- *Corresponding Author:
- A. Bissoyi
Department of Biotechnology and Medical Engineering, National Institute of Technology, Rourkela‑769 008, India
E-mail: bissoyi.akalabya@gmail.com
| Date of Submission | 01 June 2014 |
| Date of Revision | 25 January 2015 |
| Date of Acceptance | 04 August 2015 |
| Indian J Pharm Sci, 2015;77(4):453-460 |
This is an open access article distributed under the terms of the Creative Commons Attribution‑NonCommercial‑ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non‑commercially, as long as the author is credited and the new creations are licensed under the identical terms.
Abstract
Chikungunya is one of the tropical viral infections that severely affect the Asian and African countries. Absence of any suitable drugs or vaccines against Chikungunya virus till date makes it essential to identify and develop novel leads for the same. Recently, nsP2 cysteine protease has been classified as a crucial drug target to combat infections caused by Alphaviruses including Chikungunya virus due to its involvement viral replication. Here in, we investigated the structural aspects of the nsP2 protease through homology modeling based on nsP2 protease from Venezuelan equine encephalitis virus. Further, the ligands were virtually screened based on various pharmacological, ADME/Tox filters and subjected to docking with the modeled Chikungunya nsP2 protease using AutoDock4.2. The interaction profiling of ligand with the protein was carried out using LigPlot+. The results demonstrated that the ligand with PubChem Id (CID_5808891) possessed highest binding affinity towards Chikungunya nsP2 protease with a good interaction profile with the active site residues. We hereby propose that these compounds could inhibit the nsP2 protease by binding to its active site. Moreover, they may provide structural scaffold for the design of novel leads with better efficacy and specificity for the nsP2 protease.
Keywords
Chikungunya, nsP2 protease, Homology modeling, AutoDock4.2, Molecular dynamics simulation
In recent years, re‑emerging tropical infections have created a huge impact throughout the world. One such tropical infection is Chikungunya, caused by Chikungunya Virus (CHIKV) which belongs to Togaviridae family of Alphavirus genus [1,2]. CHIKV is transmitted to the humans by bite of two main vectors (mosquitoes) including Aedes aegypti and Aedes albopictus. Fever, headache, nausea, vomiting and joint pain forms the major symptoms of CHIKV infections [1]. Absence of specific antiCHIKV drugs or vaccines makes the identification and development of potential drug candidates highly essential. After infection, CHIKV majorly produces two mRNAs; one genomic and other subgenomic mRNA, similar to other alpha viruses [2]. The genomic mRNA is translated into nonstructural proteins, while subgenomic mRNA forms virion structural proteins. The translation of genomic mRNA results into long polyprotein consisting of nsP1, nsP2, nsP3 and nsP4, formed after cleavage of aforesaid polyprotein [2,3]. The nsP2 is multifunctional with nucleoside triphosphatase, helicase, and RNA‑dependent 5‑triphosphatase activities located in the N‑terminus of the protein while the proteolytic domain has been mapped to its C‑terminal part. It forms a papain‑like thiol protease with a catalytic dyad involving two conserved residues cysteine and histidine [3]. In addition, Trp1084 is a highly conserved residue of nsP2 protease active site pocket and is considered necessary for its protease activity [4]. The nsP2 protease is responsible for cleavages in the nonstructural polyprotein that are crucial for the viral replication cycle [3]. Therefore, this protease constitutes an attractive target for the development of antiviral compounds and in‑depth understanding of its structural features will help in efforts made for designing drugs against CHIKV.
In the previous reports, the researchers have demonstrated that Interferon in combination with ribavirin and mercaptopurine possessed antiviral activity against CHIKV [5,6]. In addition, arbidol, which is widely used for the treatment of influenza, also inhibited CHIKV replication [7]. The extracts of the plants namely, Flacourtia ramontchi, Anacolosa pervilleana and Trigonostemon cherrieri have been reported to have inhibitory activity against CHIKV [8‑11]. Although the function of CHIKV nsP2 (CnsP2) protease is very well‑known, yet only a few studies have focused on exploring inhibitors for the same. In a recent report, Jayaprakash et al. has demonstrated antiCHIKV activity of thiazolidinone derivatives via targeting CnsP2 [6]. Also, Nguyen et al. identified the CnsP2 protease inhibitors using the structure based approach [4]. From the pharmacological point of view, the amino acid residues of CnsP2, crucial for interaction with the ligands and their pharmacophore features have been established by Singh et al. and Bassetto et al. [3,12].
In the present investigation, we elucidated the structural aspects of nsP2 protease from CHIKV through molecular modeling based on nsP2 protein from Venezuelan Equine Encephalitis (VEE) virus. Thereafter, virtually screening of potential leads was carried out based on R.E.O.S, Lead like Soft, CAESAR mutagenic and carcinogenic filters. The screened leads were subjected to docking using Autodock4.2 and their interaction profile with CnsP2 was analyzed using PyMOL and LigPlot+. Further, molecular dynamic simulation of protein ligand complex was carried out to analyze the stability of the complex.
Materials and Methods
Homology modeling and structural validation of protein
The amino acid sequence of CnsP2 protein was obtained from the UniProt database (UniProt KB/ Swiss‑Prot entry Q8JUX6 (POLN_CHIKS)) (http:// www.uniprot.org/uniprot/Q8JUX6). The protein sequence was examined for the motifs using ExPASy PROSITE (http://prosite.expasy.org/). The amino acid sequence corresponding to CnsP2 protease was subjected to modeling using Modeller v9.12. The selection of best predicted protein model was based on DOPE score and was further refined based on multiple template and loop modeling by Ab‑initio/loop refinement method [13]. The missing side chains in the best model (if any) were predicted using SCWRL 3.0 software [14]. Thereafter, the model was validated using PROCHECK, ERRAT and VERIFY_3D present at Structural Analysis and Verification server (http://nihserver.mbi. ucla.edu/SAVES/) [15‑17]. Further, electrostatic surface potential was calculated using PyMOL software. The active site prediction in the CnsP2 protease was carried using CASTp Calculation web tool (http://sts‑fw.bioengr. uic.edu/castp/calculation.php).
Virtual screening of novel CnsP2 ligands and structure retrieval
Novel ligands for CnsP2 proteases were screened from PubChem (http://pubchem.ncbi.nlm.nih. gov/) based on their structural similarity with {3‑fluoro‑5‑ [6‑(hexylsulfanyl)‑9H‑purin‑9‑yl] t e t r a h y d r o f u r a n ‑ 2 ‑ y l m e t h a n o l ( R E F 1 ) , [5‑(5‑fluoro‑2,4‑dioxo‑3,4‑dihydropyrimidin‑1 (2H)‑yl)‑3,4‑dihydroxytetrahydrofuran‑2‑yl] methyl methyl hydrogen phosphate (REF2) and 3‑hydroxy‑5‑(3‑hydroxyphenyl)‑1‑ (2‑hydroxypropyl)‑4‑ [(4‑methylphenyl) carbonyl]‑1,5‑ dihydro‑2H‑pyrrol‑2‑one (REF3), potential inhibitor for CnsP2 protease (fig 1). The ligands were further screened on the basis of R.E.O.S and Lead‑like Soft filters present at FAF‑Drug2 (http:// mobyle.rpbs.univ‑paris‑diderot.fr/cgi‑bin/portal. py?form=FAF‑Drugs2#forms:: FAF‑Drugs2) [18‑20]. Thereafter, the selected ligands were analyzed for their mutagenicity and carcinogenicity using VEGA Non Interactive Client v1.0.8 based on CEASER Mutagenicity and Carcinogenicity model. 3D structure of leads in their energy minimized confirmation was deduced using PRODRG Server (davapc1.bioch. dundee.ac.uk/cgi‑bin/prodrg) [21].
Molecular docking
The leads were docked onto the predicted active site of CnsP2 protein based on Lamarckian Genetic Algorithm using AutoDock4.2 software as described by Agarwal et al. [22‑25]. Prior to docking, the protein structure was energy minimized using UCSF Chimera for 500 steps (http://www.cgl.ucsf. edu/chimera) [26]. Further, protein ligand interactions were evaluated using LigPlot+ [27]. The pharmacophore features of the LIG6 and LIG1 were analyzed using Accelrys Discovery Studio Client 4.1. The ADME properties of ligand showing best interaction profiles were analyzed using PreADMET web tool (http://preadmet. bmdrc.org/).
Molecular dynamic simulations
Molecular dynamic simulations of CnsP2 protease complexed with ligand were performed using the GROMACS 4.0.7 under OPLS‑AA force field. The temperature was maintained using Berendsen thermostat algorithm. The pressure was maintained to a reference pressure of 1 bar with a coupling time of 1ps. Interactions within the larger cut‑off were updated every 10 steps. The time step used was 2fs. The simulations lasted for 10ns and were carried out at temperature of 310K. The root mean square distance (RMSD) values were plotted versus the simulation time to judge whether the simulations had reached a level of equilibrium.
Results and Discussion
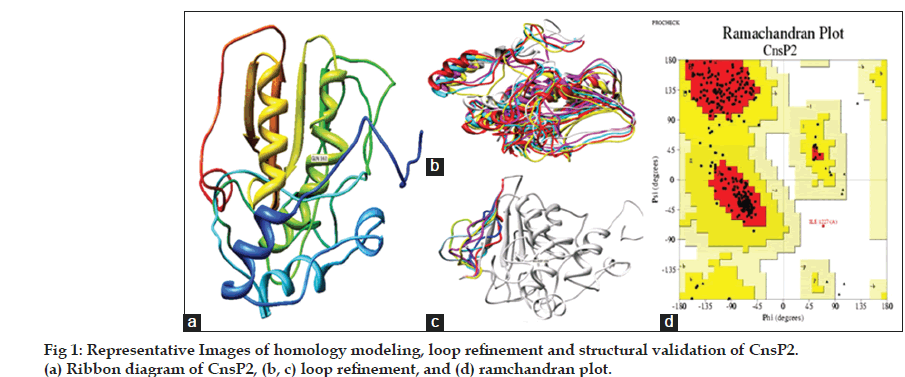
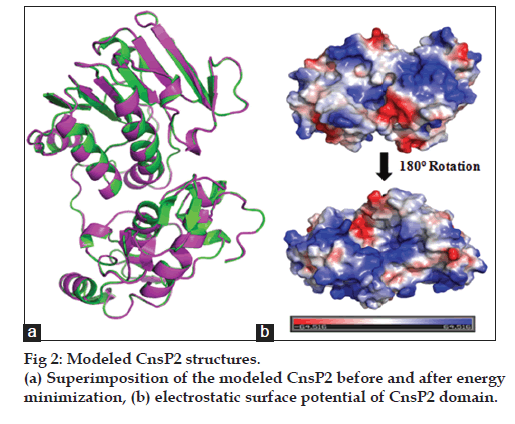
In the present investigation, an integrated computational approach to model nsP2 protease from chikungunya and to identify and design novel inhibitors of nsP2 protease, based on virtual screening was applied. The sequence scanning of CnsP2 protein (amino acids: 536‑1333) demonstrated that the protein is composed of two domains, first helicase domain (amino acids: 690‑991) and nsP2 protease domain (amino acids: 1004‑1327). The nsP2 protease sequence was used for the homology modeling. The three dimensional structure of CnsP2 protease was homology modeled based on alignment with other proteins using NCBI BLASTp. It was observed that the CnsP2 protein sequence showed a best alignment with Venezuelan Equine Encephalitis alpha virus nsP2 Protease Domain (PDB Id: 2HWK) with a percentage similarity of 41% and thus, it was selected as a template for modeling using Modeler v9.12. Further, model refining was carried out based on multiple template and loop modeling by Ab‑initio/loop refinement method using Modeler v9.12. The missing side chains (if any) were predicted and added using SCWRL 3.0 software. The model was energy minimized using UCSF Chimera and evaluated based on DOPE score, PROCHECK, ERRAT and VERIFY_3D. It was observed that the energy minimized structure 87.2%, 12.5% and 0.4% of amino acid residues in core, allowed and disallowed regions; confirmed through Ramachandran Plot (fig 1). 91.75% of amino acid residues had an average 3D‑1D score >0.2 with an overall quality factor of 86.885, confirmed through Verify_3D and Errat analysis respectively. A root mean square value (RMS value) of 0.185 was observed between modeled and energy minimized structure of CnsP2 protease (fig. 2).
Targeting the active sites for the design of potent inhibitors of the protein has proved to be a standard approach in Structure based drug designing. The ligand’s interaction with the active site of the protein results in modulation of protein’s activity, thus resulting in either gain or loss of activity. The predictions through CASTp Calculations demonstrated that the CnsP2 protease active site had a surface area of 816.7 Å2, comprising of 35 amino acid residues including Asn1011, Cys1013, Trp1014, Ile1038, Gln1039, Ala1040, Glu1043, Lys1045, Ala1046, Tyr1047, Ser1048, Val1051, Glu1055, Tyr1079, Asn1082, His1083, Trp1084, Asn1202, Leu1203, Glu1204, Leu1205, Gly1206, Ile1221, Thr1223, Pro1224, Arg1226, Val1234, Asp1235, His1236, Ala1237, Lys1239, Gln1241, Leu1243, Asp1246 and Thr1268. The ExPASy PROSITE predictions indicated Cys1013 and His1083 as catalytic amino acids for CnsP2 and forms the “catalytic diad”. Nguyen et al. demonstrated that Cys1013 and His1083 are sufficiently close to carry out the proton transfer (with only 4.8 Å distance between Cys1013 (S atom) and His1083 (N atom)) [4].
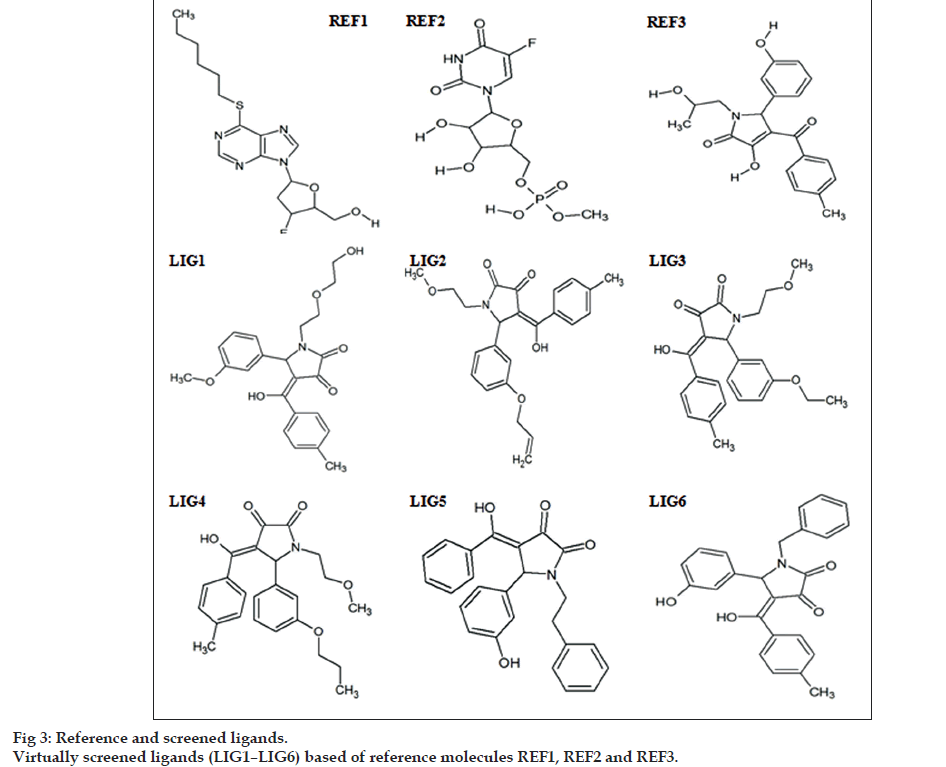
Potential CnsP2 protease inhibitors were virtually screened based on their structural similarity, molecular properties, ADME/Toxicity, mutagenicity and carcinogenicity. Limpon et al. and Singh et al., through in silico analysis have reported that REF1, REF2 and REF3 may possess potential inhibitory activity towards CnsP2 protease (fig. 3) [1,3]. Thus, they were taken as reference ligands for a 3D structural PubChem similarity search with 90% as threshold value. Simultaneously while similarity search, R.E.O.S (Rapid Elimination of Swill) screening filter (Molecular weight (MW): 200‑500, LogP: ‑ 5–5, Hydrogen bond acceptor (HBA): <=10, Hydrogen bond donor (HBD): <=5, Total Polar Surface Area (tPSA): <=150 and Rotatable bonds (RB): <=8) was applied in PubChem. A total of 506/1164 screened out after application of R.E.O.S. filter. Further, screening of the ligands was carried out based on Lead like Soft filter (MW: 150‑400, LogP: ‑ 3–4, HBA: <=7, HBD: <=4, tPSA: <=160, RB: <=9, Rigid bonds: <=30, Rings: <=4, Carbons: 3–35, Heteroatoms: 1–15, H/C Ratio: 0.1–1.1, Stereocenters <=2); present at FAF Drug2 wherein 83/189 got selected. In the lead like soft screening of ligands, the duplicates were not analyzed. Thereafter, the selected ligands were analyzed for their mutagenicity and carcinogenicity using VEGA Non Interactive Client v1.0.8 based on CEASER Mutagenicity and Carcinogenicity model. Finally a total of 6 ligands screened out from 83 ligands with good ADME properties without any carcinogenic/mutagenic effect (Table 1). It is important to mention that the final 6 ligands screened from ADME/Tox filters had structural similarity with only REF3. These molecules were further subjected to molecular docking against active site of CnsP2 proteases using AutoDock4.2.
| Smiles | REF1 C1(=NC=NC2=C1N=C[N] 2C3CC(C(O3)CO[H])F) SCCCCCC | REF2 C1(NC(C(=CN1C2C(C(C(O2) CO[P](=O)(OC)O[H])O[H]) O[H])F)=O)=O | REF3 C1(=CC=C(C=C1)C(C2=C(C(=O) N(C2C3=CC(=CC=C3)O[H]) CC(C)O[H])O[H])=O)C |
|---|---|---|---|
| Structure similarity threshold (%) | 90 | 90 | 90 |
| Similar structures (without filters) | 138 | 666 | 364 |
| R.E.O.S filtering (passed) | 92 | 133 | 281 |
| Lead like soft filtering (tested and duplicates) | 51 and 41 | 56 and 77 | 82 and 199 |
| Lead like soft filtering (passed) | 5 | 0 | 78 |
| Mutagenic and carcinogenic filtering (VEGA NIC) (passed) | 0 | 0 | 6 |
R.E.O.S: Rapid elimination of swill, VEGA NIC: VEGA noninteractive client
Table 1: Virtual Screening Of Ligands Based On Reference Ligands (Ref1, Ref2 And Ref3)
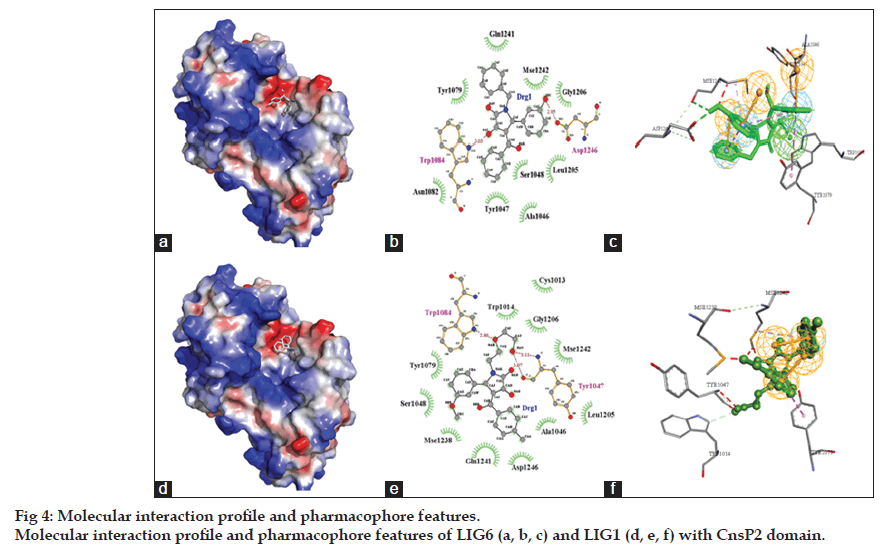
The interaction profile of these virtually screened ligands demonstrated that LIG6 with PubChem Id (CID_5808891) possessed the highest binding affinity with CnsP2 protease with a binding energy of ‑ 8.57 kcal/mol (Ki=0.527 mM). The LIG6 interacted with the protein with three hydrogen bonds with amino acid residues, Trp1084 and Tyr1047. The hydrogen bonds were formed between Trp1084:NE1‑LIG6:OAR, Tyr1047:N‑LIG6:OAU and LIG6:OAU‑Tyr1047:O with corresponding bond length of 2.86, 3.11 and 3.07 Å, respectively. Along with it, LIG1 (CID_5864277) also showed a better affinity towards CnsP2 protease with binding energy of ‑ 8.06 kcal/mol (Ki=1.25 mM). LIG1 interacted via two hydrogen bonds with Trp1084 and Asp1246 amino acid residues of CnsP2 protease. The hydrogen bonds were formed between Trp1084:NE1‑LIG1:OAT and Asp1246:OD2‑LIG1:OBC with bond length of 3.03 and 2.95 Å, respectively. The binding energies of remaining four ligands (LIG2‑4) were found to vary in the range of ‑ 7.55 to ‑ 6.75 kcal/mol. Interestingly LIG1 and LIG6 shared a common binding site at Trp1084:NE1 atom (fig. 4). Also, it is important to mention that only LIG1 interacted hydrophobically with catalytic amino acid residue, Cys1013 (Table 2). It is important to mention that His1083 (catalytic amino acid residue) was found to be in close proximity to the ligands with distance <5 Å, however, this distance is insufficient to form hydrophobic interactions. This indicates that the ligands might inhibit protease activity blocking the binding of substrate to its active site. However, Trp1084 is a highly conserved residue of nsP2 protease active site pocket and is present in close proximity to His1083 and thus considered necessary for its protease activity [4]. Interaction of the ligands with Trp1084 might also result in the reduced catalytic activity of CnsP2 protease. Apart from this, all the six ligands were found to be involved in either hydrogen bonding or hydrophobic interactions with Ser1048, Tyr1079 and Gln1241 which form important amino acid residues for creating microenvironment for the CnsP2 protease activity. None of the ligand was found to be involved in pi‑pi interaction with the CnsP2 protease. The pharmacophore features of the LIG6 and LIG1 important for the interaction with the CnsP2 mainly include hydrogen bond donors, hydrophobic aromatic and hydrophobic interactions (fig. 4c and f).
| Ligands | BE | Ki (µM) | Hydrogen bond interaction | Hydrophobic interactions |
|---|---|---|---|---|
| LIG1 | −8.06 | 1.25 | Tyr1047 (2), Trp1084 | Cys1013, Trp1014, Ala1046, Ser1048, Tyr1079, Leu1205, Gly1206, Mse1238, |
| Gln1241, Mse1242, Asp1246 | ||||
| LIG2 | −6.57 | 15.19 | Ser1048 (2), Gln1241 | Ala1046, Tyr1047, Val1051, Tyr1079, Trp1084, Glu1204, Leu1205, Mse1238, Mse1242 |
| LIG3 | −6.81 | 10.24 | Gln1241 | Ser1048, Gln1050, Val1051, Val1077, Tyr1079, Trp1084, Lys1091, Thr1268, Arg1271 |
| LIG4 | −6.59 | 14.67 | Ser1048 (2), Gln1241 | Ala1046, Tyr1047, Trp1048, Tyr1079, Leu1205, Mse1238, Mse1242, Asp1246 |
| LIG5 | −7.55 | 2.92 | Ser1048, Tyr1079 | Ala1046, Tyr1047, Val1051, Trp1084, Mse1238, Gln1241 |
| LIG6 | −8.57 | 0.526 | Trp1084, Asp1246 | Ala1046, Tyr1047, Ser1048, Tyr1079, Asn1082, Leu1205, Gly1206, Gln1241, Mse1242 |
BE: Binding energies
Table 2: Be And Amino Acid Interactions Of The Modelled Ligands
Further, the molecular features of the selected ligands were analyzed and have been compiled in Table 3. Amongst all the six selected ligands, LIG1 and LIG6 have shown a better affinity for the CnsP2, thus were analyzed for their biological attributes. The analysis revealed that both the ligands had high human intestinal absorption (>94%) and in vitro Caco‑2 cell permeability (>20 nm/sec). It is important to mention that LIG6 had about 22.5 folds higher in vitro MDCK cell permeability, owing to its higher excretion from the body. A higher in vitro plasma protein binding (1.17 folds) of LIG6 in comparision of LIG1 makes is less likely to be available to carry out therapeutic effect (Table 4). However, higher affinity and better interaction profile of LIG6 may overcome its lower availability.
| Ligand | MW | LogP | LogSw | tPSA | Lipinskiviolation | Solubility(mg/L) |
|---|---|---|---|---|---|---|
| LIG1 | 411.447 | 2.372 | −3.518 | 96.3 | 0 | 12,192.59293 |
| LIG2 | 395.448 | 3.399 | −4.136 | 76.07 | 0 | 6321.680999 |
| LIG3 | 407.459 | 3.565 | −4.246 | 76.07 | 0 | 5834.484809 |
| LIG4 | 409.474 | 3.789 | −4.394 | 76.07 | 0 | 5056.971714 |
| LIG5 | 399.438 | 3.994 | −4.764 | 77.84 | 0 | 3406.990984 |
| LIG6 | 385.412 | 3.952 | −4.73 | 80.67 | 0 | 3392.267701 |
MW: Molecular weight, tPSA: total polar surface area
Table 3: Comparitive Account Of Molecular Features Of Selected Ligands
| ADME properties | LIG1 | LIG6 |
|---|---|---|
| Human intestinal absorption (%) | 94.795095 | 95.068488 |
| In vitro Caco‑2 cell permeability (nm/s) | 21.1556 | 20.4712 |
| In vitro MDCK cell permeability (nm/s) | 7.25304 | 162.916 |
| In vitro skin permeability (logKp, cm/h) | −3.25483 | −2.62887 |
| In vitro plasma protein binding (%) | 78.11194 | 91.547559 |
| In vivo blood brain barrier penetration | 0.0180245 | 0.867311 |
| (Concentration in brain/Concentration in blood) |
ADME: Absorption, distribution, metabolism, and excretion
Table 4: Biological Properties Of The Modelled Ligands: Lig1 And Lig6
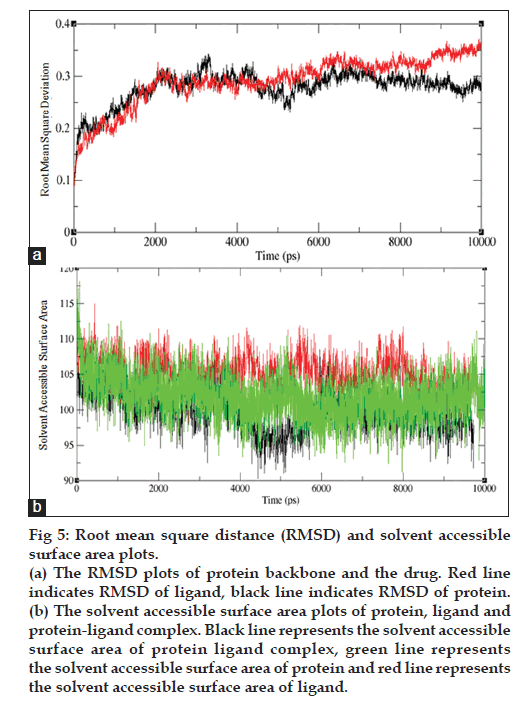
The best ligand‑receptor complex determined by docking calculation, were placed in a box of water using algorithms and simulated for 1ns after initially equilibrating with water molecules for 10 ns. An average structure was energy minimized under conjugated gradient and periodic boundary condition. The dynamic behavior and structural change of the receptor was analyzed by calculating the RMSD value for structural movement and changes in the secondary structural elements of the receptor model during the MD simulation. The structural changes of CnsP2 protease model were evaluated during 10ns MD simulation using GROMACS. The RMSD plots of protein backbone and the drug were plotted in fig 5a. It can be clearly seen from the plots that the complex as well as the protein becomes stable after 2000 ps simulation and the ligand molecule backbone reaches a constant level after 2100 ps at 3.0 Å but suddenly increases after 8500 ps at 3.5 Å and then remain same for 10 ns simulation time period. The backbone RMSD indicates that the rigid protein structure equilibrates rather quickly in this simulation. The RMSD for the drug is more variable indicative of its mobility within the binding pocket. Presence of ligand in protein decreases the solvent accessible surface area. In absence of ligand the surface area increases. Similarly, in absence of protein, solvent accessible surface increases in case of ligand (fig. 5b).
Figure 5: Root mean square distance (RMSD) and solvent accessible
surface area plots.
(a) The RMSD plots of protein backbone and the drug. Red line
indicates RMSD of ligand, black line indicates RMSD of protein.
(b) The solvent accessible surface area plots of protein, ligand and
protein-ligand complex. Black line represents the solvent accessible
surface area of protein ligand complex, green line represents
the solvent accessible surface area of protein and red line represents
the solvent accessible surface area of ligand.
In this complete study, our main objective of this work was to identify the residues involved in the cleavage mechanism through theoretical calculations. The identification of inhibitors for Chikungunya virus has been hampered but a lack of structural insight into any proteins. Therefore, we have chosen to model the nsP2 protein, which plays a vital role in activating the nonstructural protein complex by cleaving the proteins into subunits of nsP1, nsP2, nsP3 and nsP4. CnsP2 protease plays important role in viral replication and propagation, thus inhibiting this protein would be of therapeutic importance. In this regard, we report two ligands CID_5808891 and CID_5864277 that possess good affinity towards CnsP2 protease, thus providing a clue for design of novel drugs with better specificity and affinity. As per our docking analysis, the residues Cys1013, Trp1014, Ala1046, Tyr1047, Ser1048, Tyr1079, Asn1082, Trp1084, Leu1205, Gly1206, Mse1238, Gln1241, Mse1242 and Asp1246 demonstrated crucial interactions with CnsP2 protease. The work is significant as it emphasizes on modeling of CnsP2 protease structure and virtual screening of drugs that may be used as scaffolds for the development of more potent CnsP2 inhibitors.
Acknowledgements
We authors are also grateful to the developers of Autodock 4.2 docking software, UCSF Chimera and LigPlot+ for providing excellent software facilities for carrying out the in silico study.
Financial support and sponsorship
Nil.
Conflicts of interest
There are no conflicts of interest.
References
- Limpon B. Homology modeling and docking to potential novel inhibitor for chikungunya (37997) protein nsP2 protease. J Proteomics Bioinform 2012;5:54‑9.
- Khan AH, Morita K, Parquet MdMdel C, Hasebe F, Mathenge EG, Igarashi A. Complete nucleotide sequence of chikungunya virus and evidence for an internal polyadenylation site. J Gen Virol 2002;83 (Pt 12):3075‑84.
- Singh KD, Kirubakaran P, Nagarajan S, Sakkiah S, Muthusamy K, Velmurgan D, et al. Homology modeling, molecular dynamics, e‑pharmacophore mapping and docking study of Chikungunya virus nsP2 protease. J Mol Model 2012;18:39‑51.
- Nguyen PT, Yu H, Keller PA. Identification of chikungunya virus nsP2 protease inhibitors using structure‑base approaches. J Mol Graph Model 2015;57:1‑8.
- Briolant S, Garin D, Scaramozzino N, Jouan A, Crance JM. In vitro inhibition of Chikungunya and Semliki Forest viruses replication by antiviral compounds: Synergistic effect of interferon‑alpha and ribavirin combination. Antiviral Res 2004;61:111‑7.
- Jadav SS, Sinha BN, Hilgenfeld R, Pastorino B, de Lamballerie X, Jayaprakash V. Thiazolidone derivatives as inhibitors of chikungunya virus. Eur J Med Chem 2015;89:172‑8.
- Delogu I, Pastorino B, Baronti C, Nougairède A, Bonnet E, de Lamballerie X. In vitro antiviral activity of arbidol against Chikungunya virus and characteristics of a selected resistant mutant. Antiviral Res 2011;90:99‑107.
- Konishi E, Hotta S. Effects of tannic acid and its related compounds upon Chikungunya virus. MicrobiolImmunol 1979;23:659‑67.
- Bourjot M, Leyssen P, Eydoux C, Guillemot JC, Canard B, Rasoanaivo P, et al.Flacourtosides A‑F, phenolic glycosides isolated from Flacourtiaramontchi. J Nat Prod 2012;75:752‑8.
- Bourjot M, Leyssen P, Eydoux C, Guillemot JC, Canard B, Rasoanaivo P, et al. Chemical constituents of Anacolosapervilleana and their antiviral activities. Fitoterapia 2012;83:1076‑80.
- Allard PM, Leyssen P, Martin MT, Bourjot M, Dumontet V, Eydoux C, et al. Antiviral chlorinated daphnanediterpenoidorthoesters from the bark and wood of Trigonostemoncherrieri. Phytochemistry 2012;84:160‑8.
- Bassetto M, De Burghgraeve T, Delang L, Massarotti A, Coluccia A, Zonta N, et al. Computer‑aided identification, design and synthesis of a novel series of compounds with selective antiviral activity against chikungunya virus. Antiviral Res 2013;98:12‑8.
- Fiser A, Do RK, Sali A. Modeling of loops in protein structures. Protein Sci 2000;9:1753‑73.
- Canutescu AA, Shelenkov AA, Dunbrack RL Jr. A graph‑theory algorithm for rapid protein side‑chain prediction. Protein Sci 2003;12:2001‑14.
- Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: A program to check the stereochemical quality of protein structures. J ApplCrystallogr 1993;26:283‑91.
- Colovos C, Yeates TO. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci 1993;2:1511‑9.
- Bowie JU, Lüthy R, Eisenberg D. A method to identify protein sequences that fold into a known three‑dimensional structure. Science 1991;253:164‑70.
- Walters WP, Namchuk M. Designing screens: How to make your hits a hit. Nat Rev Drug Discov 2003;2:259‑66.
- Baell JB. Broad coverage of commercially available lead‑like screening space with fewer than 350,000 compounds. J ChemInf Model 2013;53:39‑55.
- Oprea TI, Davis AM, Teague SJ, Leeson PD. Is there a difference between leads and drugs? A historical perspective. J ChemInfComputSci 2001;41:1308‑15.
- Schüttelkopf AW, van Aalten DM. PRODRG: A tool for high‑throughput crystallography of protein‑ligand complexes. ActaCrystallogr D BiolCrystallogr 2004;60 (Pt 8):1355‑63.
- Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, et al. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J ComputChem 1998;19:1639‑62.
- Agarwal T, Asthana S, Gupta P, Khursheed A. In Silico study to elucidate inhibitory effect of thiazides on plasmepsins: Implications of new antimalarial drug design. Int J Pharm PharmSci 2014;6:379‑82.
- Agarwal T, Singh A, Asthana S. Study of binding mode of betulin (lup‑20(29)‑ene‑3β, 28‑diol) on plasmepsin II from Plasmodiumfalciparum: An antimalarial studies. Int J Curr Res 2012;4:505‑8.
- Asthana S, Agarwal T, Banerjee I, Ray SS. In silico screening to elucidate the therapeutic potentials of Asparagamine. Int J Pharm PharmSci 2014;6:247‑53.
- Pettersen EF, Goddard TD, Huang CC, Couch GS, Greenblatt DM, Meng EC, et al. UCSF Chimera – A visualization system for exploratory research and analysis. J ComputChem 2004;25:1605‑12.
- Laskowski RA, Swindells MB. LigPlot+: Multiple ligand‑protein interaction diagrams for drug discovery. J ChemInf Model 2011;51:2778‑86.