- *Corresponding Author:
- P. S. Gatne
Department of Pharmaceutical Chemistry, Bombay College of Pharmacy, Kalina, Santacruz, Mumbai-400 098
E-mail: parag507@gmail.com
| Date of Submission | 21 October 2009 |
| Date of Revision | 1 July 2010 |
| Date of Acceptance | 1 October 2010 |
| Indian J. Pharm. Sci., 2010, 72 (5): 663-667 |
Abstract
Eight novel 1-(substituted acetyl)-4-(10-bromo-8-chloro-5,6-dihydro-11H-benzo[5,6] cyclohepta [1,2-b] pyridine-11-ylidene)piperidines were designed by incorporating zinc binding groups to enhance activity. The designed molecules were synthesized and were evaluated for antitumor activity in vitro in five cell lines and for farnesyl protein transferase inhibition. Test compounds (6a-h) exhibited antitumor activity in most of the cell lines but were less potent than adriamycin. Compound 6e was most active with IC 50 values of <15 μM in two cell lines tested. Test compounds also exhibited potent FPT inhibitory activity and 6c was most potent with IC 50 value of <30 μM.
Keywords
Anticancer agents, benzocycloheptapyridines, farnesyl protein transferase inhibitors
An important therapeutic approach in cancer is to inhibit farnesylation of ras and this can be achieved by inhibition of farnesyl protein transferase (FPT) enzyme using inhibitors (FTIs) [1]. FTIs belonging to various structural classes were developed and among these, lonafarnib® (benzocycloheptapyridines) [2], tipifarnib® (4,6-disubstituted-quinoline-2-ones) [3] and BMS-241662 (1,3,4-trisubstituted-2,3,4,5-tetrahydro- 1,4-benzodiazepines) [4] are potent molecules and are currently undergoing phase II and III clinical trials.
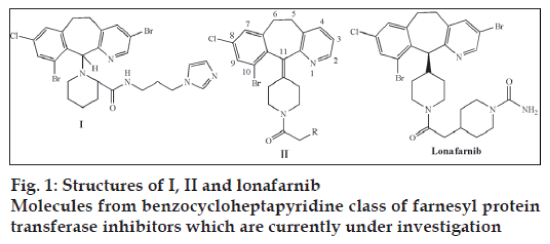
Among benzocycloheptapyridine class of FTIs, the tricyclic nucleus, a bromine at C10 position on the ring, a piperidine ring at C11 position linked to heteroaryl systems by a –COCH2 linker are important structural parameters for activity. It is also reported that the incorporation of groups with zinc binding ability at 2-position of the piperidine ring can enhance activity (structure I, fig. 1) [5].
Hence in the present work, it is proposed to develop a series of novel benzocycloheptapyridines with structure II (fig. 1) where R is a group of heteroaryl ring systems. It is also proposed that the group R in structure II would occupy the zinc binding region in FPT and thereby exhibit potent activity. Hence for designing of novel molecules, the ‘Flexible Alignment’ tool of Molecular Operating Environment 2006.08 (MOE, Chemical Computing group, USA) software was used on a Windows platform with a Pentium IV Dual Core processor computer (2.9 GHz and 1 GB RAM).
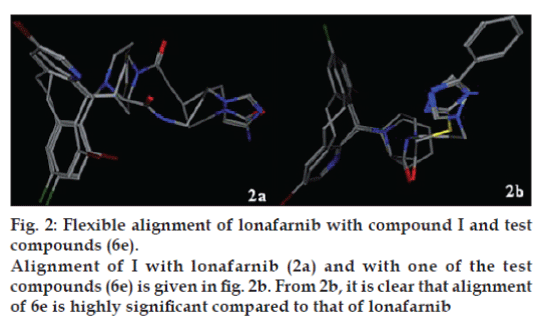
Conformers of I, lonafarnib® and test molecules (6a -h) (general structure II) were generated by using dynamic simulations at the temperature of 310º and the sampling time was 5×10-4 s and the lowest energy conformations were identified for each molecule. Keeping I as template, lonafarnib and 6a-h were superimposed with similarity terms being Aromaticity, Hydrophobe, LogP (o/w) and Hydrogen bond acceptor. Similarity index (F); Average strain energy (U) and Alignment score (S) were measured and are given in Table 1. Lower scores indicate greater similarity and better alignment. Alignment of I with lonafarnib (fig. 2a) and with one of the test compounds (6e) is given in fig. 2b. From fig. 2b, it is clear that alignment of 6e is highly significant compared to that of lonafarnib.
| Compound | Scores | |||
|---|---|---|---|---|
| F | S | U | ||
| 6a | 179.31 | 293.51 | 114.20 | |
| 6b | 176.94 | 298.13 | 126.61 | |
| 6c | 170.74 | 272.01 | 113.10 | |
| 6d | 172.10 | 286.50 | 109.86 | |
| 6e | 167.72 | 276.68 | 103.95 | |
| 6f | 165.33 | 273.55 | 108.22 | |
| 6g | 169.72 | 276.25 | 106.00 | |
| 6h | 174.30 | 281.84 | 101.42 | |
| Lonafarnib | 164.95 | 253.45 | 78.65 | |
Table 1: Scores obtained for flexible alignment study
LoratadineTM was obtained as a gift sample from Themis laboratories, Thane, India and was used as a starting material for the synthesis of 6a-h. 3 [H] FPP, H-ras protein and FPT required for the assay were purchased from Sigma Aldrich (USA). Desloratadine was obtained as a gift sample from Glenmark Pharmaceuticals, Mahape, Navi Mumbai, India.
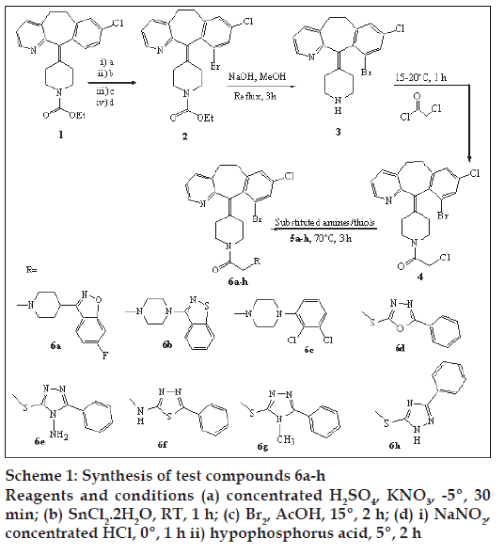
10-Bromodesloratadine (3) was prepared from loratadine (1) by nitration using conc. sulfuric acid and potassium nitrate at -10º for 30 min to get a mixture of two nitro isomers (9-nitro- and 7-nitroloratadine). The nitro group in the mixed isomers was then reduced to amine using stannous chloride dihydrate in ethyl acetate at room temperature. The mixed amines formed were then brominated using bromine in acetic acid at 15-20° to achieve bromination at the C10 position on ring. Diazotization of amine function with sodium nitrite and concentrated HCl at 0° followed by treatment with hypophosphorous acid at 5° gave 10-bromoloratadine (2) as a single isomer. 10-bromoloratadine was decarboethoxylated using sodium hydroxide in methanol at reflux to get 10-bromodesloratadine. Further reaction with chloroacetyl chloride gave an intermediate (4), which was then condensed with various substituted amines or thiols (5a-h) in dimethyl formamide (DMF) in presence of base like potassium carbonate or sodium hydride to get test compounds 6a-h (Scheme 1).
Substituted amines/thiols 5d-5f and 5h were synthesized as per literature methods [6,7], while 4-amino-5-phenyl-3-thiol-1,2,4-triazole (5g) was synthesized from 5-phenyl-1,3,4-oxadiazole-2-thiol by reacting it with 40% methyl amine solution under microwave conditions. Amines 5a-5c were obtained as gift samples from RPG Lifesciences, Pawane, Navi Mumbai, India.
The yield, mp, IR and NMR and mass spectral characteristics of 10-bromoloratadine, 10-bromodesloratadine and test compounds 6a-h are given as follows, 10-bromoloratadine (2), Yield: 65%; mp: 174-176°; IR (KBr): 3059 (C-H stretch, Ar), 2972,1429 (C-H stretch, aliph), 1695 (C=O stretch, ester), 1224 (C-O stretch, acetate), 769 (C-Cl stretch), 525 (C-Br stretch); NMR (CDCl3): 8.47 (s, 1H, Ar), 7.46 (s, 1H, Ar), 7.37 (d, 1H, Ar), 7.20 (s, 1H, Ar), 7.1 (m, 1H, Ar), 4.15 (q, 2H, Aliph), 3.84 (s, 2H, Aliph), 3.14-3.50 (m, 4H, Aliph), 2.8 (m, 2H, Aliph), 2.6 (m, 1H, Aliph), 2.3-2.4 (m, 2H, Aliph), 2.0 (m, 1H, Aliph), 1.25 (t, 3H, Aliph); GC-MS (ES)- 462.5.
10-bromodesloratadine (3), Yield: 65%; mp: 114-116°; IR (KBr): 3508-3385 (N-H stretch, secondary aliphatic amines), 3059 (C-H stretch, Ar), 2928, 1429 (C-H stretch, Aliph), 1103 (C-N stretch, aliphatic amine), 798 (C-Cl stretch), 525 (C-Br stretch); LC-MS (ES)- 388.98, 390.96, 392.95, 393.97.
6a, Yield: 80%; mp: 116-118º; IR (KBr): 3061 (C-H stretch, Ar), 2928,1446 (C-H stretch, Aliph), 1643 (C=O, amide stretch), 1300 (C-N stretch, tertiary amine), 1000 (C-O stretch, ether), 787 (C-Br stretch), 675 (C-Cl stretch); 1H-NMR (CDCl3): 8.5 (s, 1H, Ar), 7.7 (s, 1H, Ar), 7.5 (s, 1H, Ar), 7.37 (d, 1H, Ar), 7.26 (m, 2H, Ar), 7.1 (m, 2H, Ar), 3.9 (s, 2H, Aliph), 3.2-3.6 (m, 6H, Aliph), 3.1 (d, 3H, Aliph), 2.2-2.9 (m, 7H, Aliph), 2.1 (m, 5H, Aliph).
6b, Yield: 70%; mp: 110-112º; IR (KBr): 3059 (C-H stretch, Ar), 2903, 1423 (C-H stretch, Aliph), 1641 (C=O stretch, amide), 1350 (C-N stretch, tertiary amine), 787 (C-Cl), 525 (C-Br stretch); 1H-NMR (CDCl3): 8.5 (s, 1H, Ar), 7.9 (m, 2H, Ar), 7.4 (d, 2H, Ar), 7.36 (m, 2H, Ar), 7.26 (m, 2H, Ar), 3.85 (m, 2H, Aliph), 3.2-3.6 (10H, Aliph), 2.9 (m, 7H, Aliph), 2.4 (m, 2H, Aliph), 2.1 (m, 1H, Aliph); LC-MS (ES)- 647.99, 649.8, 650.04 (MH+).
6c, Yield: 70%; mp: 104-106º; IR (KBr): 3059 (C-H stretch, Ar), 2920, 1448 (C-H stretch, Aliph), 1641 (C=O stretch, amide), 1275 (C-N stretch, tertiary amine), 787 (C-Br stretch), 711 (C-Cl stretch); 1H-NMR (CDCl3): 8.5 (s, 1H, Ar), 7.5 (s, 1H, Ar), 7.43 (d, 1H, Ar), 7.2(m, 5H, Ar), 4.1 (m,1H, Aliph), 3.9(m, 1H, Aliph), 3.4 (m, 6H, Aliph), 3.1 (m, 4H, Aliph), 2.9 (m, 7H, Aliph), 2.5 (m, 2H, Aliph), 2.1 (m, 1H, Aliph).
6d, Yield: 60%; mp: 118-122º; IR (KBr): 3059 (C-H stretch, Ar), 2987, 1467 (C-H stretch, Aliph), 1643 (C=O stretch, amide), 1525(C=N stretch, imines), 1320 (C-N stretch, tertiary amine), 990 (C-O stretch, ethers), 764 (C-Br stretch), 673 (C-Cl stretch); 1H-NMR (CDCl3): 8.5 (d, 1H, Ar), 8.0 (d, 2H, Ar), 7.6 (m, 4H, Ar), 7.4(d, 1H, Ar), 7.35 (s, 1H, Ar), 7.15( m, 1H, Ar), 4.4 (s, 2H, Aliph), 4.10 (m, 1H, Aliph.), 3.8 (m, 1H, Aliph), 3.3-3.8 (m, 4H, Aliph), 2.8 (m, 3H, Aliph), 2.4 (m, 2H, Aliph), 2.1 (m, 1H, Aliph).
6e, Yield: 65%; mp: 154-157º; IR (KBr): 3314-3182 (N-H stretch for amines), 3061 (C-H stretch, aromatic), 2924, 1457 (C-H stretch, Aliph), 1590 (C=N stretch, imines), 1639 (C=O stretch, amide), 1545 (N-H bending vibrations), 771 (C-Cl stretch), 694 (C-Br stretch); 1H-NMR (CDCl3) 8.35 (d, 1H, Ar), 8.15 (d, 2H, Ar), 7.45 (m, 4H, Ar), 7.26 (s, 1H, Ar), 7.20 (m, 2H, Ar), 5.45 (s, 2H, NH2, D2O exchangeable), 4.1(s, 2H, Aliph), 3.8 (m, 2 H, Aliph), 3.30 (m, 4H, Aliph), 2.7 (m, 4H, Aliph), 2.07(m, 2H, Aliph).
6f, Yield: 50%; mp: 170-174º; IR (KBr): 3061 (C-H stretch, aromatic), 2924, 1457 (C-H stretch, Aliph), 1552 (C=N stretch), 1639 (C=O stretch, amide), 771 (C-Cl stretch), 694 (C-Br stretch); 1H-NMR (CDCl3): 8.75 (d, 1H, Ar), 7.63 (d, 2H, Ar), 7.57 (m, 4H, Ar), 7.10(s, 1H, Ar), 7.05 (m, 2H, Ar), 5.9 (1H, N-H) 3.9 (s,2H, Aliph), 3.4 (5 H, Aliph), 2.80 (m, 3H, Aliph), 2.4 (m, 2H, Aliph), 2.1 (m, 2H, Aliph).
6g, Yield: 65%; mp: 157-160°; IR (KBr): 3061 (C-H stretch, aromatic), 2924, 1457 (C-H, Aliph), 1712 (C=N stretch), 1639 (C=O, amide), 771 (C-Cl stretch), 694 (C-Br stretch); 1H-NMR (CDCl3): 8.5 (d, 1H, Ar), 8.15 (d, 2H, Ar), 7.45 (m, 4H, Ar), 7.26 (s, 1H, Ar), 7.20 (m, 2H, Ar), 4.1(s, 2H, Aliph), 3.95 (s, 3H, Aliph), 3.8 (m, 2H, Aliph), 3.30 (m, 4H, Aliph), 2.7 (m, 3H, Aliph), 2.07(m, 2H, Aliph), 2.1 (m, 1H, Aliph).
6h, Yield: 65%; mp: 164-167°; IR (KBr): 3433 (N-H stretch, amines), 3067 (C-H stretch, aromatic), 2926, 1442 (C-H stretch, Aliph), 1545 (C=N stretch), 1631 (C=O stretch, amide), 785 (C-Cl stretch), 694 (C-Br stretch); 1H-NMR (CDCl3): 8.5 (d, 1H, Ar), 8.15 (d, 2H, Ar), 7.45 (m, 4H, Ar), 7.26 (s, 1H, Ar), 7.20 (m, 2H, Ar), 5.32 (s, 1H, N-H, D2O exchangeable), 4.1(s, 2H, Aliph), 3.8 (m, 2H, Aliph), 3.30 (m, 4H, Aliph), 2.7 (m, 3H, Aliph), 2.07(m, 2H, Aliph), 2.1 (m, 1H, Aliph).
Test compounds (6a-h) were evaluated in MCF7 breast, HOP62 lung, MIA-PA-CA-2 pancreatic, Colo205 and HT29 colon cancer cell lines (human) by following the sulforhodamine-B (SRB) semi automated assay protocol using adriamycin as a standard drug [8]. The assay was carried out in a 96 well plate (each well of 100 μl capacity) using 90 μl of tumor suspension containing 5×103 cells. Adriamycin and test compounds (6a-h) were tested at 15, 30, 60 and 120 μM concentrations. All experiments were done in triplicate. Results are average of three experiments. The average value was used to express activity as % inhibition as compared to control and then IC50 value was determined. The results are given in Table 2.
| Compound | IC50 Values in μM concentrations | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Anticancer evaluation | FPT inhibition | ||||||||
| MCF | Hop62 | Colo205 | HT29 | Miapaca-2 | |||||
| 6a | 45 | 120 | 90 | 45 | 33 | 32 | |||
| 6b | 22 | 60 | 90 | <15 | <15 | 34 | |||
| 6c | 41 | >120 | 105 | 45 | 45 | <30 | |||
| 6d | >120 | >120 | >120 | >120 | >120 | 33 | |||
| 6e | 27 | 53 | <15 | <15 | 37 | 40 | |||
| 6f | 110 | >120 | >120 | >120 | >120 | 55 | |||
| 6g | 70 | >120 | >120 | 65 | 60 | 60 | |||
| 6h | 80 | >120 | >120 | 80 | 42 | 32 | |||
| Desloratidine | 25 | 50 | 85 | 20 | 25 | 60 | |||
| 10-Bromodesloratadine | <15 | 27 | 60 | <15 | 18 | 53 | |||
| Adriamycin | <15 | <15 | <15 | <15 | <15 | -- | |||
Table 2: Ic50 values for anticancer evaluation and fpt inhibition
Test compounds 6a-h, desloratadine and 10-bromodesloratadine were evaluated for FPT inhibition by following standard protocol [1] and were tested at 30 and 60 μM concentrations. Desloratadine was used as a standard. Experiments were carried out in duplicate and the average value (% inhibition) was calculated relative to the vehicle control and then IC50 value was determined. Data is given in Table 2.
Test compounds 6a-h in general exhibited dose dependent increase in anticancer activity in the cell lines but were less potent than adriamycin. Amongst the test compounds, 6e was most active with IC50 values of <15 μM in two cell lines tested, while in remaining three cell lines its IC50 values were in the range of 27-53 μM. Adriamycin had IC50 values of < 15 μM concentration. Adriamycin (doxorubicin) does not act by inhibiting FPT, but by interacting with DNA by intercalation [9]. Hence it was not used as a standard in the FPT inhibition study.
Test compounds 6a-h exhibited potent FPT inhibitory activity at the doses studied and had IC50 values in the range of 32-40 μM except for 6f which had IC50 of 55 μM. In comparison, des loratadine had IC50 of 60 μM and 10-bromodesloratadine had IC50 of 53 μM. 10-bromodesloratadine was evaluated to examine the effect of introduction of bromine in desloratadine. The results show that 10-bromodesloratadine was more potent compared to desloratadine indicating that activity increases with bromination at C10 position.
It is clear that test compounds 6a-h in general are more potent than 10-bromodesloratadine and this enhanced activity can be attributed to the presence of zinc binding groups in these molecules.
Acknowledgements
The authors wish to thank the All India Council of Technical Education (AICTE) for providing funds for this project.
References
- Mallams AK, Njoroge FG, Doll RJ, Snow ME, Kaminski JJ, Rossman RR, et al. Antitumor 8-chlorobenzocycloheptapyridines: A new Class of Selective, Nonpeptidic, Nonsulfhydryl Inhibitors of RasFarnesylation. Bioorg Med Chem 1997;5:93-9.
- N joroge FG, Taveras AG, Kelly J, Remiszewski S, Mallams AK, Wolin R, et al. (+)-4-[2-[4-(8-Chloro-3,10-dibromo-6,11-dihydro-5H-benzo[5,6]cyclohepta[1,2-b]-pyridin-11(R)-yl)-1-piperidinyl]-2-oxo-ethyl]-1-piperidinecarboxamide (SCH-66336): A Very Potent Farnesyl Protein Transferase inhibitor as a Novel Antitumour agent. J Med Chem 1998;41:4890-902.
- End DW, Smets G, Todd AV, Applegate TL, Fuery CJ, Angibaud P, etal. Characterization of the Antitumor Effects of the Selective FarnesylProtein Transferase Inhibitor R115777 in vivo and in vitro. Cancer Res 2001;61:131-7.
- Hunt JT, Ding CZ, Batorsky R, Bednarz M, Bhide R, Cho Y, etal. Discovery of (R)-7-Cyano-2,3,4,5-tetrahydro-1-(1H-imidazol-4-ylmethyl)-3-(phenylmethyl)-4-(2-thienylsulfonyl)-1H-1,4-benzodiazepine (BMS-214662), a Farnesyltransferase Inhibitor with potent Preclinical Antitumor Activity. J Med Chem 2000;43:3587-95.
- Njoroge FG, Vibulbhan B, Pinto P, Strickland C, Kirschmeier P, Bishop WR, et al.Farnesyl Protein Transferase inhibitors targeting the catalytic zinc for enhanced activity. Bioorg Med ChemLett 2004;14:5877-80.
- Boyer JH, Elderfield RC. Heterocyclic compounds. New York: John Wiley and Sons; 1961. p. 525.
- Young RW, Wood KH, Eichler JA, Vaughan JR Jr., Anderson GW. 1,3,4-Thiadiazole- and Thiadiazolinesulfonamides as Carbonic Anhydrase Inhibitors. Synthesis and Structural Studies. J Am ChemSoc 1956;78:46-9.
- Smith V, Rowlands MG, Barrie E, Workman P, Kelland LR. Establishment and Characterization of Acquired Resistance to the Farnesyl Protein Transferase Inhibitor R115777 in a Human Colon Cancer Cell Line. Clin Cancer Res 2002;8:2002-09.
- Fornari FA, Randolph JK, Yalowich JC, Ritke MK, Gewirtz DA. Interference by Doxorubicin with DNA Unwinding in MCF-7 Breast Tumor Cells. MolPharmacol 1994;45:649-56.