ;52:197-203.
- McLeod MC, McHenry RS, Beckman EJ, Roberts CB. Synthesis and stabilization of silver metallic nanoparticles and premetallic intermediates in perfluoro polyether/CO2 reverse micelle systems. J Phys Chem B 2003;107:2693-700.
- Vedpriya A. Living Systems: eco-friendly nanofactories. Dig J Nanomater Biostruct 2010;5:9-21.
- Ahmad A, Senapati S, Khan MI, Kumar R, Ramani R, Srinivas V. Intracellular synthesis of gold nanoparticles by a novel alkalotolerant actinomycetes Rhodococcusspecies. Nanotechnology 2003;14:824.
- Bull AT, Ward AC, Goodfellow M. Search and discovery strategies for biotechnology: the paradigm shift. Microbiol Mol Biol Rev 2000;64:573-606.
- Ip M, Lui SL, Poon VK, Lung I, Burd A. Antimicrobial activities of silver dressings: an in vitro comparison. J Med Microbiol 2006;55:59-63.
- Singh S, Kumar P, Gopalan N, Shrivastava B, Kuhad RC, Chaudhary HS. Isolation and partial characterization of actinomycetes with antimicrobial activity against multidrug resistant bacteria. Asian Pac J Trop Biomed 2012;2:S1147-50.
- Silver S. Bacterial silver resistance: molecular biology and uses and misuses of silver compounds. FEMS Microbiol Rev 2003;27:341-53.
- Duran N, De Souza GIH, Alves OL, Esposito E, Marcato PD. Antibacterial activity of silver nanoparticles synthesized by Fusarium oxysporum strain. J Nanotechnol 2003;122-8.
- Zhang L, Jiang Y, Ding Y, Povey M, York D. Investigation into the antimicrobial behaviour of suspension of ZnO nanoparticles (ZnO nano fluids). J Nanopart Res 2007; 9:479-89.
- Roe D, Karandikar B, Bonn-Savage N, Gibbins B, Roullet JB. Antimicrobial surface functionalization of plastic catheters by silver nanoparticles. J Antimicrob Chemother 2008;61:869-76.
- Mitsuiki S, Sakai M, Moriyama Y, Goto M, Furukawa K. Purification and some properties of keratinolytic enzyme from an alkaliphilic Nocardiopsis sp. TOA-1. Biosci Biotechnol Biochem 2002;66:164-7.
- Krajewski S, Prucek R, Panacek A, Avci-Adali M, Nolte A, Straub A. Hemocompatibility evaluation of different silver nanoparticle concentrations employing a modified Chandler-loop in vitro assay on human blood. Acta Biomater 2013;9:7460-8.
- Kawamoto I. Genus Micromonospora. In: Williams ST, Sharpe ME, Holt JG, editors. Bergey’s Manual of Systematic Bacteriology. Baltimore: The Williams and Wilkins Co.; 1989. p. 2442-50.
- Laidi RF, Kansoh AL, Elshafei AM, Cheikh B. Taxonomy, Identification and biological activities of novel isolate of Streptomyces tendae. Arab J Biotech 2006;9:427-36.
- Manikandan S, Ganesapandian S, Manoj Singh, Kumaraguru AK. Antimicrobial susceptibility pattern of urinary tract infection causing human pathogenic bacteria. Asian J Med Sci 2011;3:56-60.
- Mandal P, Kapil A, Goswami K, Das B, Dwivedi SN. Uropathogenic Escherichia coli causing urinary tract infections. Indian J Med Res 2001;114:207-11.
- Krumperman PH. Multiple antibiotics resistance indexing of E. coli to identify high-risk sources of fecal contamination of foods. Appl Environ Microbiol 1983;46:165-70.
- Obritsch MD, Fish DN, Maclaren R, Jung R. National surveillance of antimicrobial resistance in Pseudomonas aeroginosa isolates obtained from intensive care unit patients from 1993-2002. Antimicrob Agents Chemother 2004;48:4606-10.
- Eta S, Kawahara M. Antimicrobial susceptibilities of organisms isolated from patients with urinary tract infections in 2002. Jpn J Antibiot 2006;59:21-8.
- David MZ, Daum RS. Community-associated Methicillin-Resistant Staphylococcus aureus: epidemiology and clinical consequences of an emerging epidemic. Clin Microbiol Rev 2010;23:616-87.
- Appelbaum PC. Reduced glycopeptide susceptibility in Methicillin-Resistant Staphylococcus aureus (MRSA). Int J Antimicrob Agents 2007;30:398-408.
- Koehn FE, Carter GT. The evolving role of natural products in drug discovery. Nat Rev Drug Discov 2005;4:206-20.
- Schiewe HJ, Zeeck A. Cineromycin, Y-butyrolactones and ansamycins by analysis of the secondary metabolite pattern created by a single strain of Streptomyces. J Antibiot (Tokyo) 1999;52:635-42.
- Sastry M, Ahmad A, Khan IM, Kumar R. Biosynthesis of metal nanoparticles using fungi and Actinomycetes. Curr Sci 2003;85:162-70.
- Durán N, Marcarto PD, De Souza GIH, Alves OL, Esposito E. Antibacterial effect of silver nanoparticles produced by fungal process on textile fabrics and their effluent treatment. J Biomed Nanotechnol 2007;3:203-8.
- Tsibakhashvili NY, Kirkesali EI, Pataraya DT, Gurielidze MA, Kalabegishvili TL, Gvarjaladze DN. Microbial synthesis of silver nanoparticles by Streptomyces glaucus and Spirulina platensis. Adv Sci Lett 2011;4:1-10.
- Jain N, Bhargava A, Majumdar S, Tarafdar JC, Panwar J. Extracellular biosynthesis and characterization of silver nanoparticle using Aspergillus flavus NJP08: a mechanism perspective. Nanoscale 2011;3:635-41.
- Zayed MF, Eisa WH, Shabaka AA. Malva parviflora extract assisted green synthesis of silver nanoparticle. Spectrochim Acta A Mol Biomol Spectrosc 2012;98:423-8.
- Kalimuthu K, Suresh Babu R, Venkataraman D, Bilal M. Biosynthesis of silver nanocrystals by Bacillus licheniformis. Colloids Surf B Biointerfaces 2008;65:150-3.
- Panacek A, Kvítek L, Prucek R, Kolar M, Vecerova R, Pizúrova N, et al. Silver colloid nanoparticles: synthesis, characterization and their antibacterial activity. J Phys Chem B 2006;110:16248-53.
- Richard JW, Spencer BA, McCoy LF, Carina E, Washington J, Edgar P, et al. Acticoat versus Silverlon: the truth. J Burns Surg Wound Care 2012;1:11-20.
- Choi J, Reipa V, Hitchins VM, Goering PL, Malinauskas RA. Physicochemical characterization and in vitro haemolysis evaluation of silver nano particles. Toxicol Sci 2011;123:133-43.
- Waksman SA. The Actinomycetes: classification, identification and description of genera and species. Baltimore: The Williams and Wilkins Co.; 1961.