- Corresponding Author:
- S. Saxena
Department of Biotechnology, Thapar University, Patiala-147 004, India
E-mail: sanjaibiotech@yahoo.com
| Date of Submission | 01 January 2015 |
| Date of Revision | 11 October 2015 |
| Date of Acceptance | 29 November 2015 |
| Indian J Pharm Sci 2015;77(6):758-763 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms.
Abstract
Fem proteins are the essential structural proteins of various gram-positive bacteria. These are of three different types namely FemX (FmhB), FemA and FemB. Only two Fem protein crystallographic structures are available till date, one for FemA in Staphylococcus aureus and another for FemX in Weissella viridescensis. In this study, computational methods are used to evaluate interaction of FemA protein with catechin and epicatechin analogues. The interaction of FemA protein with catechin and epicatechin analogues are confirmed by binding energy and scores given by Autodock Vina and UCSF Dock docking softwares, which is followed by Lipinski filters and toxicity studies using online Lipinski server of SCFBIO and OSIRIS. Catechin gallate has been found as the best ligand for FemA protein in all aspects and it has outperformed all catechin and epicatechin isomers.
Keywords
Fem, virtual screening, gram-positive, bacteria
Methicillin resistance in Staphylococcus aureus (MRSA) is a major cause of various nosocomial infections in humans, which are responsible for morbidity and morality. Target site of ß-lactam antibiotics predominantly is PBP-2 protein. Production of additional low-affinity PBP-2a or PBP-2’ protein by mecA chromosomal structural gene is responsible for the resistance of S. aureus. Fem genes are the other additional chromosomal genes, which are responsible for methicillin resistance [1]. Fem genes belong to FemXAB family and produces three proteins namely, FemX (fmhB), FemA and FemB. Fem proteins are responsible for sequential building of interpeptide bonds in cell wall of Staphylococcus aureus. FemX (fmhB) forms first glycine-lysine bond [2,3], followed by next two bonds by FemA and last two by FemB protein [4]. This penta-glycine chain is essential for integrity of cell wall of S. aureus. Inhibition of either of these proteins can acts as a target for new drug development against gram-positive bacteria [5].
In the present work, FemA has been considered as a potential target to identify inhibitors for overcoming resistance to ß-lactam antibiotics in S. aureus. FemA is the only fem protein of S. aureus for which x-ray crystallography structure is available (1LRZA) [5]. It is composed of 426 amino acid residues and has a molecular weight of 50.27kDa.
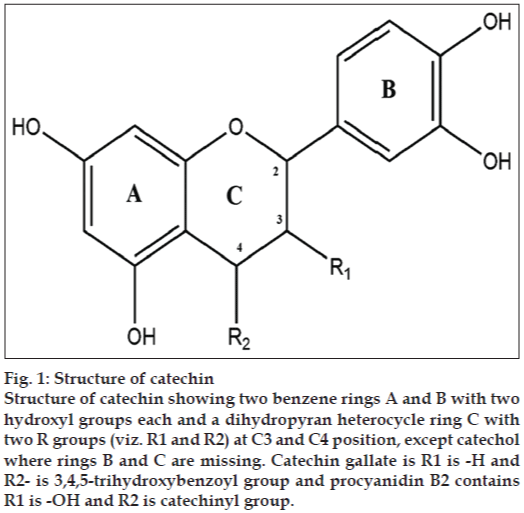
Flavanoids like epicatechin gallate and epigallocatechin gallate, which are constituents of green tea have been reported to change the architecture of the cell wall of S. aureus [6,7]. Moreso, analogs of these compounds viz. cis and trans forms of catechin and epicatechin also exist as naturally occurring flavanoids. Generally the catechin comprise of three aromatic rings. The first two rings are benzene rings with hydroxyl groups while the third is dihydropyran heterocycle with hydroxyl group at carbon 3. As C2 and C3 are chiral, these structure exhibit (fig. 1) stereoisomerism. Scientists have synthesized various analogues by substitution of these hydroxyl groups with other alkyl groups or other chemical moiety. For example by addition of gallate group at C3 catechin gallate, epicatechin gallate, EGCG, has been formed, which are stored in various structure databases like Zinc [8] and Pubchem [9]. Thus in this study, all synthetic and semi-synthetic analogs of catechins have been evaluated as inhibitor to assess their potential to bind with FemA protein.
Figure 1: Structure of catechin
Structure of catechin showing two benzene rings A and B with two hydroxyl groups each and a dihydropyran heterocycle ring C with two R groups (viz. R1 and R2) at C3 and C4 position, except catechol where rings B and C are missing. Catechin gallate is R1 is -H and R2- is 3,4,5-trihydroxybenzoyl group and procyanidin B2 contains R1 is -OH and R2 is catechinyl group.
Materials and Methods
Ligand and protein preparation
FemA protein having 1lrz PDB id was retrieved from RCSB-Protein data bank (www.rcsb.org). Downloaded 1lrz structure was prepared for docking in mol2 and pdbqt format files by deleting all hetero atoms and water molecules by chimera [10] and MGL [11] tools, respectively. Pdb format file of 1lrz having no hydrogen atom was also prepared by Chimera. This file was used as input for generation of ms format file by DMS tool. Similarly, ms format file was used to generate sph format file by sphgen tool.
Ligand structures were retrieved from Pubchem database. Pubchem database consists of 15 catechin and epicatechin analogues, which are available in chemical form. All these molecules were downloaded in sdf format. Similar to protein all ligands were prepared in mol2, pdbqt and pdb format.
Docking studies
It was carried out by two tools, Autodock Vina and UCSF dock. Autodock Vina [12] uses pdbqt input files both for ligands as well as protein. Center grid coordinates used in this study, for Autodock Vina were 35.829×63.416×94.89 while size used for x, y, z were 76×102×20, respectively. Spacing used in Autodock Vina was 1.000 at different exhaustiveness values. While, in case of UCSF Dock 6.5 [13] mol2 files were used and moreover instead of using user defined center coordinates, different sphere clusters were calculated by sphgen software. Sphgen software used dms software output file, which consists of molecular surface of receptor. Total 78 clusters were calculated, out of these, cluster 1 having 111 spheres with their coordinates was used for generation of grid and box files for 1lrz protein. Docking was performed with all ligand files using grid spacing of 0.3 A°.
Rigid docking was performed by both the softwares. The docking orientations are ranked based on a molecular mechanics-like scoring function known as Dock score in case of UCSF Dock, binding energy in case of Autodock Vina. Protein residues forming hydrogen and hydrophobic interactions with ligand molecule is further calculated by Ligplot+ software [14].
Toxicological studies
To determine the toxic level of best FemA inhibitors, toxicity studies was performed by online server, OSIRIS property explorer [15] (http://www.organicchemistry.org/prog/peo/) during toxicological analysis various in silico tests were performed to detect the risk of mutagenicity, carcinogenicity, teratogenicity, irritants and reproductive effects. These predictions depend on comparison between precomputed set of structural moieties whose properties are already known with structural moieties of molecules loaded by user on server.
Lipinski filters
Lipinski filters or Lipinski rule of five distinguishes between drug like and nondrug like molecules. It is also known as Pfizer’s rule of Five. It is a thumb rule to predict the probability of a chemical to be orally active drug in humans. According to this rule molecular mass of molecule should be less than 500 Dalton, lipophilicity represented by LogP should be less than 5, hydrogen bond donors (HBD) should be less than 5, hydrogen bond acceptors (HBA) should be less than 10 and molar refractivity (MR) should be lies between 40-130. Molecule failing in more than 2 rules is considered as nondrug molecule. Lipinski filtering was performed by Lipinski filter tool of SCFBIO at IITDelhi server [16].
Results and Discussion
Docking of FemA protein with catechin and epicatechin analogues have exhibited 100% results with Autodock Vina while 50% with UCSF DOCK. Ligands were ranked on the basis of binding energy and grid score given by Autodock Vina and UCSF Dock 6.5, respectively. As shown in Table 1 both docking tools has predicted (-)catechin gallate with cid 6419835 as best inhibitor having best binding affinity of -8.7 kcal/mol with Autodock Vina and best dock score of -45.19 with UCSF Dock among all 15 analogues. Best conformations of best ligands predicted by both the softwares are shown in fig. 2a and b.
| CID of Pubchem | Name | Grid_score | Binding affinity (kcal/mol) | Dock rank | Vina rank |
|---|---|---|---|---|---|
| 6419835 | (-)-catechingallate | -45.195 | -8.7 | 1 | 1 |
| 73160 | (-)-catechin; catechin L-form | -34.034 | -8.7 | 6 | 2 |
| 5276454 | (-)-catechingallate; (2R,3S)-2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-3, | -44.642 | -8.5 | 2 | 3 |
| 4-dihydro-2Hchromen-3-yl 3,4,5-trihydroxybenzoate; XEG | |||||
| 122738 | Procyanidin B2; Procyanidol B2 | Out of grid | -8.2 | Nil | 4 |
| 182232 | (+)-epicatechin; 35323-91-2; ent-epicatechin | -37.057 | -8.1 | 4 | 5 |
| 12309507 | L-epicatechin | -34.499 | -8.1 | 5 | 6 |
| 9064 | Catechin; cianidanol; (+)-Catechin; D-caechin;(+) cyanidanol | Out of grid | -8.1 | Nil | 7 |
| 65064 | (-)-epigallocatechingallate | Out of grid | -8 | Nil | 8 |
| 107957 | Catechin hydrate;(+)-catechin hydrate | Out of grid | -7.7 | Nil | 9 |
| 107905 | Epicatechingallate; (-)-epicatechingallate; (-)-epicatechin-3-gallate | -43.823 | -7.6 | 3 | 10 |
| 367141 | (-)-epicatechingallate; NSC636594 | Out of grid | -7.4 | Nil | 11 |
| 1203 | DL-catechin; NSC81746; L-epicatechin | Out of grid | -7.3 | Nil | 12 |
| 72276 | Epicatechin; (-)-epicatechin | Out of grid | -7.3 | Nil | 13 |
| 255538 | L-Epicatechin; epicatechol | Out of grid | -7.3 | Nil | 14 |
| 289 | Catechol; pyrocatechol | -20.687 | -4.1 | 7 | 15 |
CID: Compound identifier
Table 1: Docking results of autodock vina and ucsf dock
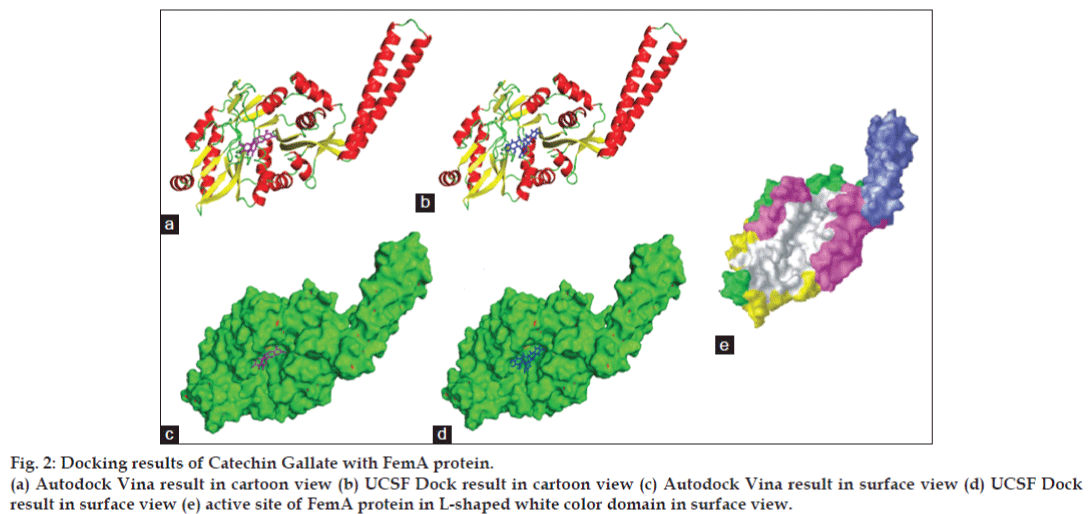
Figure 2: Docking results of Catechin Gallate with FemA protein.
(a) Autodock Vina result in cartoon view (b) UCSF Dock result in cartoon view (c) Autodock Vina result in surface view (d) UCSF Dock result in surface view (e) active site of FemA protein in L-shaped white color domain in surface view.
FemA protein structure consists of one helical arm and one globular domain. Globular domain is further divided into two sub-domains and some secondary structure elements. Helical arm is formed of residues 246-307 and in case of globular domain residues 1–110, 129–144, 396–401 forms first sub-globular domain, and residues 145–166, 189–245and 308–395 forms second sub-globular domain and rest 111–128, 167–188, and 402–412 residues are part of secondary structure elements [5]. Globular domain is similar to HAT domain of Tetrahymena GCN5. Globular domain of FemA protein consists of five additional structural elements that are not found in the structure of GCN5. Two ß strands that extends from first sub-domain of globular domain and two a helices that lies on the top of second sub-domain of globular domain and one more C-terminal a helix. HAT domain of Tetrahymena GCN5 binds with two substrates: Coenzyme A and Peptide [17]. Similar pockets also exist in both sub-domains of globular domain of FemA protein. These pockets form a deep L-shaped channel, which is a suitable site of binding for FemA protein in S. aureus [5]. Helical domain of FemA protein is similar to seryl-tRNA synthetases in bacteria [18,19]. Function of helical domain in FemA and seryl-tRNA synthetases is just to holding of charged amino acid t-RNA molecule during addition of glycine in penta-glycine chain [20-23]. Therefore it is not available for inhibitors, only pockets in L-shaped channel of sub-domains of globular protein is available for inhibitors (fig. 2e).
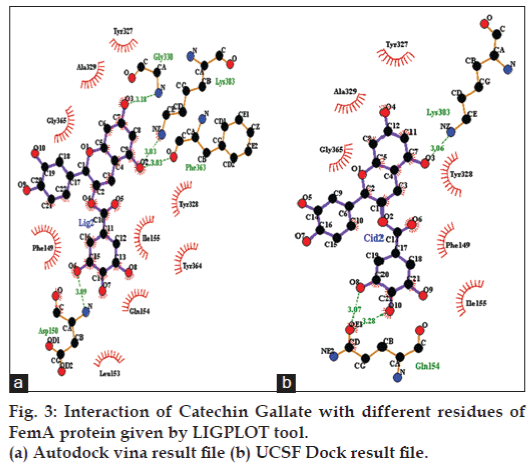
As shown in fig. 2c and d according to both the softwares catechin gallate is binding exactly between the active sites in L-shaped channel of first sub-globular domain of FemA protein. Best ligand conformation predicted by both the docking tools were further used to form complex with protein and these were further used with Ligplot+ to calculate hydrogen and hydrophobic interactions with protein residues. As shown in fig. 3a, according to conformation given by Autodock Vina catechin gallate form hydrogen bonds with GLY 330, LYS 383, PHE 363, ASP 150 residues of FemA protein with distance of 3.18, 3.03, 3.03 and 3.09, respectively (shown in green color), while residues like GLY 365, ALA 329, TYR 327, TYR 328, ILE 155, TYR 364, GLN 154, LEU 153, PHE 149 forms hydrophobic interactions. Similarly, UCSF Dock predicted conformation forms hydrogen bond with LYS 383 and GLN 154 with bond distance 3.06, 3.07 and 3.28, respectively, while there are six residues, which forms hydrophobic interactions, residues are GLY 365, ALA 329, TYR 327, TYR 328, PHE 149 and ILE 155 (fig. 3b). Both softwares predicted catechin gallate as most efficient ligand for FemA protein as compared to other catechin and epicatechin derivatives or flavanoids and have shown LYS 383 residue of FemA protein as a common residue, which forms hydrogen bond and all 6 hydrophobic interacting residues, which were predicted by UCSF Dock were also predicted in Autodock Vina result. It appears that LYS 383 is the most essential residue of FemA protein for interaction with catechin gallate.
Toxicity analysis was performed by OSIRIS tool. According to OSIRIS if score for any category is 1, then it indicates no risk, if it lies between 0.6-0.8 then the chemical may have medium risk but if score is less than 0.6 it is considered as high risk chemical. As shown in Table 2 all the analogues except catechol (s=0.6), have score equal to one for all four risk categories viz.-mutagenicity, tumorigenicity, irritation and reproductive effects. Similarly as predicted by other two categories of OSIRIS drug likeness and drug score, which determine whether particular molecule is similar to the known drugs, same molecule catechol was discarded due to poor scores and rest all the molecules are non-hazardous as well as similar to good drugs. Therefore, except catechol; rest 14 molecules are safe and can be used as future drugs. In toxicity analysis catechin gallate has passed all the tests, therefore it may be a ray of hope for various health problems caused by gram-positive bacteria.
| CID of Pubchem | Name | Drug likeliness | MR | TR | IE | RE | Drug score | |
|---|---|---|---|---|---|---|---|---|
| 6419835 | (-)-catechingallate | 2.81 | (score=0.94) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.74 |
| 73160 | (-)-catechin; catechin L-form | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| 5276454 | (-)-catechingallate | 2.81 | (score=0.94) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.74 |
| 122738 | Procyanidin B2 | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.534 |
| 182232 | (+)-epicatechin; 35323-91-2; | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| ent-epicatechin | ||||||||
| 12309507 | L-epicatechin; | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| 9064 | Catechin; cianidanol; (+) catechin; (+) | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| cyanidanol | ||||||||
| 65064 | (-)-epigallocatechingallate | 1.57 | (score=0.83) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.69 |
| 107957 | Catechin hydrate; (+) catechin hydrate | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| 107905 | Epicatechingallate; (-)-epicatechin | 2.81 | (score=0.94) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.74 |
| gallate | ||||||||
| 367141 | (-)-epicatechingallate; NSC636594 | 1.39 | (score=0.8) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.68 |
| 1203 | DL-catechin; NSC81746; L-epicatechin | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| 72276 | Epicatechin; (-)-epicatechin | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| 255538 | L-epicatechin; epicatechol | 1.92 | (score=0.87) | No (score=1) | No (score=1) | No (score=1) | No (score=1) | 0.86 |
| 289 | Catechol; pyrocatechol | -2.25 (score=0.09) | Yes (score=0.6) | Yes (score=0.6) | Yes (score=0.6) | No (score=1) | 0.112 | |
CID: Compound identifier, MR: mutagenicity risk, TR: tumorigenicity risk, IE: irriating effects, RE: reproductive effects
Table 2: Toxicity results predicted by osiris
As shown in Table 3, two molecules have violated two Lipinski rules, while 6 molecules have violated one rule and rest seven have not violated any Lipinski rule. All the violations are shown in red color and summarized in last category under number of Lipinski value (LV). Pyrocyanidin B2 and EGCG have high chances of failure in preclinical and clinical trials, while rest 13 molecules seems to have very less or no chances of failure.
| CID of Pubchem | Name | Molecular weight | HBA | HBD | LogP | MR | Number of LV |
|---|---|---|---|---|---|---|---|
| 6419835 | (-)-catechingallate | 442 | 3 | 7 | 3 | 96.3 | 1 |
| 73160 | (-)-catechin; catechin L-form | 290 | 1 | 5 | 0.52 | 64.6 | 0 |
| 5276454 | (-)-catechingallate;(2R,3S)-2-(3,4-dihydroxyphenyl)-5,7-dihydroxy-3, | 442 | 3 | 7 | 3 | 96.3 | 1 |
| 4-dihydro-2Hchromen-3-yl 3,4,5-trihydroxybenzoate | |||||||
| 122738 | Procyanidin B2; procyanidol B2; | 578 | 2 | 10 | 0.9 | 108 | 2 |
| 182232 | (+)-epicatechin; 35323-91-2; ent-epicatechin | 290 | 1 | 5 | 0.52 | 64.6 | 0 |
| 12309507 | L-epicatechin | 290 | 1 | 5 | 0.52 | 64.6 | 0 |
| 9064 | Catechin; cianidanol; (+)-catechin; D-catechin;(+) cyanidanol | 290 | 6 | 5 | 0.4 | NA | 0 |
| 65064 | (-)-epigallocatechingallate | 458 | 11 | 8 | 1.2 | NA | 2 |
| 107957 | Catechin hydrate; (+)-catechin hydrate | 308 | 7 | 6 | NA | NA | 1 |
| 107905 | Epicatechingallate; (-)-epicatechingallate; (-)-epicatechin-3-gallate | 442 | 3 | 7 | 3 | 96.3 | 1 |
| 367141 | (-)-epicatechingallate; NSC636594 | 442 | 10 | 7 | 1.5 | NA | 1 |
| 1203 | DL-catechin; NSC81746; L-epicatechin | 290 | 6 | 5 | 0.4 | NA | 0 |
| 72276 | Epicatechin; (-)-epicatechin | 290 | 1 | 5 | 0.52 | 64.6 | 0 |
| 255538 | L-epicatechin; epicatechol | 290 | 1 | 5 | 0.52 | 64.6 | 0 |
| 289 | Catechol; pyrocatechol | 110 | 0 | 2 | 1.1 | 29.8 | 1 |
CID: Compound identifier, HBA: hydrogen bond acceptors, HBD: hydrogen bond donors, MR: molar refractivity, LV: Lipinski value, NA: not available
Table 3: Lipinski results
Ligand molecule catechin gallate Pubchem cid 6419835 has been predicted as a potential inhibitor of FemA protein; by two best tools in terms of binding affinity and docking scores and has also shown nontoxic behavior in toxicity tests and also follows Lipinski rule of five. Therefore, if it can be analysed in in vitro studies and chances of failure in preclinical and clinical trials seem to be very less and it can be a ray of hope for the treatment of various S. aureus infections.
Acknowledgement
We are thankful to UCSF Lab and Scripps Research Institute Lab as well as OSIRIS and SCFBIO web servers for providing these softwares academically free. Special thanks to Ligplot team for providing academic license of this software.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
References
- Mallorquí-Fernández G, Marrero A, García-Piquè S, García-Castellanos R, Gomis-Rüth FX. Staphylococcal methicillin resistance: Fine focus on folds and functions. FEMS MicrobiolLett 2004;235:1-8.
- Ehlert K, Schröder W, Labischinski H. Specificities of FemA and FemB for different glycine residues: FemB cannot substitute for FemA in staphylococcal peptidoglycan pentaglycine side chain formation. J Bacteriol 1997;179:7573-6.
- Rohrer S, Ehlert K, Tschierske M, Labischinski H, Berger-Bächi B. The essential Staphylococcus aureus gene fmhB is involved in the first step of peptidoglycan pentaglycineinterpeptide formation. ProcNatlAcadSci USA 1999;96:9351-6.
- Strandén AM, Ehlert K, Labischinski H, Berger-Bächi B. Cell wall monoglycine cross-bridges and methicillin hypersusceptibility in a femAB null mutant of methicillin-resistant Staphylococcus aureus. J Bacteriol 1997;179:9-16.
- Benson TE, Prince DB, Mutchler VT, Curry KA, Ho AM, Sarver RW, et al. X-ray crystal structure of Staphylococcus aureusFemA. Structure 2002;10:1107-15.
- Stapleton PD, Shah S, Ehlert K, Hara Y, Taylor PW. The beta-lactam-resistance modifier (-)-epicatechingallate alters the architecture of the cell wall of Staphylococcus aureus. Microbiology 2007;153:2093-103.
- Ando C, Kono K, Tatara I, Takeda S, Arakawa K. Antibacterial activity of epigallocatechingallate against staphylococci. Med Bull Fukuoka Univ 1999;26:195-7.
- Bolton E, Wang Y, Thiessen PA, Bryant SH. PubChem: Integrated Platform of Small Molecules and Biological Activities. Chapter 12 in Annual Reports in Computational Chemistry. Washington, DC: American Chemical Society; 2008.
- Irwin JJ, Shoichet BK. ZINC – A free database of commercially available compounds for virtual screening. J ChemInf Model 2005;45:177-82.
- Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, et al. UCSF Chimera – A visualization system for exploratory research and analysis. J ComputChem 2004;25:1605-12.
- Johnson GT, Autin L, Goodsell DS, Sanner MF, Olson AJ. ePMV embeds molecular modeling into professional animation software environments. Structure 2011;19:293-303.
- Trott O, Olson AJ. AutoDockVina: Improving the speed and accuracy
- of docking with a new scoring function, efficient optimization, and multithreading. J ComputChem 2010;31:455-61.
- Lang PT, Brozell SR, Mukherjee S, Pettersen EF, Meng EC, Thomas V, et al. DOCK 6: Combining techniques to model RNA-small molecule complexes. RNA 2009;15:1219-30.
- Laskowski RA, Swindells MB. LigPlot+: Multiple ligand-protein interaction diagrams for drug discovery. J ChemInf Model 2011;51:2778-86.
- Sander T, Actelion Pharmaceuticals Ltd., Gewerbestrasse 16, 4123 Allschwil, Switzerland. Organic Chemistry Portal; 2014. Available from: http://www.organic-chemistry.org/prog/peo/. [Last accessed on 2013 Jan 01].
- Jayaram B. Scfbioiitd Web Server. Available from: http://www.scfbioiitd. res.in/utility/lipinskiFilters.jsp. [Last accessed on 2013 Jan 01].
- Rojas JR, Trievel RC, Zhou J, Mo Y, Li X, Berger SL, et al. Structure of Tetrahymena GCN5 bound to coenzyme A and a histone H3 peptide. Nature 1999;401:93-8.
- Cusack S, Berthet-Colominas C, Härtlein M, Nassar N, Leberman R. A second class of synthetase structure revealed by X-ray analysis of Escherichia coli seryl-tRNAsynthetase at 2.5 A. Nature 1990;347:249-55.
- Fujinaga M, Berthet-Colominas C, Yaremchuk AD, Tukalo MA, Cusack S. Refined crystal structure of the seryl-tRNAsynthetase from Thermusthermophilus at 2.5 A resolution. J MolBiol 1993;234:222-33.
- Kamiryo T, Matsuhashi M. Sequential addition of glycine from glycyl-tRNA to the lipid-linked precursors of cell wall peptidoglycan in Staphylococcus aureus. BiochemBiophys Res Commun 1969;36:215-22.
- Kamiryo T, Matsuhashi M. The biosynthesis of the cross-linking peptides in the cell wall peptidoglycan of Staphylococcus aureus. J BiolChem 1972;247:6306-11.
- Katz W, Matsuhashi M, Dietrich CP, Strominger JL. Biosynthesis of the peptidoglycan of bacterial cell walls. IV. Incorporation of glycine in Micrococcus lysodeikticus. J BiolChem 1967;242:3207-17.
- Thorndike J, Park JT. A method for demonstrating the stepwise addition of glycine from transfer RNA into the murein precursor of Staphylococcus aureus. BiochemBiophys Res Commun 1969;35:642-7.