- *Corresponding Author:
- S. M. Mandal
Central Research Facility, Indian Institute of Technology Kharagpur, Kharagpur-721 302, India
E-mail: mandalsm@gmail.com
| Date of Submission | 28 December 2016 |
| Date of Revision | 17 August 2017 |
| Date of Acceptance | 01 April 2018 |
| Indian J Pharm Sci 2018;80(3):547-551 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
Sepsis is one of the major cause of morbidity and mortality from infants to adults. It is most important to determine the infected bacterial species found in bloodstream and its antibiotic susceptibility pattern for appropriate treatment. Overall fifteen hundred patients’ data were incorporated into our study, collected from Sanjiban Hospital, Howrah, West Bengal, India. Out of 1500 samples, 190 neonates and 250 adults were positive for bacterial sepsis. The strains expressing extended-spectrum β-lactamases are major threat as therapeutic options are limited. Among the isolates, Gram-positive bacteria were predominated (53.36 %) over Gram-negative bacteria (46.64 %) and Staphylococcus aureus was the most frequent isolate (37.6 %) followed by Pseudomonas aeruginosa (16 %), Escherichia coli (13.60 %), Klebsiella sp. (12.8 %), coagulase-negative Staphylococci (12.4 %), Acinetobacter sp. (4 %) and Enterococci sp.(2.8 %). Levofloxacin was revealed to be more active against all the Gram-negative isolates along with carbapenems, aminoglycosides (except Klebsiella sp.) and polymyxin-resistant strains. Levofloxacin gave superior coverage against both Gram-positive and Gram-negative bacteria whereas most of the penicillins and cephalosporins were found to be ineffective against both Gram-positive and Gram-negative isolates.
Keywords
Bacteraemia, antibiotic resistance, blood stream infection, levofloxacin
Blood stream infection (BSI) is a significant reason of increased morbidity and mortality, one of the most frequently observed cases in the tertiary hospitals [1]. It causes millions of deaths globally each year [2]. In case of neonatal sepsis it contributes nearly 13-15 % of all deaths and in developing countries where it contributes between 30-50 % [3]. The causative microorganisms of BSI have been found to be associated with increased drug resistance propensity to the conventionally used antibiotics and emergent global threats to control those multi-drug resistant pathogens [4]. Several antibiotics have been successfully used for the treatment of sepsis, but recently most of the Gram-negative bacilli responsible for BSI became a serious therapeutic challenge due to development of multidrug resistance [5]. In most cases of the neonatal sepsis, extended-spectrum β-lactamase (ESBL) producing Gram-negative bacilli and methicillin-resistant Staphylococcus aureus (MRSA) further complicated the treatment resulting in higher mortality and morbidity rate [6-8]. In addition, Klebsiella pneumoniae carbapenemases, an ESBL capable of hydrolysing a broad spectrum β-lactam antibiotics including third-generation cephalosporins. These are plasmid-mediated enzymes, which can be transferred to other Gram-negative species, such as Escherichia coli and Pseudomonas sp. [9,10]. Such plasmidmediated enzymes further complicate antimicrobial therapy [11]. Such isolates often exhibit resistance to other classes of drugs such as aminoglycosides (gentamicin, amikacin), cotrimoxazole, tetracycline and fluoroquinolones, which create a dreadful challenge with restricted therapeutic options especially in India [12]. Some studies revealed that few bacteria (E. coli and K. pneumoniae) become highly resistant as they produce metallo-β-lactamases, which make the bacteria resistant to a broad range of β-lactams including the carbapenems [13]. Presence of efflux pumps also explains high-level intrinsic resistance against a broad variety of substrates. The overproduction of the efflux pump is usually accompanied by an increase in resistance to two or more structurally unrelated antibiotics and significantly contribute to the emergence and spread of multi-drug resistance pathogens [14]. The expression of these efflux pumps are important for pathogens, which can be coupled to other mechanisms of resistance [15,16].
Therefore, these hospital-based descriptive studies are essential for determining the pathogens prevalent in the respective hospital community and their resistance pattern to various antibiotics need to be documented. This will in turn help in formulating guidelines for management of BSI in local hospitals and the choice of antibiotic therapy in the community as well.
This study was done using the data of blood culture and antibiograms carried out at Sanjiban Hospital in Howrah district, West Bengal, India. All data were retrospectively collected and de-identified to ensure patient confidentiality. All the media, reagents and antibiotic-discs were procured from HiMedia, Mumbai, India.
A total number of 1500 blood culture samples were included in this study. This study assessed resistance in a total number of 30 antimicrobials. Two sets of blood specimens were collected from independent venepuncture sites of about 08-10 ml of blood for adult and for a child depending upon age but less than 1 % of total blood volume. The samples were sent to the laboratory as soon as possible after blood was aseptically introduced into aerobic media, brain heart infusion broth with sodium polyanethol sulfonate [17]. The inoculated media were incubated for 7 d at 37° and blood cultures were considered negative if there was no growth after subcultures being made each day. For positive cultures, the identification of the isolates was performed through biochemical analysis and antimicrobial sensitivity test of the bacterial isolates were determined following Kirby-Bauer disc diffusion method as per CLSI guidelines [14-18].
ESBL detection was carried out using routine susceptibility test following the method recommended by the NCCLS. Two discs, ceftazidime (30 μg) and cefotaxime (30 μg), were used. An inhibition zone of ≤22 mm for ceftazidime and ≤27 mm for cefotaxime indicated that the strain probably produced ESBL.
Confirmation test was also conducted following NCCLS recommendations, on Mueller-Hinton agar. Four discs, containing cefotaxime (30 μg), cefotaxime/ clavulanic acid (30/10 μg), ceftazidime (30 μg) and ceftazidime/clavulanic acid (30/10 μg), were used. An inhibition zone of ≥5 mm increase in diameter for either antimicrobial tested in combination with clavulanic acid versus its zone when tested alone confirmed ESBL production [19].
Blood culture samples were taken from patients who were admitted with clinical features suggestive of sepsis in our different wards including intensive care unit and neonatal care unit. A total of 1500 samples were critically screened and analysed. Among them, 190 samples were from neonatal patients with 15.2 % positive (38) for bacterial growth and out of the total blood cultures, 250 (16.67 %) were positive for bacterial growth, among them 123 (E. coli: 11, Klebsiella sp.: 27, Acinetobacter sp.: 7, Pseudomonas sp.: 21 and S. aureus: 57) isolates were ESBL positive and 2 were positive for Candida sp.
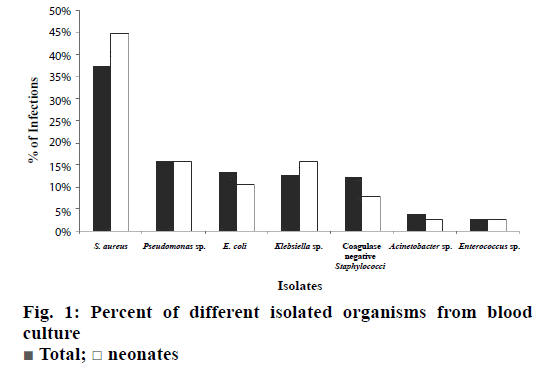
Amongst the 250 positive cultures, Gram-positive sp. (53.36 %) was predominant over Gram-negative sp. (46.64 %). By analysing the data, it was found that S. aureus was the most common bacterial pathogen causing BSI and accounted for 37.6 % of the total isolates. The other bacteria were, Pseudomonas sp. 16 %, E. coli 13.6 %, Klebsiella sp. 12.8 %, coagulase-negative Staphylococci 12.4 %, Acinetobacter sp. 4 % and Enterococcus sp. 2.8 % (Figure 1).
The antibiotic sensitivity patterns of the isolates were shown in Table 1. S. aureus was most sensitive to imipenem (98.76 %), meropenem (98.24 %), levofloxacin (97.56 %) but resistant to cefepime (18.75 %). The antibiotic susceptibility pattern of Pseudomonas sp. revealed sensitivity to levofloxacin (97.36 %), along with antiPseudomonas β-lactams, ceftazidime and cefoperazone-sulbactam. Most β-lactams were found ineffective against Pseudomonas sp. In case of E. coli, all the isolates were sensitive to levofloxacin, most were sensitive to amikacin and less than 40 % isolates were found sensitive to β-lactam antibiotics. Almost similar results were found with the Pseudomonas isolates. The sensitivity profile of Klebsiella isolates were quite unlike comparing with Pseudomonas and E. coli isolates. Most of the Klebsiella isolates were found sensitive to imipenem, meropenem and polymixins. Nearly 60 % of the isolates were found sensitive to levofloxacin and 25 % strains were sensitive to amikacin. Most β-lactams were found ineffective against Klebsiella isolates. Like S. aureus, coagulase-negative Staphylococci were found susceptible to levofloxacin along with meropenem, imipenem, vancomycin, linezolid, clindamycin amoxicillin-clavulinic acid. The antibiotic sensitivity pattern of Acinetobacter isolates indicated sensitivity to tigecycline, doxycycline, polymyxin B, colistin (100 %), and a few of them were also sensitive to meropenem, imipenem, and levofloxacin. In case of Enterococci, it was found that all the isolates were sensitive in vitro to imipenem, vancomycin, linezolid, levofloxacin, gentamycin. Very few number of isolates were found sensitive to erythromycin (40 %), azithromycin (20 %).
| Bacteria | PI | AMC | CXM | CX | CTR | CPM | CTX | IPM | MRP | E | AZM | VA | CD | LZ |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| S. aureus | - | 77.38 | 59.37 | 50 | 87.5 | 18.75 | 73.07 | 98.76 | 98.24 | 41.11 | 43.75 | 97.81 | 81.05 | 97.8 |
| Psudomonas sp. | 42.1 | 37.14 | 61.11 | - | 64.87 | 24.13 | 72.73 | 83.33 | 97.14 | - | - | - | - | - |
| E. coli | 41.18 | 60.72 | 23.33 | - | 34.75 | 22.22 | 46.15 | 96 | 89.47 | - | - | - | - | - |
| Klebsiella sp. | 14.32 | 20.68 | 10.35 | 31.04 | 17.39 | 29.3 | 95.28 | 60 | ||||||
| Staphylococcus | - | 85.18 | 66.67 | 29 | 71.42 | 100 | 100 | 42.4 | 45.83 | 100 | 90.33 | 100 | ||
| Coagulase (-ve) | ||||||||||||||
| Acinetobacter sp. | 11.25 | 20 | - | - | 10 | - | - | 86.5 | 87.5 | - | - | - | - | - |
| Enterococci | - | 85.71 | 100 | 33.33 | - | - | 83.33 | 100 | 83.33 | 40 | 20 | 100 | 42.86 | 100 |
| Bacteria | LE | GEN | RIF | COT | PIT | CZ | CAZ | CFS | OF | TGC | DO | CL | AK | |
| S. aureus | 97.56 | 92.77 | 82.3 | 68.23 | - | - | - | - | - | - | 97.59 | - | - | |
| Psudomonas sp. | 97.36 | - | - | 78.37 | 60.53 | 8 | 48.48 | 44.44 | 11.9 | 100 | 91.42 | 68 | - | |
| E. coli | 100 | 76.93 | - | 64.28 | 68.96 | 42.86 | - | - | 53.58 | 100 | 85.72 | - | 96.3 | |
| Klebsiella sp. | 65.25 | 32.14 | 38.46 | 22.24 | 8.1 | - | - | 52 | 100 | 76.92 | 70.25 | 25.2 | ||
| Staphylococcus, Coagulase (-ve) | 96.42 | 88.89 | 87.88 | 69.23 | - | - | - | - | - | - | 96.29 | - | - | |
| Acinetobacter sp. | 77.75 | 25 | - | 28.5 | 22.5 | - | - | - | 44.5 | 100 | 100 | 100 | 66.75 | |
| Enterococci | 100 | 100 | - | - | - | - | - | - | - | - | 100 | - | - | |
Abbreviations are AMC: amoxicillin-clavulanate, PI: piperacillin, PIT: piperacillin-tazobactam, CZ: cefazolin, CXM: cefuroxime, CTR: ceftriaxone, CTX: cefotaxime, CAZ: ceftazidime, CFS: cefoperazone-sulbactam, CPM: cefepime, CX: cefoxitin, IPM: imipenem, MRP: meropenem, OF: ofloxacin, LE: levofloxacin, GEN: gentamicin, AK: amikacin, CL: colistin, DO: doxycycline, TGC: tigecycline, E: erythromycin, AZM: azithromycin, VA: vancomycin, CD: clindamycin, LZ: linezolid, RIF: rifampicin, COT: cotrimoxazole
Table 1: Percent antimicrobial susceptibility of different isolates
The antimicrobial resistance patterns of ESBLproducing isolates were presented in Table 2. Among the total bacterial isolates (250), ESBL producing strains (123) were 49.20 % where individual strains were E. coli (39.28 %), Klebsiella sp. (77.14 %), Acinetobacter sp. (100 %), Pseudomonas sp. (52.5 %) and S. aureus (56 %), a significant number of percentage. All the ESBL positive strains are resistant to cephalosporins, ceftazidime and cefotaxime and sensitive to carbapenems, meropenem or imipenem. Few of them showed resistant to both cephalosporins and carbapenems, which could be the cause of crossresistance or by the production of other β-lactamases.
| Antibacterial drugs | E. coli (n=11) | Klebsiella sp. (n=27) | Acinetobacter sp. (n=7) | Pseudomonas sp. (n=21) | S. aureus (n=57) |
|---|---|---|---|---|---|
| Ampicillin | 81.81 | 100 | 100 | 90.47 | 82.45 |
| Piperacillin | 63.63 | 74.07 | 100 | 80.95 | 71.92 |
| Ticarcillin | 63.63 | 29.69 | 100 | 52.38 | 73.68 |
| Amox/clavulanic acid | 27.27 | 7.40 | 100 | 47.61 | 36.84 |
| Ticarcillin/clavulanic acid | 36.36 | 7.40 | 100 | 38.09 | 45.61 |
| Piperacillin/tazobactem | 27.27 | 40.74 | 100 | 42.85 | 35.08 |
| Imipenam | 0 | 7.40 | 14.28 | 14.28 | 12.28 |
| Meropenam | 0 | 7.40 | 14.28 | 9.52 | 7.01 |
| Ceftazidime | 72.72 | 77.77 | 71.28 | 52.38 | 43.85 |
| Cephotaxime | 72.72 | 70.37 | 71.28 | 47.61 | 36.84 |
| Cefixime | 36.36 | 25.95 | 71.28 | 61.90 | 57.89 |
| Ceufuroxime | 18.18 | 11.11 | 57.14 | 47.61 | 36.84 |
| Cefpodoxime | 18.18 | 22.22 | 57.14 | 47.61 | 43.85 |
| Cotrimoxazole | 45.45 | 40.74 | 85.71 | 38.09 | 49.12 |
| Levofloxacin | 0 | 0 | 28.51 | 19.04 | 5.26 |
| Amikacin | 9.09 | 11.11 | 71.48 | 23.80 | 15.78 |
| Tobramycin | 18.18 | 22.22 | 71.48 | 33.33 | 43.85 |
Table 2: Percent antimicrobial resistance of esbl producers from different isolated bacterial pathogens
The Surviving Sepsis Guidelines state that initial empiric antibiotic therapy is one or more drugs, which are active against all likely pathogens (bacterial and/ or fungal or viral) and that penetrate in adequate concentrations into tissues presumed to be the source of sepsis (grade 1B). To ensure both the above, locally prevalent pathogens and their antibiograms should be made available to practice evidence-based medicine. The present study revealed a change in the sensitivity pattern of the common pathogens to commonly used antibiotics in these hospitals. In the last decade, ESBLs were of great medical importance as these hydrolyse -lactam antibiotics. In recent years, the spread and incidence of ESBLs have increased due to the overuse of expanded-spectrum cephalosporins in the hospital setting. It is a major therapeutic challenge to treat infections by strains of Gram-negative bacteria that express ESBLs.
Third generation quinolone such as levofloxacin was the cheapest amongst the most potent antimicrobial agents effective against both Gram-negative and positive organisms in this unit and the least sensitive antibiotics were ampicillin and the first generation cephalosporins. The third generation cephalosporins are commonly used as second line antibiotics in many centers (and even as first line therapy in some centers) have been observed to become increasingly ineffective as shown in the present study, which is comparable to published reports [12,20]. However, Klebsiella sp. showed different results, 25 % sensitivity to amikacin and 32.14 % sensitivity to gentamicin that supported some earlier reports. Increasing resistance pattern of aminoglycosides to Klebsiella sp. has also been reported by other observers [21].
Therefore, the quinolones, especially levofloxacin was found to be the most potent empirical antibiotic for sepsis patients. The major limiting factors of the third generation cephalosporins were high-price and toxicity. Amongst aminoglycosides, even in the face of gentamicin resistance, amikacin might be recommended because of its good susceptibility patterns. The carbapenems especially meropenem was found to be collectively effective in almost all types of Gram-positive and negative bacterial isolates in this investigation. However, the high cost of therapy with meropenem advocate the use of levofloxacin as an initial empirical therapy in and around this region. Furthermore, steps needed to be taken to prevent or control the emergence of resistance strains by using antibiotics rationally. Laws therefore should be enforced to discourage the indiscriminate use of antibiotics seen commonly in India as well as discourage inadequate doses through proper training of personnel in order to prevent emergence of resistant strains. In the present study resistance to three or more antibiotics was common in both ESBL producers and non-ESBL producers. About 80 % strains were resistant to penicillins and cephalosporins. Septicaemia is a leading cause of mortality when caused by ESBLpositive strains. The enormous use of cephalosporins might become one of the major factor for increasing rate of ESBL producing microorganisms [22].
In conclusion, there is an overall shift in the antibiotic susceptibility to the commonly isolated bacterial pathogens. Antibiotic resistance becoming a major public health concern and hazard all over the world, so it is imperative to take more caution and evidencebased judgment, while starting antibiotics therapy. From this study, it could be concluded that levofloxacin is the drug that should be advocated when starting antibiotic empirically for BSI followed by amikacin as these were found to give good coverage against both Gram-positive and Gram-negative organisms. Increased awareness of the ESBL problem among hospital settings is crucial to control them as soon as possible.
Acknowledgements
Authors acknowledge the Director of Sanjiban Hospital for granting permission and support. Special thanks are to Mr. Amit Chakraborty and Miss Anannya Ghosh for their help throughout the work.
Conflict of interest
Authors have no any conflict of interest and none financial involvement in this work.
Financial support and sponsorship
Nil.
References
- Del Rio A, Cervera C, Moreno A, Moreillon P, Miro JM. Patients at risk of complications of Staphylococcus aureus bloodstream infection. Clin Infect Dis 2009;48:S246-S53.
- Dellinger RP, Levy MM, Rhodes A, Annane D, Gerlach H, Opal SM, et al. Surviving Sepsis Campaign: international guidelines for management of severe sepsis and septic shock. Crit Care Med 2008;34:17-60.
- West BA, Peterside O. Sensitivity pattern among bacterial isolates in neonatal septicaemia in Port Harcourt. Ann Clin Microbiol Antimicrob 2012:11:7-13.
- Marin H, Kollef MH, Golan Y, Micek ST, Shorr AF, Restrepo MI. Appraising contemporary strategies to combat multidrug resistant gram-negative bacterial infections–proceedings and data from the Gram-Negative Resistance Summit. Clin Infect Dis 2011;53:S33-S5.
- Suárez CJ, Lolans K, Villegas MV, Quinn JP. Mechanisms of resistance to beta lactams in some common gram negative bacteria causing nosocomial infection. Expert Rev Anti Infect Ther 2005;3:915-22.
- Jain A, Roy I, Gupta MK, Kumar M, Agarwal SK. Prevalence of extended-spectrum beta-lactamase-producing Gram-negative bacteria in septicaemic neonates in a tertiary care hospital. J Med Microbiol 2003;52:421-25.
- Kim YK, Pai H, Lee HJ, Park SE, Choi EH, Kim J, et al. Bloodstream infections by extended-spectrum beta-lactamase-producing Escherichia coli and Klebsiella pneumoniae in children: epidemiology and clinical outcome. Antimicrob Agents Chemother 2002;46:1481-91.
- Kapil A. Risk factors associated with MRSA infection in Children. Indian Pediatr 2015;52:22-4.
- Navon-Venezia S, Chmelnitsky I, Leavitt A, Schwaber MJ, Schwartz D, Carmeli YA. Plasmid-mediated imipenem-hydrolyzing enzyme KPC-2 among multiple carbapenem-resistant Escherichia coli clones in Israel. Antimicrob Agents Chemother 2006;50:3098-101.
- Villegas MV, Lolans K, Correa A, Kattan JN, Lopez JA, Quinn JP. Nosocomial Resistance Study Group: First identification of Pseudomonas aeruginosa isolates producing a KPC-type carbapenem-hydrolyzing beta-lactamase. Antimicrob Agents Chemother 2007;51:1553-55.
- Pitout JD. The latest threat in the war on antimicrobial resistance. Lancet Infect Dis 2010;9:578-9.
- Chandel DS, Johnson JA, Chaudhry R, Sharma N, Shinkre N, Parida S, et al. Extended-spectrum beta-lactamase producing Gram-negative bacteria causing neonatal sepsis in India in rural and urban settings. J Med Microbiol 2011;60:500-07.
- Kumarasamy KK, Toleman MA, Walsh TR, Bagaria J, Butt F, Balakrishnan R, et al. Emergence of a new antibiotic resistance mechanism in India, Pakistan, and the UK: a molecular, biological, and epidemiological study. Lancet Infect Dis 2010;10:597-602.
- Sarkar SK, Bhattacharyya A, Mandal SM. YnfA, a SMR family efflux pump is abundant in Escherichia coli isolates from urinary infection. Indian J Med Microbiol 2015;33:139-42.
- Wen-Sen L, Wei-Yu C, Tsong-Yih O, Fu-Lun C, Yung-Wei L. Malakoplakia in a patient with complicated urinary tract infection caused by extended-spectrum β-lactamase-producing Escherichia coli. J Microbiol Immunol Infect 2015;48:345-6.
- Doumith M, Ellington MJ, Livermore DM, Woodford N. Molecular mechanisms disrupting porin expression in carbapenem-resistant Klebsiella and Enterobacter sp. clinical isolates from the UK. J Antimicrob Chemother 2009;63:659-7.
- Wasilauskas BL, Floyd J, Richard Roberts T. Use of Sodium Polyanethol Sulfonate in the preparation of 5% Sheep Blood Agar Plates. Appl Microbiol 1974;28:91-4.
- Performance Standards for antimicrobial susceptibility testing, Seventeenth informational supplement. Clinical and Laboratory Standards Institute 2007;M-100 S-16(26):3.
- Deepti R, Deepthi N. Extended-spectrum β-lactamases in Gram Negative Bacteria. J Glob Infect Dis 2010;2:263-74.
- Gea-Banacloche JC, Opal SM, Jorgensen J, Carcillo JA, Sepkowitz KA, Cordonnier C. Sepsis associated with immunosuppressive medications: An evidence-based review. Crit Care Med 2004;32:578-90.
- Livermore DM, Yuan M. Antibiotic resistance and production of extended-spectrum Beta lactamases amongst Klebsiella spp. from intensive care units in Europe. J Antimicrob Chemother 1996;38:409-24.
- Paterson DL, Bonomo RA. Extended-spectrum beta-lactamases: a clinical update. Clin Microbiol Rev 2005;18:657-86.
 Total;
Total;  neonates
neonates