- *Corresponding Author:
- A. Kumar
Centre for Bioinformatics, M. D. University, Rohtak‑124 001, India
E‑mail: akumar.cbt.mdu@gmail.com
| Date of Submission | 26 February 2012 |
| Date of Revision | 03 January 2013 |
| Date of Acceptance | 07 January 2013 |
| Indian J Pharm Sci 2013;75(1):23-30 |
Abstract
The structure-function correlation of membrane proteins have been a difficult task, particularly in context to transient protein complexes. The molecular simulation of ternary complex of Rab7::REP1::GGTase-II was carried out to understand the basic structural events occurring during the prenylation event of Rab proteins, using the software YASARA. The study suggested that the C-terminus of Rab7 has to be in completely extended conformation during prenylation to reach the active site of RabGGTase-II. Also, attempt was made to find putative drug binding sites on the ternary complex of Rab7::REP1::GGTase-II using Q-SiteFinder programme. The comprehensive consensus probe generated by the program revealed a total of 10 major pockets as putative drug binding sites on Rab7::REP:: GGTase-II ternary complex. These pockets were found on REP protein and GGTase protein subunits. The Rab7 was found to be devoid of any putative drug binding sites in the ternary complex. The phylogenetic analysis of 60 Rab proteins of human was carried out using PHYLIP and study indicated the close phylogenetic relationship between Rab7 and Rab9 proteins of human and hence with further in silico study, the present observations can be extrapolated to Rab9 proteins. The study paves a good platform for further experimental verifications of the findings and other in silico studies like identifying the potential drug targets by searching the putative drug binding sites, generating pharmacophoric pattern, searching or constructing suitable ligand and docking studies.
Keywords
AMBER, GGTase‑II, molecular simulation, prenylation, Rab7, REP, YASARA
Vesicular trafficking plays a central role in the formation and establishment of different compartments as well as in the communication between the cells with each other and with the environment [1]. Rab proteins are the largest subfamily of Ras-related GTPase super family that regulates a large number of processes like responses to external stimuli, cytoskeletal remodelling and intracellular transport. Rab proteins are localised in membrane by the virtue of a well‑recognised and versatile posttranslational event of protein prenylation. The prenylation event of protein encompasses enzymatic conjugation of either a farnesyl chain (15‑carbon long) or a geranylgeranyl (GG) chain (20‑carbon long) with the C‑terminal cysteine residue of the protein. RAb geranyl geranyl transeferase II (RabGGTase‑II) is one of the three protein prenyltransferase in eukaryotic system and is unique in having strict substrate preference, acting only on those members of the RabGTPase family which play a central role in membrane trafficking [2]. RabGGTase‑II (a heterodimeric protein with α‑ and β‑subunits), for its catalytic action, requires integration to the catalytic core of RabGTPases along with an additional unique factor called Rab escort protein (REP) [3]. REP carries a nascent RabGTPase and presents it to RabGGTase for prenylation, by binding to its α‑subunit. After the prenylation event, the Rab:REP complex moves to the destined organelle’s membrane and the Rab is stably associated with the membrane due to the insertion of GG moieties into lipid bilayers [4,5].
More than 60 human Rab proteins have been identified and have been reported to perform significant role in the regulation of vesicle trafficking processes like vesicle formation, motility, tethering and fusion to the acceptor membrane and signalling to other organelles of cellular regions along with cell growth, survival and development [6‑8]. Rab proteins and their associated regulators are reportedly involved in many diseases, such as cancer, pigmentation disorder, neuropathy, lipid metabolism disorders and many other pathogenic diseases [9‑12]. It has been established that many drug targets are localised to particular subcellular compartments, but still many of the drug design strategies are focussed on tissue targeting. Insights into how the cell traffics its constituents to different intracellular compartments could improve drug design [13]. Therefore, it has been the need of the hour to understand the complete biomolecular events associated with Rab proteins and its associated effector and affector molecules, Rab::REP::GGTase ternary complex being one of them.
Structural biology investigation of the Rab::REP:: GGTase ternary complex has been futile because of limitations of nuclear mangentic resonance, X‑ray crystallography and electron microscopy in the field of transient membrane protein complexes study. Also, it is hard to reproduce the membrane molecular events by these methods [14]. Furthermore, the experimental structural view of a molecular event is often constrained by the lack of understanding the time dependent mechanism bridging structure and the execution of the biological function. Understanding protein motion is essential for deciphering functional mechanisms including folding [15,16] along with the complementary unfolding [17,18], allosteric regulation [19], catalysis [20], functional plasticity [21], protein–DNA interactions [22] and protein–protein interactions [23].
The information from the different static structures can be suitably analysed to yield a dynamic path between structures [24], however, simulation of static structure of complex in a time‑dependent manner is generally required to have dynamic insight of the structural framework. Two crystal structures of REP in complex with either Rab7 and RabGGTase‑II have been described earlier [4,25]. The interface between Rab7 and REP has been analysed individually [26], while similar analysis for REP–GGTase‑II complex has also been done earlier [4]. However, a comprehensive structural dynamic analysis of the ternary complex of Rab7::REP1::GGTase‑II is scanty [14]. So, the present in silico study was carried out for generating a computational model of Rab7::REP1::GGTase‑II ternary complex and understand the molecular dynamics of the complex by simulation up to 0.5 ns, and identifying putative drug binding sites (DBSs) on the ternary complex. A phylogenetic study of different human Rab proteins was also carried out to extend the findings of present investigation to Rab proteins other than Rab7.
Materials and Methods
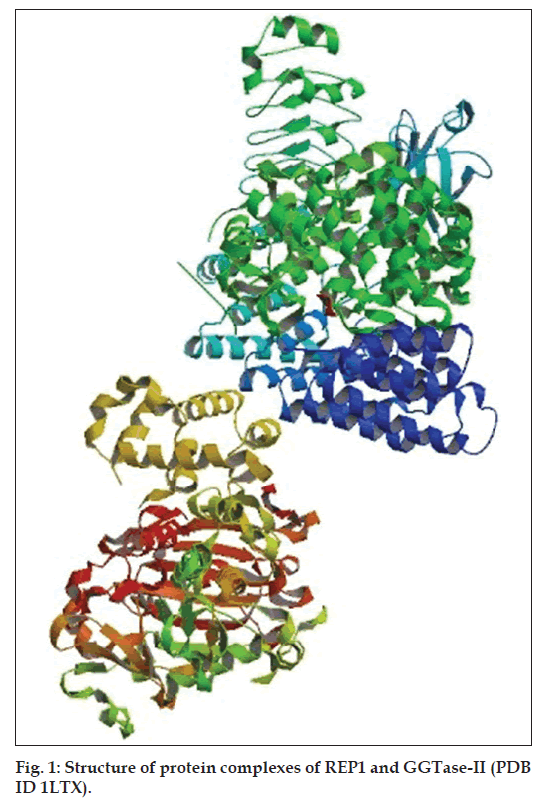
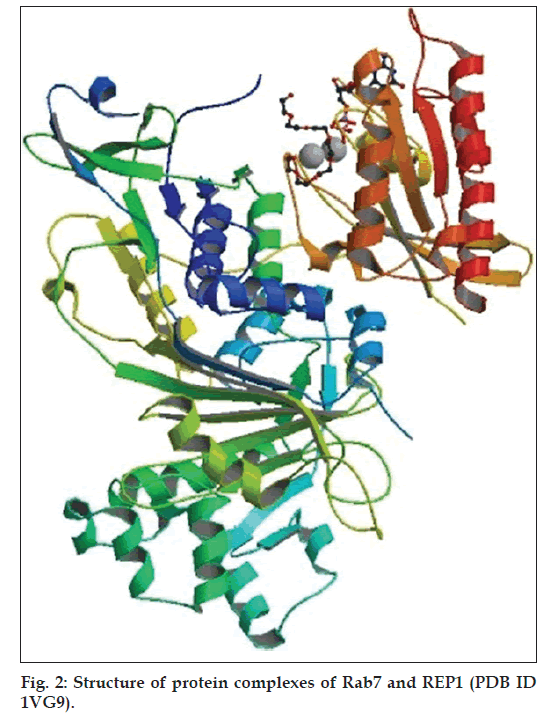
The protein complex models of REP1:GGTase‑II (Protein Data Bank [PDB] ID 1LTX) (fig. 1) and that of Rab7:REP1 (PDB ID 1VG9) (fig. 2) were obtained from PDB (http://www. rcsb.org/pdb) for the present study. The software Yet Another Scientific Artificial Reality Application (YASARA) Dynamics and Structure (YASARA Biosciences GmBH, Vienna, Austria) (v 10.12.1) was used for generating the molecular model of ternary complex of Rab7::REP1::GGTase‑II and also for its molecular dynamic simulation study. The online Q‑SiteFinder programme (University of Leeds, Leeds, UK) [27] was used for identification of putative DBSs over the ternary complex and PyMOL (Schrödinger, USA) [28] (an open source software) was used for the molecular visualisation. For phylogenetic analysis, 60 human Rab proteins were obtained from UniProt (http://www.uniprot.org) and the softwares used for the study were ClustalW and PHYLIP (University of Washington, Seattle, USA).
Construction of model of ternary complex of Rab7::REP1::GGTase‑II:
The structure files of protein complex models of REP1::GGTase‑II (PDB ID 1LTX) (fig. 1) and that of Rab7::REP1 (PDB ID 1VG9) (fig. 2) were downloaded from PDB and the two complexes were joined using YASARA by selecting the strands of REP1 molecule in both the complexes, a single complex was generated. Since the REP1 molecule was repeated, one of the REP1 molecules in the combining complexes was deleted by individually picking up the peptide chain. The ternary complex of Rab7::REP1::GGTase‑II thus obtained was subjected to solvation option of YASARA to add water molecules. To study the solvation effects on side chains of the protein complex, electrostatic interactions were screened. A null model pKa giving the lowest rootmean- square deviation (RMSD) without any shift in value was taken, where pKa values were set as 3.22, 4.09, 6.20, 10.8 and 10.76 for Asp, Glu, His, Tyr and Lys residues, respectively.
Simulation of ternary complex of Rab7::REP1::GGTase‑II
The constructed and solvated system of ternary complex of Rab7::GGTase‑II::REP was subjected to energy minimization without any constraints using steepest descent method followed by simulated annealing method. pH of the system was set to 3.4 (acidic) for better results. Simulation temperature and pressure were set to 323 ºK and 1 atm, respectively. Particle mesh Ewald method was employed to calculate the electrostatic interactions with a cut‑off of 10 Ǻ. Simulation snapshots were taken every 10,000 simulation steps, i.e., with a timestep of 2.5 fs, i.e., 10,000*2.5=25 ps. For these activities YASARA simulation software was used.
The constructed ternary complex of Rab7::GGTase‑II::REP was initially subjected to energy minimisation by AMBER force field, using Dynamics or Structure module of YASARA. It was done by loading the ternary complex structure into the module. Then, after clicking onto Simulation → force field, YAMBER99 force field was selected in YASARA Dynamics. With the selection being made, simulation was run with command – Options → Macro and Movie → Play macro and double‑clicking the standard macro ‘em_ run.mcr’.
The molecular dynamics simulation was then carried out in YASARA Dynamics or Structure, where a project directory was created and the structure of ternary complex of Rab7::GGTase‑II::REP was stored after energy minimisation. The molecular dynamics simulation was carried out after selecting the saved target molecule and following the command – Options → Macro and Movie → Play macro and double‑click the standard macro ‘md_run.mcr’. The simulation performance was increased (when needed) by increasing the simulation steps per screen update using ‘simulation>timestep’ module.
Analysis of simulation trajectory of ternary complex of Rab7::GGTase‑II::REP:
The data obtained after simulations were analysed for trajectory projection. This was done by using 'Macro and Movie' option and playing the standard analysis macro. This macro created a self‑explanatory table and was saved in project directory with ‘.tab’ ‑ file extension. The table was then imported in the data visualisation program (MS‑Excel) that included energies and RMSDs from the starting structure during simulation. The standard analysis macro also calculated the time average structure, whose atoms had the B‑factors (calculated from the root mean square fluctuations) during simulation and the minimum energy structure was saved as ‘energymin. sce’ in the project directory.
Identification of putative DBS of Rab7::REP1::GGTase‑II:
To find the putative DBSs of Rab::REP::GGTase‑II complex, Q‑SiteFinder programme was used in the present study. The Q‑SiteFinder web site (www. modelling.leeds.ac.uk/qsitefinder) was visited and the PDB file of the ternary complex was uploaded and job was submitted. The Q‑SiteFinder programme uses several separate procedures to perform ligand binding site prediction such as LigandSeek, Liggrid and Probe‑Clustering programme. The probe clusters were ranked according to their total interaction energies, with the most favourable being identified as the first predicted binding site. The clustering programme also calculated site volume, and identified the protein atoms that were within a defined range of cluster sites. The programme generated the 10 best predicted sites (interconnected dots) and any ligands (coloured by atoms) were displayed as result. Predicted binding site were listed in the display sites (atoms forming the binding sites on the protein) window according to the likelihood (energy) of the cluster of probes (Site 1, Site 2). Q‑SiteFinder results were downloaded and viewed in PyMOL.
Phylogenetic analysis of different Rab proteins found in humans
All together 60 different human Rab proteins, relevant to the present study, were analysed for evolutionary relationship by multiple sequence alignment (MSA) of their amino acid sequences. The amino‑acid sequences of these proteins were obtained from UniProt (www.uniprot.org). These sequences were then arranged in FASTA format for MSA by ClustalW. The output file (.phy) was imported to the PHYLIP for constructing the phylogenetic tree using neighbour‑joining method.
Results and Discussion
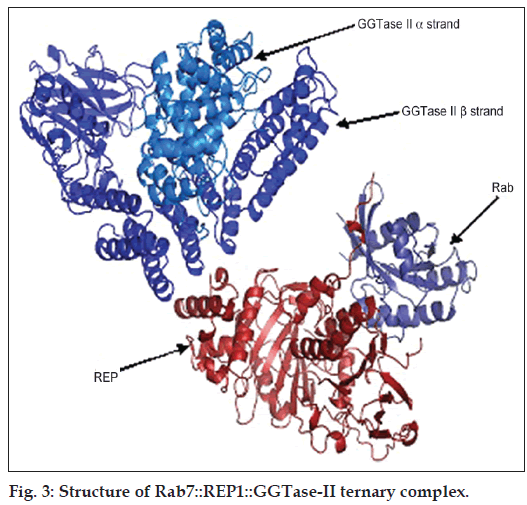
The molecular model of the ternary complex of Rab7::REP1::GGTase‑II (fig. 3) was obtained using YASARA edit option and was subjected to solvation. While studying the solvation effects on the side chains of protein complex, it was observed that the predicted pKa range of Lys, Asp, Glu, Tyr and His were 10.46‑11.84, 2.82‑5.09, 3.74‑5.08, 10.8 and 6.34‑7.42, respectively. The results thus obtained were in good accordance of assumed null model and was thus subjected to further energy minimisation programme and force field calculations
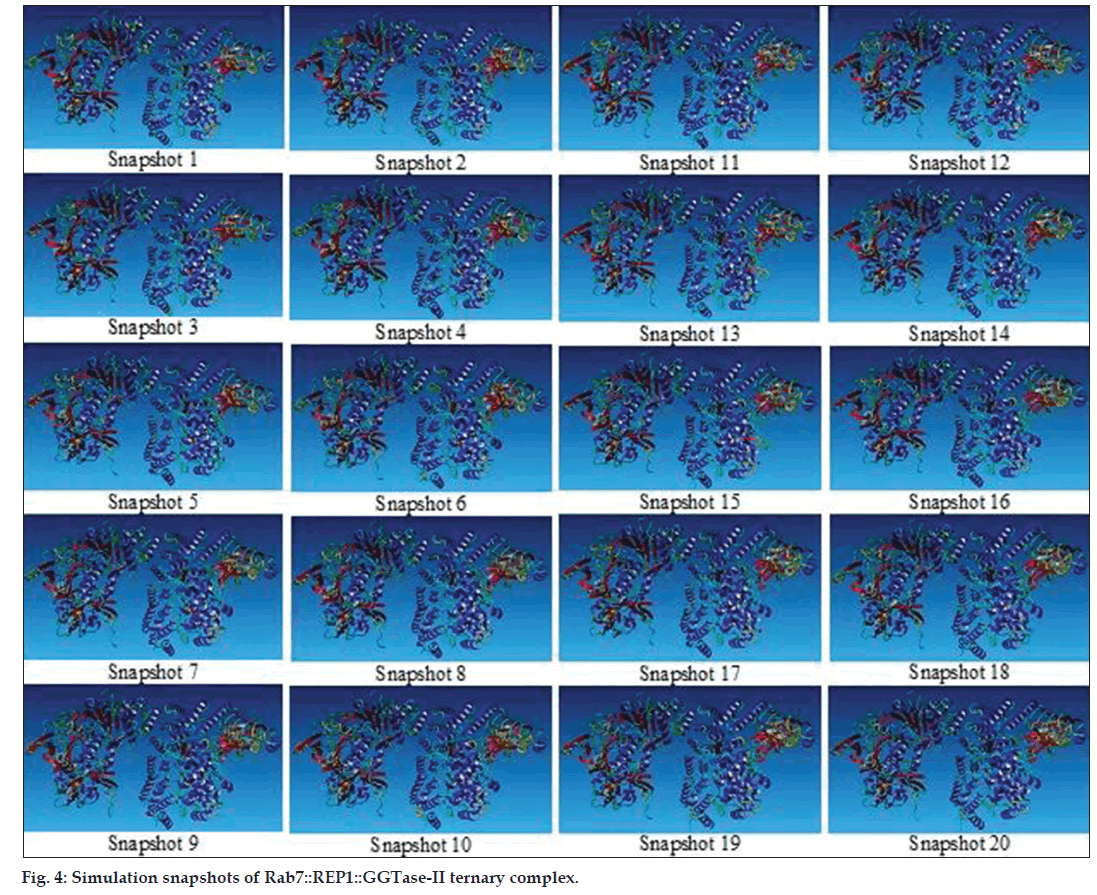
After running the energy minimisation program, the ternary complex was subjected to simulation up to 500 ps (0.5 ns). The different snapshots of simulating steps were saved with respect to set time‑intervals (fig. 4). The movie, thus, generated from these snapshots, reveals a very clear image of Rab7, GGTase‑II and REP1 interaction.
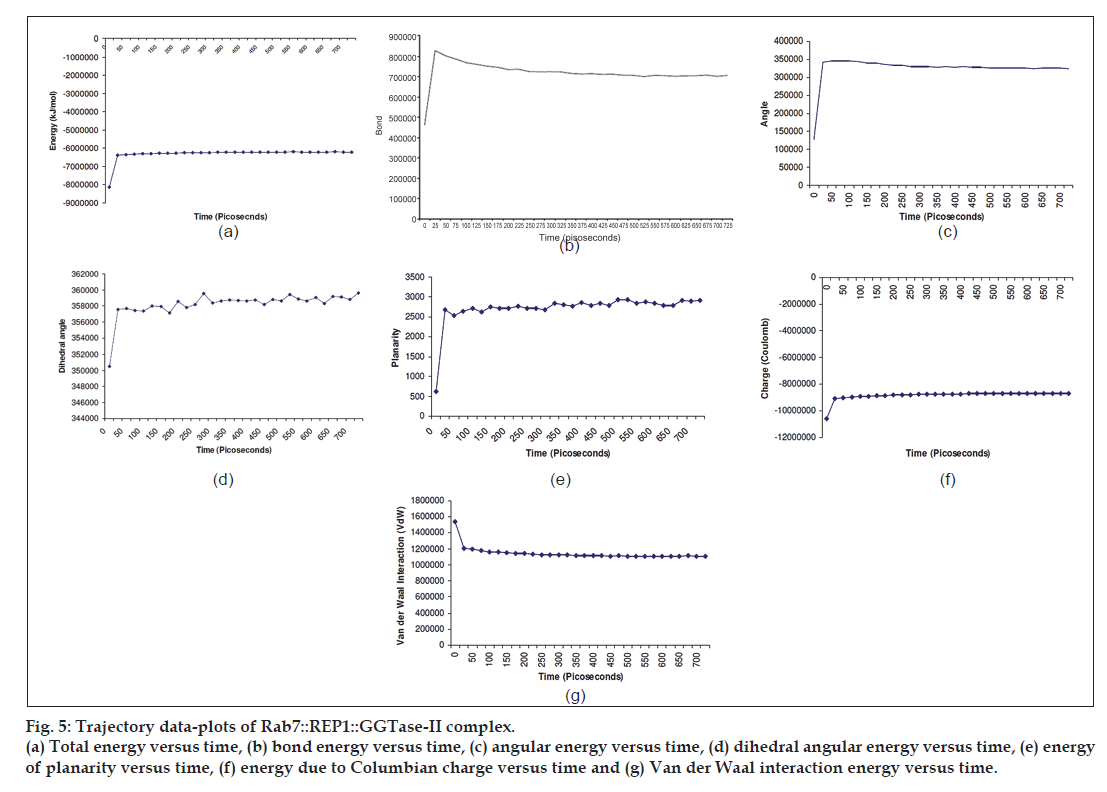
The trajectory was obtained for overall energy simulation of the ternary complex of Rab7:GGTase‑II: REP for 500 ps and it revealed that overall energy stabilised after a peak of ‑ 6363401.4 kJ/mol at 25 ps and tended to remain in plateau phase further for rest of the period (fig. 5a). This reflected that simulation was achieved with stable energy for rest of the period (50‑700 ps) for the said ternary complex. Almost, the similar trajectory was obtained for the plots of different energy contributors against simulation run time. The contribution due to steric parameters like bond, angle, dihedral angle and planarity was found maximum at 25 ps with the values 828405, 341330.665, 357593.243 and 2679.375 kJ/mol, respectively, which stabilised further to a stationary phase for rest of the period (50‑700 ps), similar to the trajectory pattern of overall energy curve (fig. 5b‑e). The contribution due to Columbian charge and van der Waal interactions was also in accordance to the above results and was maximum at 25 ps with the values of ‑ 9042365.5 and 1193953.512 kJ/mol (fig. 5f and g). The plots tended to stabilise further and attain the plateau phase for rest of the period (50‑700 ps). These trajectory patterns supports and validates the simulation profile of the ternary complex of Rab7::REP1::GGTase‑II.
The simulation model investigation of the ternary complex of Rab7::REP1::GGTase‑II suggests that the C‑terminus of Rab7 has to be in completely extended conformation during prenylation to reach the active site of RabGGTase‑II. The results obtained in the present study are in good accordance with previous investigations [14] and has provided a good picture of molecular dynamics of ternary complex of Rab7::REP1::GGTase‑II and the molecular events during interaction of these three proteins.
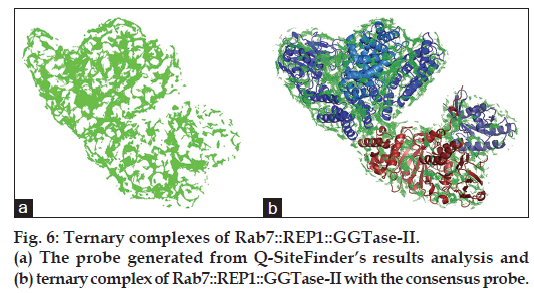
All together of about 60‑70 putative DBSs were generated by Q‑SiteFinder programme. To predict the DBS precisely, these sites were comparatively analysed using an inhouse Perl algorithm. The comparative analysis resulted in construction of comprehensive probe based on clustering method of energy criteria. The comprehensive probe thus generated (fig. 6a) was used to overlap with the ternary complex of Rab7::REP1::GGTase‑II. The probe was found to conjugate fit with the Rab7::REP1:: GGTase‑II ternary complex (fig. 6b) and thus was a good primary indication of the correctness of the generated comprehensive probe.
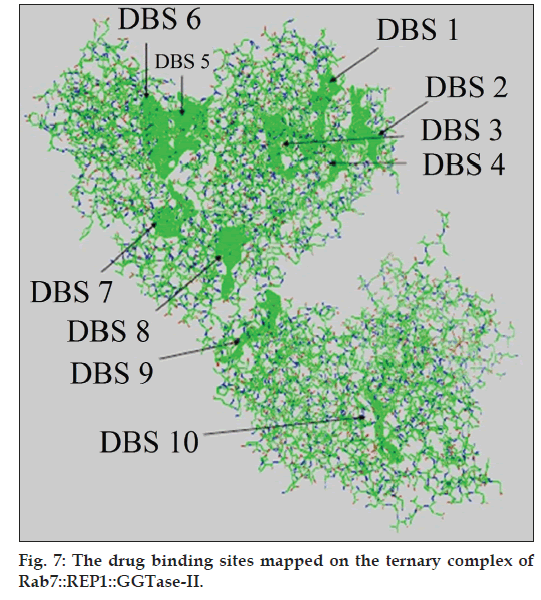
The comprehensive probe was used to integrate the putative DBSs being generated by different Q‑SiteFinder programme, as mentioned earlier. After an iterative process of selection and omission of different predicted DBSs based on statistical parameters, a total of 10 major pockets were identified as putative DBSs on Rab7::REP1::GGTase‑II complex (fig. 7). These pockets reflected the energy of interactions of different probes being generated by the different used algorithms, viz.; carbohydrate binding propensity, drug‑like compound binding propensity, interacting pocket size threshold of radius 3 Å in three‑dimension.
These 10 major putative DBSs thus identified were then mapped onto the Rab7::REP1::GGTase‑II ternary complex (fig. 7) which revealed that a total of two major pockets of DBS (one in the β‑barrel and one in α‑helical portion) were located on the REP protein of Rab7::REP1::GGTase‑II ternary complex. Of these two DBS, one was found to be located within the major cavity produced at the interaction point of Rab7‑REP complex with GGTase‑II. The other eight major putative DBSs identified were found to be located on GGTase portion of the ternary complex. The α‑strand of GGTase was found to have two major pockets of putative DBSs while the β‑strand possessed the other six pockets of putative DBSs. All these putative DBS mapped completely over the Rab7::REP1::GGTase‑II ternary complex, which was an indirect evidence of completeness of the computational calculation. These results have paved the path for further investigation into assigning rigidity to these putative DBSs and develop pharmacophoric patterns for the same. These pharmacophore can be further used for virtual screening of ligand databases (viz., PubChem, ZINC) and are being presently investigated in our lab, apart from generating combinatorial libraries.
After performing MSA of the selected 60 different human Rab proteins and constructing the phylogenetic tree by neighbour‑joining algorithm of PHYLIP of the same, it was observed that human Rab7a, Rab7b, Rab9a and Rab9b share the common ancestor and are the nearest neighbours during the course of evolution. So, the molecular dynamics studies of Rab7::GGTase‑II::REP ternary complex will also provide a good platform for further investigation of related human Rab proteins such as Rab7a, Rab7b, Rab9a and Rab9b. Rab7 and Rab9 are related to each other being localised to late endosomes and thus the phylogenetic analysis in the present study is coherent with the earlier results [29‑31].
To conclude, we can say that the present study has produced a good insight into the molecular events occurring during the ternary complex formation of Rab7::REP1::GGTase‑II. The study also draws its significance because the protein studied is a membrane protein whose structural analysis is difficult through experimental way. The present in silico study was a primary and initial attempt in this direction and has paved pathway for further in silico studies like identifying the potential drug targets by searching the putative DBSs, generating pharmacophoric pattern, searching or constructing suitable ligand, docking studies.
References
- Huotari J, Helenius A. Endosome maturation. EMBO J 2011;30:3481‑500.
- Stenmark H, Olkkonen VM. The Rab GTPase family. Genome Biol 2001;2:3007.
- Wu YW, Tan KT, Waldmann H, Goody RS, Alexandrov K. Interaction analysis of prenylated Rab GTPase with Rab escort protein and GDP dissociation inhibitor explains the need for both regulators. Proc Natl Acad Sci USA 2007;104:12294‑9.
- Pylypenko O, Rak A, Reents R, Niculae A, Sidorovitch V, Cioaca MD, et al. Structure of Rab escort protein‑1 in complex with Rab geranylgeranyltransferase. Mol Cell 2003;11:483‑94.
- Ghomashchi F, Zhang X, Liu L, Gelb MH. Binding of prenylated and polybasic peptides to membranes: Affinities and intervesicle exchange. Biochemistry 1995;34:11910‑8.
- Pfeffer SR. Unsolved mysteries in membrane traffic. Annu Rev Biochem 2007;76:629‑45.
- Somsel Rodman J, Wandinger‑Ness A. Rab GTPases coordinate endocytosis. J Cell Sci 2000;113:183‑92.
- Snider MD. A role for Rab7 GTPase in growth factor‑regulated cell nutrition and apoptosis. Mol Cell 2003;12:796‑7.
- Seabra MC, Mules EH, Hume AN. RabGTPases, intracellular traffic and disease. Trends Mol Med 2002;8:23‑30.
- Brumell JH, Scidmore MA. Manipulation of rab GTPase function by intracellular bacterial pathogens. Microbiol Mol Biol Rev 2007;71:636‑52.
- Corbeel L, Freson K. Rab proteins and Rab‑associated proteins: Major actors in the mechanism of protein‑trafficking disorders. Eur J Pediatr 2008;167:723‑9.
- Zhang M, Chen L, Wang S, Wang T. Rab7: Roles in membrane trafficking and disease. Biosci Rep 2009;29:193‑209.
- Rajendran L, Knölker HJ, Simons K. Subcellular targeting strategies for drug design and delivery. Nat Rev Drug Discov 2010;9:29‑42.
- Wu YW, Goody RS, Abagyan R, Alexandrov K. Structure of the disordered C terminus of Rab7 GTPase induced by binding to the Rab geranylgeranyl transferase catalytic complex reveals the mechanism of Rab prenylation. J Biol Chem 2009;284:13185‑92.
- Daggett V, Fersht A. The present view of the mechanism of protein folding. Nat Rev Mol Cell Biol 2003;4:497‑502.
- Muñoz V. Downhill protein folding under pressure. Nat Methods 2009;6:490‑1.
- Dobson A. Ecology metalife! Science 2003;301:1525‑6.
- Daggett V. Protein folding‑simulation. Chem Rev 2006;106:1898‑916.
- Popovych OV, Hauptmann C, Tass PA. Control of neuronal synchrony by nonlinear delayed feedback. Biol Cybern 2006;95:69‑85.
- Benkovic SJ, Hammes‑Schiffer S. Biochemistry. Enzyme motions inside and out. Science 2006;312:208‑9.
- Kobilka BK, Deupi X. Conformational complexity of G‑protein‑coupled receptors. Trends Pharmacol Sci 2007;28:397‑406.
- Sarai A, Siebers J, Selvaraj S, Gromiha MM, Kono H. Integration of bioinformatics and computational biology to understand protein‑DNA recognition mechanism. J Bioinform Comput Biol 2005;3:169‑83.
- Bonvin AM. Flexible protein‑protein docking. Curr Opin Struct Biol 2006;16:194‑200.
- Flores MV, Lam EY, Crosier P, Crosier K. A hierarchy of Runx transcription factors modulate the onset of Chondrogenesis in craniofacial endochondral bones in zebrafish. Dev Dyn 2006;235:3166‑76.
- Rak A, Pylypenko O, Niculae A, Pyatkov K, Goody RS, Alexandrov K. Structure of the Rab7:REP‑1 complex: Insights into the mechanism of Rab prenylation and choroideremia disease. Cell 2004;117:749‑60.
- Guo Z, Wu YW, Das D, Delon C, Cramer J, Yu S, et al. Structures of RabGGTase‑substrate/product complexes provide insights into the evolution of protein prenylation. EMBO J 2008;27:2444‑56.
- Laurie AT, Jackson RM. Q‑SiteFinder: An energy‑based method for the prediction of protein‑ligand binding sites. Bioinformatics 2005;21:1908‑16.
- DeLano WL. The PyMOL Molecular Graphics System. San Carlos, CA, USA: DeLano Scientific; 2002. Available from: http://www.pymol.org [Last accessed on 2012 Dec 6].
- Soldati T, Rancaño C, Geissler H, Pfeffer SR. Rab7 and Rab9 are recruited onto late endosomes by biochemically distinguishable processes. J Biol Chem 1995;270:25541‑8.
- Chavrier P, Simons K, Zerial M. The complexity of the Rab and Rho GTP‑binding protein subfamilies revealed by a PCR cloning approach. Gene 1992;112:261‑4.
- Brighouse A, Dacks JB, Field MC. Rab protein evolution and the history of the eukaryotic endomembrane system. Cell Mol Life Sci 2010;67:3449‑65.