- *Corresponding Author:
- S. B. Kalidhar
Department of Chemistry, Haryana Agricultural University, Hisar-125 004, India
E-mail: kalidhar@hau.ernet.in
| Date of Submission | 25 June 2007 |
| Date of Decision | 22 January 2008 |
| Date of Acceptance | 14 August 2008 |
| Indian J Pharm Sci, 2008, 70 (4): 517-519 |
Abstract
Phytochemical examination of the peel of grapefruit resulted in the isolation of five compounds namely friedelin, β -sitosterol, 7(3',7',11',14'-tetramethy)pentadec-2',6',10'-trienyloxycoumarin, limonin and cordialin B. These compounds have been characterized on the basis of spectral data, and 7(3',7',11',14'-tetramethy)pentadec-2',6',10'-trienyloxycoumarin is a hitherto unreported compound.
Keywords
Citrus paradisi, Rutaceae, coumarin, friedelin, β-sitosterol, limonin, cordialin B
Citrus paradisi commonly known as grapefruit and pahari nimbu belongs to the family Rutaceae. It is synonymous with C. decumana var. racemosa, C. decumana var. paradisi Nichols and C. racemosa Macf. The juice of the grapefruit is used for building up resistance to common cold and wound infections [1]. There is report of only one compound i.e. 6,7- dimethoxycoumarin [2] from grapefruit peel. In view of existing scanty data, the present study was undertaken (fig 1).
Melting points were determined on Ganson Electrical Melting Point Apparatus. IR spectra were recorded on Hitachi 570 Infrared Spectrophotometer using KBr. 1H NMR spectra were recorded on Brucker AC-300F 300 MHz NMR Spectrophotometer using TMS as an internal standard. Chemical shifts are given in δ (ppm) and CDCl3 was used as a solvent for recording NMR spectra. Fruits of C. paradisi (10 kg) were procured from Landscape, HAU, Hisar.
The peel of the fruits was removed and air dried. The peel was then refluxed with hot methanol for 6 h and the process was repeated 4 times. Extractives were concentrated on water bath under reduced pressure and the viscous mass thus obtained was mixed with silica gel (60-120 mesh), dried on water bath and subjected to column chromatography. Petroleum ether was used as a solvent in packing the column. Five compounds were isolated.
Compound A crystallized from methanol, 25 mg, mp 2680 (literature mp 267.3-269.5º) [3]. It gave pale brown color on reaction with Ac2O/H2SO4; IRνmax (KBr, cm-1): 1720; 1H NMR (δ, CDCl3): 2.25-1.20 (25 H, m, 11*CH2 and 3*CH), 1.17 (3 H, s, CH3), 1.04 (3H, s, CH3), 1.00 (6H, s, 2*CH3), 0.95 (3 H, s, CH3), 0.86 (6H, s, 2*CH3), 0.72(3H, s, CH3); GCMS: 426 (M+). This data settled compound A to be friedelin (1)3.
Compound B crystallized from methanol, 20 mg, mp 1360 (literature mp 136-137º) [4]. It responded to Liebermann-Burchard Reaction. IRνmax (KBr, cm-1): 3427; 1H NMR (δ, CDCl3): 5.34 (1H, br, H-6), 3.51 (1H, m, H-3), 2.28-1.13 (29H, m, 11*CH2, 7*CH), 0.92 (6H, s, 2*CH3), 0.83 (3H, s, CH3), 0.80 (3H, s, CH3), 0.78 (3H, s, CH3), 0.68 (3H, s, CH3); GCMS: 414 (M+). This data confirmed compound B to be β-sitosterol (2) [5] using a direct comparison.
Compound C (3) crystallized from methanol, 50 mg, mp 70º. IRνmax (KBr, cm-1): 1700; 1H NMR (δ, CDCl3): 7.64 (1H, d, J 10 Hz, H-4), 7.36 (1H, d, J 7.5 Hz, H-5), 6.82 (2H, m, H-6, H-8), 6.25 (1H, d, J 10 Hz, H-3), 5.50 (1H, 1H, t, J 7.0 Hz, H-2’), 5.10 (2H, t, J 7.0 Hz, H-6’, H-10’), 4.61 (2H, d, J 7.0 Hz, 2*H-1’), 2.34 (4H, m, 2*H-5’, 2*H-9’), 2.10 (6H, t, J 7.0 Hz, 2*H-4’, 2*H-8’, 2*H-12’), 1.75 (3H, s, CH3), 1.72 (3H, s, CH3), 1.65 (3H, s, CH3), 1.30-0.86 (9H, m, 2*CH3, CH2, CH); GCMS: 422 (M+).
Compound D crystallized from methanol, 40 mg, mp 297 (literature mp 298) [3]. IRνmax (KBr, cm-1): 1758,1708; 1H NMR (δ, CDCl3): 7.41 (2H, m, H-2’, H-5’), 6.30 (1H, m, H-3’), 5.46 (1H, s, OCHCOO), 4.77-4.47 (2H, m, 2*OCH), 4.02 (2H, s, OCH2), 2.98-1.77 (10H, m, 4*CH2, 2*CH), 1.29 (3H,s, CH3), 1.17 (6H, s, 2*CH3), 1.07 (3H,s, H3); GCMS: 470 (M+). The data proposed compound D to be limonin (4)3.
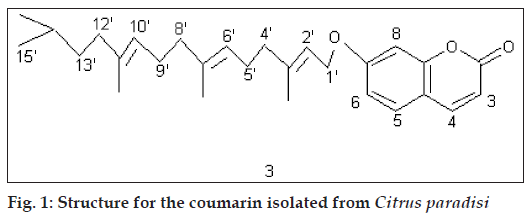
Compound E crystallized from methanol, 50 mg, mp 1140 (literature mp 114-1150) [3]. It responded to Liebermann-Burchard Reaction. IRνmax (KBr, cm-1): 3358; Acetate: compd.+Ac2O+Py; 1H NMR (δ, CDCl3): 5.34 (1H, m, =CH), 4.28 (1H, m, CH-O), Fig. 1: Structure for the coumarin isolated from Citrus paradisi 4.10 (1H, m, CH-O), 3.65(1H, m, CH-O), 3.39(1H, m, CH-O), 2.32-1.08 (20H, m, 8*CH2, 4*CH), 2.08 (3H, s, OAc), 2.04 (3H, s, OAc), 2.01 (3H, s, OAc), 2.00 (3H, s, OAc), 1.23 (3H, s, CH3), 1.10 (3H, s, CH3), 0.98 (3H, s, CH3), 0.92 (3H, s, CH3), 0.88 (3H, s, CH3), 0.86 (3H, s, CH3), 0.67 (3H, s, CH3), GCMS: 490 (M+). A comparison with literature data [3] suggested this compound to be cordialin B (5).
Compound C showed fluorescence under UV indicating it to be a coumarin. The IR spectrum of the compound exhibited the presence of a carbonyl group at 1700 cm-1 which was a further support towards the coumarin nucleus. MS suggested its molecular mass to be 422. 1H NMR of the compound in CDCl3 showed a doublet at δ 7.64 (J 10 Hz) which was typical of H-4 of a coumarin. Another doublet was observed at δ 7.36 (J 7.5 Hz) which could be H-5 of a coumarin. There was a multiplet at δ 6.82 for two protons which represented H-6 and H-8 of the nucleus. A doublet at δ 6.25 (J 10 Hz) was assignable to H-3. A triplet at δ 5.50 (J 7.0 Hz) could be H-2’ of an aliphatic chain attached at C-7 of the nucleus. Another triplet at δ 5.10 (J 7.0 Hz) for two protons could be due to H-6’ and H-10’. A doublet at δ 4.61 (J 7.0 Hz) was assignable to 2 protons at C-1’. A multiplet at δ 2.34 for 4 protons could be 2*H-5’ and 2*H-9’. A triplet at δ 2.10 (J 7.0 Hz), representing 6 protons, could be 2*H-4’, 2*H-8 and 2*H-12’. Three singlets at δ 1.75, 1.72 and 1.65, each for 3 protons, could be due to 3 methyls, one each at C-3’, C-7’ and C-11’. A multiplet in the range 1,30-0.86 for 9 protons could be 2 methyls, one methylene and one methine. The data suggested the compound to be 7(3’,7’,11’,14’-tetramethy) pentadec-2’,6’,10’-trienyloxycoumarin (3) which is a hitherto unreported compound.
Acknowledgements
The financial assistance provided by ICAR, New Delhi, is gratefully acknowledged.
References
- Bakshi GDN, Sensarama P, Pal DC. A lexicon of medicinal plants in India. Vol. I. Calcutta (India): Naya Prokash; 1999.
- Rastogi RP, Mehrotra BN. Compendium of Indian medicinal plants. Vol. II. New Delhi: Publication and Information Directorate; 1990.
- Sukhdev. Handbook of Terpenoids. Vol. I. Florida (USA): CRC press Inc. Bota Raton; 1989.
- Heilbron I, Cook AH, Bunbury HM, Hey DH. Dictionary of organic compounds. London (UK): Eyre and Spottiswoode; 1965.
- Meera, Meenarani, Kumar S, Kalidhar SB. Phytochemical investigation of Parkinsonia aculeata. Indian J Pharm Sci 1999;61:315-6.