- *Corresponding Author:
- T. K. Raja
Departments of Plant Molecular Biology and Biotechnology, Centre for Plant Molecular Biology, Tamil Nadu Agricultural University, Coimbatore-641 003, India
E-mail: tkraja@gmail.com
| Date of Submission | 16 October 2012 |
| Date of Revision | 20 November 2013 |
| Date of Acceptance | 25 November 2013 |
| Indian J Pharm Sci 2014;76(1):10-18 |
Abstract
The genome of the virus H1N1 2009 consists of eight segments but maximum number of mutations occurs at segments 1 and 4, coding for PB2 subunit of hemagglutinin. Comparatively less number of mutations occur at segment 6, coding for neuraminidase. Two antiviral drugs, oseltamivir and zanamivir are commonly prescribed for treating H1N1 infection. Alternate medical systems do compete equally; andrographolide in Siddha and gelsemine in Homeopathy. Recent studies confirm the efficacy of eugenol from Tulsi and vitamins C and E against H1N1. As the protein structures are unavailable, we modeled them using Modeller by identifying suitable templates, 1RUY and 3BEQ, for hemagglutinin and neuraminidase, respectively. Prior to docking simulations using AutoDock, the drug likeness properties of the ligands were screened using in silico techniques. Docking results showed interaction between the proteins individually into selected ligands, except for gelsemine and vitamin E no interactions were shown. The best docking simulation was reported by vitamin C interacting through six hydrogen bonds into proteins hemagglutinin and neuraminidase with binding energies -4.28 and -4.56 kcal/mol, respectively. Furthermore, vitamin C showed hydrophobic interactions with both proteins, two bonds with Arg119, Glu120 of HA, and one bond with Arg74 of NA. In silico docking studies thus recommend vitamin C to be more effective against H1N1.
Keywords
Swine flu, H1N1, Tamiflu, vitamin C, hemagglutinin, neuraminidase
A global outbreak of a new strain of H1N1-2009 influenza virus, often referred to as “swine flu virus” is well-known for causing a huge number of deaths both in human and swine in recent years. The World Health Organization (WHO) reported about 15,174 deaths due to the pandemic influenza virus H1N1 [1]. A pathogenic swine viral of H1N1 subtype has been proven to cause an outbreak of respiratory disease in both human and swine. During 1918-1919, influenza was first described as a disease of swine [2]. The first influenza virus was isolated from swine in 1930; the same was isolated from a human between 1974 and 2005 [3]. H1N1 virus results when a previous triple fusion of bird, pig, and human flu viruses further combines with a Eurasian pig flu virus [4]. While the virus is a major pathogen to humans, it does not disproportionately infect adults older than 60 years, which is an unusual and characteristic feature [5]. Similar to other influenza viruses, H1N1 is transmitted through respiratory droplets and not by eating pork or pork products. Similar to other influenza viruses, H1N1 also contains two surface antigens, namely hemagglutinin (HA) and neuraminidase (NA) [6]. The major role of HA involves in the viral entry mechanism and immune recognition through two subunits namely HA1 containing the receptor binding domain and HA2 responsible for the fusion of the virion with the endosomal membrane in the host cell [7]. Among the eight segments present in the genome of H1N1 virus, mutation is found to occur only at three segments. Maximum number of mutations occur at segments 1 and 4, coding for PB2 subunit of HA and comparatively less mutations occur at segment 6, coding for NA [8]. Among 144 combinatorial possibilities from 16 subtypes of HA and 9 subtypes of NA, only H1N1, H2N2, and H3N2 are human adapted viruses [9].
NA inhibitor antiviral drugs such as oseltamivir (Tamiflu) and zanamivir (Relenza) are widely used in allopathy to control this influenza virus [10]. Recent researches also report that zanamivir is comparatively more effective in controlling the pathogenicity of this virus [10,11]. Yet both the drugs are notorious for their strong side effects. Other alternative medicine systems practiced in India do compete equally in preventing/curing swine flu. A popularly known oldest system of medicine in India namely Siddha prescribes “Nilavembu kudineer”, the extract from the plant Andrographis paniculata against H1N1; the active ingredient is andrographolide [12]. Another famous alternative medicine system widely practiced even in modern India is Homeopathy. Gelsemine, a homeopathic medicine can both prevent and cure swine flu according to the physicians who practice homeopathy [13]. Ayurveda another ancient medicine system of India reports the best form of herbal remedies for swine flu. This corroborates with a natural product called eugenol from Tulsi (holy basil) for curing swine flu. Eugenol is an essential oil present in high concentration in Tulsi, which has both antiviral and antiinflammatory properties [14]. Even certain vitamins like vitamin C [15,16] and vitamin E [16] have been reported to be effective in controlling H1N1 virus.
Research on prevention and cure of swine flu still remains as an open challenge in the field of pharmacogenomics and bioinformatics. A comparative study on the effectiveness of the known drugs can identify the drug with maximum interaction with the receptor protein among the available drugs. Such knowledge is essential for the researchers to discover a potentially new drug for swine flu. The comparative study on the effectiveness of drugs with similar pharmacological actions is common among the researchers to find the most potent drug [17,18]. In our study, we compare the effectiveness of the available drugs in terms of binding energy and hydrogen bond formation between the proteins (HA and NA) and the known drugs/ligands using in silico techniques. We adopted the in silico techniques such as homology modeling to model HA and NA proteins and docking to find the most potent drug among the available drugs. The outcome of our research can give better understanding for the researchers aiming to discover a new drug compound with improved efficacy against swine-flu.
Materials and Methods
Modeling
The amino acid sequences for NA and HA of swine influenza virus subtype H1N1 of A/Hong Kong/2369/2009 (H1N1) were retrieved from protein sequence database situated at NCBI (http://www. ncbi.nlm.nih.gov/). NA sequence consists of 469 amino acids with the molecular weight of about 51.6379 kDa, whereas HA consists of 566 amino acids with the molecular weight of 63.2395 kDa. These protein sequences have GenBank accession number; ACT10319.1 for NA and ACT10316.1 for HA. Templates for protein modeling were retrieved by running the PDB BLAST using the protein sequences.
Though the sequences of HA and NA were retrieved from NCBI database, their protein structural data from PDB database are unavailable. We therefore carried out a comparative modeling for the proteins to predict their 3D structure. We started with BLAST search against PDB in order to find suitable templates for modeling HA. The resulting templates were used for structure modeling using MODELER 9V2. The sequences of targets and templates structures have then been aligned and models were built. As the modeled structures contain few amino acids in the disallowed region of the Ramachandran plot, validation is required to minimize the energy and to move the amino acids into the allowed region. We carried out the loop refinement to bring the amino acid residues present originally in the disallowed region back into the allowed regions on the Ramachandran plot. Force field calculation was used to estimate the energy and stability of modeled structures. To make a stable structure, the energy was minimized in Steepest Descent followed by conjugate gradient method using the default parameters. Finally, we validated the modeled structures by submitting to Procheck-Protein Model Check. Ramachandran plot for the modeled structure is used to view the number of residues in the most favored region, additional allowed region, generously allowed region, and disallowed region. We repeated the same procedure to model the protein NA. Finally we used SPDBV, a freeware to minimize the energy of the modeled structures. The same tool can also be used for loop building, with which we can build the loop for the amino acids, so that it enters into the allowed region.
Ligand selection
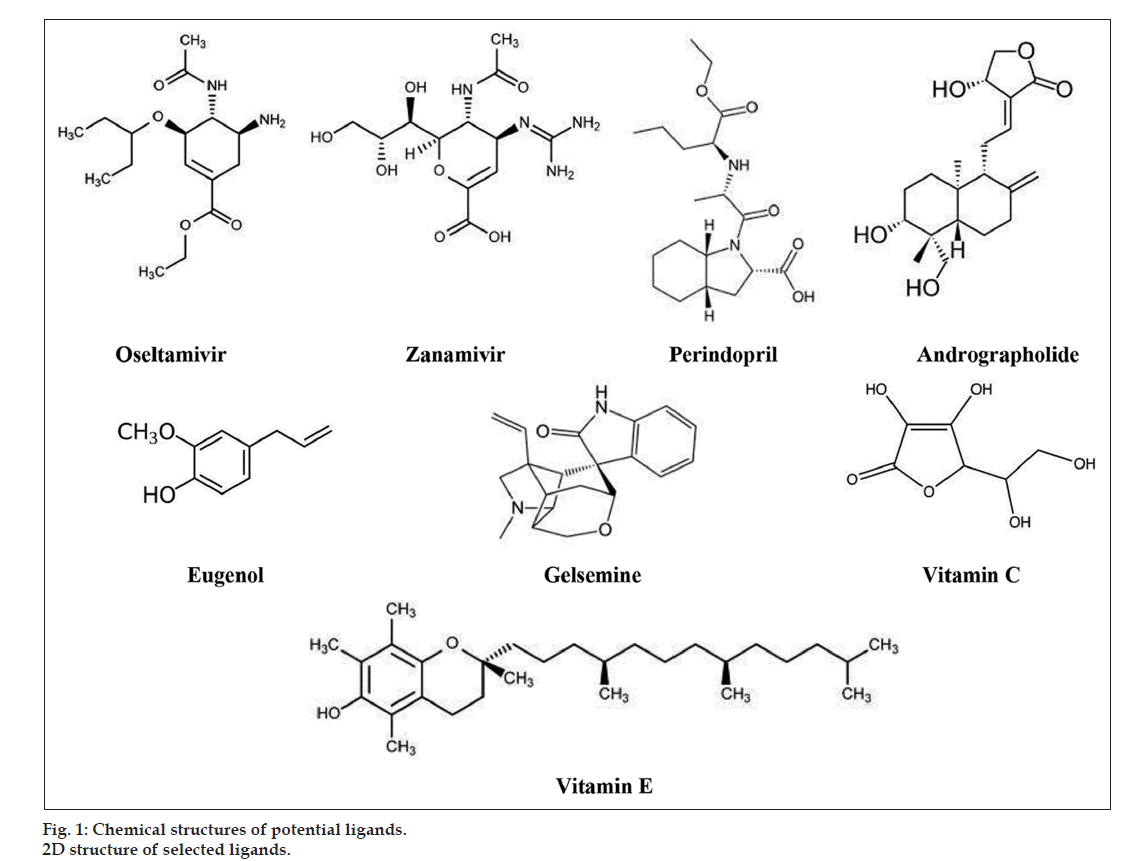
Ligands for our study include two drug molecules from allopathy namely oseltamivir and zanamivir. A structurally similar drug molecule to zanamivir by name perindopril was also selected as a potential ligand. Small molecules from other alternate medicine systems, andrographolide from Siddha, gelsemine from Homeopathy, eugenol from Ayurveda, and two natural products namely vitamin C and vitamin E were selected as potential ligands. We generated the 2D and 3D structures of the proposed ligands using PubChem, a free online database for small molecules (fig. 1). We then saved the structures in pdb format for further docking studies.
Lipinski rule of 5 screening
An essential screening methodology for the rational drug design is by Lipinski rule of 5 [19-21]. The rule states that poor absorption or permeation are more likely when a ligand molecule violates Lipinski rule of 5, that is, has more than five hydrogen bond donors, the molecular weight is over 500, the log P is over 5 and the sum of N and O is over 10 [22,23]. This screening methodology was implemented to analyze the drug likeness of the proposed ligands as it influences the behavior of molecule in a living organism, including bioavailability, transport properties, affinity to proteins, reactivity, toxicity, metabolic stability, and many more. We screened the ligands against Lipinski rule of 5 using Molinspiration (http://www.molinspiration.com/) and their drug likeness was further confirmed using MarvinSketch (http://www.chemaxon.com/marvin/sketch/index.jsp).
ADMET/Bioactivity screening
Many in silico technologies in drug discovery have increased the possibility of finding the new lead compounds at much shorter time period [24]. However, unfavorable ADMET properties have been identified as a major cause of failure even for very promising drug candidate molecules [24,25]. Consequently, earlier prediction of ADMET properties through in silico methods can increase the success rate of candidate molecules prior to reaching development. We utilized an online server FAF-Drugs (http://mobyle.rpbs. univ-paris-diderot.fr/cgi-bin/portal.py?form= admetox) for screening the ADMET properties of the proposed ligands. In addition to the drug likeness screening, Molinspiration implements a sophisticated Bayesian statistics technique to calculate the bioactivity, a number typically between -3 and +3. The bioactivity of the selected ligands toward GPCR ligands, ion channel modulators, kinase inhibitors, and nuclear receptors was calculated as a sum of activity contributions of the sub-structures of each individual ligands using Molinspiration.
Protein-ligland docking
Prior to docking, the two receptor proteins HA, NA and the potential ligands were optimized for proper geometry using AutoDock [26-28], a suite of automated docking tool, which operates on Lamarckian genetic algorithm (LGA) [29]. The grid parameter file for each protein was generated by fixing the number of grid points in x, y, and z-axes to 80×80×80, though the size was changed depending on the ligand size and the distance between two connecting grid points to 0.375 Å. The maximum number of energy evaluations before the termination of LGA run was 5 500 000 and the maximum number of generations of the LGA run before termination was 1000. Other docking parameters were set to the software’s default values. After docking, the ligands were ranked according to their protein-ligand affinity. The accuracy of AutoDock 4.0 results were confirmed by considering clusters of 50 runs of conformations/orientations with the RMSD value of less than 2.0 Å in addition to the lowest binding free energy and hydrogen bonds between the macromolecule and the ligands. Further, the docked conformations were energetically and statistically validated.
Results and Discussion
BLAST was used to retrieve the structural templates from PDB to model the proteins HA and NA since their protein structures are unavailable. The templates identified were 1RUY for HA and 3BEQ for NA with e-values 2e-170, 0.0 and identity scores 57%, 82%, respectively. Models were built for the target proteins from the template structure using MODELLER 9V2, which was then followed by the energy minimization to find the stabilization energy of the predicted target structures. The result of RAMPAGE server confirmed the presence of more than 90% of amino acid residues of both HA and NA in favored and allowed regions of Ramachandran plot. The two models thus obtained were used for the docking study.
Lipinski rule is a rule of thumb to evaluate drug likeness, or determine if a chemical compound with a certain pharmacological or biological activity has properties that would make it a likely orally active drug in humans. The molecular weight, Log P value, number of donors and acceptors for each ligand were generated using Molinspiration. The output confirms that most of the ligands fall within the proposed limit confirming their drug likeness, except zanamivir and vitamin E (Table 1). Zanamivir exceeds the limit of hydrogen bond donor, 9 (must be <=5) and hydrogen bond acceptor, 11 (must be <=10) whereas vitamin E exceed Log P value 9.043 (must be <=5). The output from MarvinSketch (Table 1) for all the ligands was found to be 1 except for zanamivir and vitamin E showing 0, which further confirms that both zanamivir and vitamin E fail to obey Lipinski rule of 5. Thus the drug likeness of zanamivir and vitamin E is not satisfactory and so both of them cannot be prescribed as a drug against H1N1. Cross verification of zanamivir with the FDA approved drugs further highlights that zanamivir has received a warning from the FDA [30]. The prescreening of drugs for Lipinski rule of 5 predicted maximum number of hydrogen bond donors and acceptors in vitamin C with the lowest log P value when compared with other ligands obeying Lipinski rule of 5 (Table 1).
| Drug name | Prediction with Molinspiration | |||
|---|---|---|---|---|
| Molecular weight | H-Bond donors | H-Bond acceptors | Log P (≤5) | |
| (≤500) | (≤5) | (≤10) | ||
| Oseltamivir | 312.41 | 3 | 6 | 0.852 |
| Zanamivir | 332.313 | 9 | 11 | -3.642 |
| Perindopril | 368.474 | 2 | 7 | 2.019 |
| Andrographolide | 350.455 | 3 | 5 | 1.051 |
| Gelsemine | 322.408 | 1 | 4 | 2.501 |
| Eugenol | 164.204 | 1 | 2 | 2.1 |
| Vitamin C | 176.124 | 4 | 6 | -2.693 |
| Vitamin E | 430.717 | 1 | 2 | 9.043 |
Virtual screening using molinspiration
Table 1: Screening Of Ligands For Lipinski Rule
FAF-Drugs, an online ADMET prediction server was used to perform the in silico analysis on the ADMET properties for all the potential ligands under study. Among the various parameters of ADMET in FAF-Drugs, 4 parameters of Lipinski rule also find a place. We thus sub-divided the ADMET parameters into two categories namely, parameters of Lipinski Rule of 5 and other ADMET parameters for easy analysis (Table 2). Failure of zanamivir and vitamin E is finally confirmed for the third time by the ADMET output generated by FAF-DRUGS. In addition to failure to obey 2 parameters of Lipinski rule, number of H-bond donors, 7 (0.0 to 6.0) and Log P value, -3.17 (0.0 to -2.0), zanamivir further fails in one of the ADMET parameters, polar surface area (PSA), 200.72 (0.0 to 150.0). Vitamin E also fails to obey ADMET property by exceeding the value of Log P, 9.95 (0.0 to -2.0) (Table 2). Prescreening of the selected ligands for Lipinski rule of 5 and ADMET properties clearly confirm the failure of both zanamivir and vitamin E to satisfy the drug-likeness properties. Thus zanamivir and vitamin E cannot be used or prescribed as drug to prevent/cure swine flu.
| ADMET properties | Ligands | |||||||
|---|---|---|---|---|---|---|---|---|
| Oseltamivir | Zanamivir | Perindopril | Andrographolide | Gelsemine | Eugenol | Vitamin C | Vitamin E | |
| Flexible bonds (0.0–15.0) | 8 | 6 | 6 | 3 | 1 | 3 | 2 | 12 |
| Rigid bonds (0.0–50.0) | 9 | 10 | 14 | 19 | 27 | 7 | 6 | 11 |
| Ring number (0.0–7.0) | 1 | 1 | 1 | 3 | 6 | 1 | 1 | 2 |
| Ring length (0.0–12.0) | 6 | 6 | 6 | 6 | 6 | 6 | 5 | 6 |
| Carbons (5.0 min) | 16 | 12 | 19 | 20 | 20 | 10 | 6 | 29 |
| Noncarbons (2.0 min) | 6 | 11 | 7 | 5 | 4 | 2 | 6 | 2 |
| Ratio (no carbons/carbons) (0.1–1.0) | 0.375 | 0.916667 | 0.3684 | 0.25 | 0.2 | 0.2 | 1 | 0.068966 |
| Number of charges (0.0–3.0) | 0 | 0 | 2 | 0 | 0 | 0 | 0 | 0 |
| Total charges (-2.0 to 2.0) | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| Polar surface area (PSA) (0.0–150.0) | 90.65 | 200.72 | 95.94 | 86.99 | 41.57 | 29.46 | 107.22 | 29.46 |
Computational approach to derive ADMET properties of the selected ligands
Table 2: Prescreening Of Ligands For Admet Properties
Bioactivity prediction using Molinspiration proves that most of the potential ligands of our study show higher bioactivity toward GPCR ligand, the highest value being 0.08 for gelsemine (Table 3). The highest bioactivity toward ion channel modulators was reported by both perindopril (0.12) and andrographolide (-0.52). Both zanamivir and vitamin E were omitted for bioactivity screening and further docking studies as these two drug molecules failed to pass Lipinski Rule of 5.
| Doed complex | No. of H - bond | Hydrogen bond donor | Hydrogen bond acceptor | Hydrogen bond distance | Lowest binding free energy (kcal/mol) |
|---|---|---|---|---|---|
| Scaling factor of hydrogen bond distance is less than 2.90 Å | |||||
| HA into Oseltamivir | 3 | Tamiflu:: UNK0:H: | Hema::GLU116:OE2 | 1.766 | -3.58 |
| Hema::ARG119:HH22: | Tamiflu::UNK0:O | 2.033 | |||
| NA into Oseltamivir | Hema::PHE112:HN: | Tamiflu::UNK0:O | 1.816 | ||
| 3 | Tamiflu:: UNK0:H: | Neura: ASP69:OD2 | 1.925 | -5.33 | |
| Neura::ARG74:HH11: | Tamiflu::UNK0:O | 1.756 | |||
| Tamiflu::UNK0:H: | Neura: GLU146:OE2 | 2.213 | |||
| HA into Zanamivir | 4 | Zana::UNK0:H: | Hema:: GLY110: O | 1.978 | -5.10 |
| Hema::PHE112:HN: | Zana:: UNK0: O | 1.951 | |||
| Hema::GLY110:HN: | Zana:: UNK0: O | 1.836 | |||
| Zana::UNK0:H: | Hema::ASP111:OD1 | 2.235 | |||
| NA into Zanamivir | 0 | - | - | - | - |
| HA into Perindopril | 2 | Hema:: THR106;HN: | Perindopril::UNK0:O | 1.948 | -6.13 |
| Hema::ARG119:HH22: | Perindopril::UNK0:O | 2.126 | |||
| NA into Perindopril | 3 | Neura::ARG 7:HH22: | Perindopril::UNK0:O | 2.241 | -6.84 |
| Neura::ARG74:HH11: | Perindopril::UNK0:O | 1.949 | |||
| Perindopril::UNK0:H: | Neura::GLU146:OE2 | 1.744 | |||
| HA into Andrographolide | 5 | Hema::THR437:HN: | Andro::LIG1:O: | 2.194 | -6.48 |
| Andro::LIG1:H: | Hema::THR437:OGI | 1.888 | |||
| Andro::LIG1:H: | Hema::ASP434:ODI | 1.932 | |||
| Hema::THR437:HG1: | Andro::LIG1:O: | 2.012 | |||
| NA into Andrographolide | 3 | Neura::ARG74:HH21: | Andro::LIG1:O: | 1.628 | -7.04 |
| Andro::LIG1:H: | Neura::GLU195:OE2 | 2.054 | |||
| Andro::LIG1:H: | Neura::GLU37:OE2 | 2.14 | |||
| HA into Eugenol | 2 | Eugenol::LIG: H: | Hema::ASP103:O | 2.034 | -4.77 |
| Hema::GLY105:HN: | Eugenol::LIGI: O | 2.003 | |||
| NA into Eugenol | 2 | Eugenol::LIG1:H: | Neura::ASP69:OD2 | 1.854 | -4.31 |
| Neura::ARG74:HH11: | Eugenol::LIG1:O | 1.884 | |||
| HA into Vitamin C | 6 | Vitamin C::UNK0:H: | Hema::ILE435:O | 2.062 | -4.28 |
| Hema::ARG119:HH12: | Vitamin C::UNK0:O | 2.046 | |||
| Hema::SER123:HG: | Vitamin C::UNK0:O | 1.986 | |||
| Vitamin C::UNK0:H: | Hema::ASP434:OD1 | 2.023 | |||
| Vitamin C::UNK0:H: | Hema::ILE435:O | 2.247 | |||
| Hema::ILE435:HN: | Vitamin C::UNK0:O | 2.214 | |||
| NA into Vitamin C | 6 | Vitamin C::UNK0:H: | Neura::ASP69:OD2 | 1.986 | -4.56 |
| Neura::ARG74::HH12: | Vitamin C::UNK0:O | 2.174 | |||
| Neura::ARG74::HH21: | Vitamin C::UNK0:O | 2.175 | |||
| Vitamin C::UNK0:H: | Neura::ASP69:OD2 | 2.097 | |||
| Vitamin C::UNK0:H: | Neura::GLU37:OE2 | 2.048 | |||
| Neura::ARG74::HH11: | Vitamin C::UNK0:O | 2.168 | |||
Docking results for 2 proteins vs. 6 ligands using AutoDock 4. HA is hemagglutinin and NA is neuraminidase
Table 3: Summary Of Molecular Interactions Of Ha And Na With Prescreened Ligands
Molecular docking simulations of both proteins NA and HA individually with all prescreened potential ligands namely, oseltamivir, zanamivir, perindopril, andrographolide, gelsemine, eugenol, vitamin C, and vitamin E were conducted with the AutoDock 4.0. There were 50 runs of docking for each of the docking simulation. The accuracy of the AutoDock 4.0 results were confirmed by considering clusters of 50 runs of conformations/orientations with the RMSD value of less than 2.0 Å in addition to the lowest binding free energy and hydrogen bonds between macromolecules. Further, the docked conformations were energetically and statistically validated. Except gelsemine and vitamin E, all other ligands show docking interaction with both HA and NA. Even though docking simulations of HA and NA individually into gelsemine show very low binding energy, -8.57 and -8.29 kcal/mol, respectively, the lack of hydrogen bond formation provide evidence that the interaction is possible only at the surface level. The possibility of van der Waals’ interactions at the surface level is higher than the stronger hydrogen bond formation. Such an interaction is more prone to environmental changes and can alter the drug action.
In addition to hydrogen bond formation, our in silico analysis showed hydrophobic interactions between the two target proteins and the potential ligands except for vitamin E (Table 4). While oseltamivir showed maximum number of hydrophobic interactions, that is, five bonds with HA and seven bonds with NA, vitamin C showed minimum number of hydrophobic interactions, that is, two bonds with HA and one bond with NA. We also observed that amino acid Arg is very commonly involved in hydrophobic interaction except for eugenol with NA interaction and gelsemine with both HA and NA interaction.
| Ligands | Hemagglutanin | Neuraminidase | ||
|---|---|---|---|---|
| Bonds | Amino acids in bond formation | Bonds | Amino acids in bond formation | |
| Oseltamivir | 5 | Try436 | 7 | Asp69 |
| Arg119 | Arg74 | |||
| Glu120 | Arg36 | |||
| Glu116 | Arg143 | |||
| Asp434 | Ile141 | |||
| Arg70 | ||||
| Alu146 | ||||
| Zanamivir | 4 | Arg119 | 4 | Asp69 |
| Ile435 | Glu146 | |||
| Trp436 | Ile141 | |||
| Asp434 | Arg143 | |||
| Perindopril | 5 | Glu116 | 5 | Arg36 |
| Arg119 | Tyr320 | |||
| Ile435 | Lys350 | |||
| Trp436 | Arg286 | |||
| Asp434 | Pro349 | |||
| Andrographolide | 4 | Arg119 | 5 | Glu196 |
| Thr437 | Arg36 | |||
| Ile435 | Tyr320 | |||
| Trp436 | Ile141 | |||
| Arg143 | ||||
| Eugenol | 4 | Ile435 | 3 | Gln348 |
| Glu116 | Gly65 | |||
| Glu120 | Thr66 | |||
| Arg119 | ||||
| Gelsemine | 3 | Asp434 | 3 | Glu196 |
| Trp436 | Tyr320 | |||
| Ile435 | Asp69 | |||
| Vitamin C | 2 | Arg119 | 1 | Arg74 |
| Glu120 | ||||
| Vitamin E | 0 | - | 0 | - |
Hydrophobic interaction of hemagglutinin HA and neuraminidase NA with potential ligands
Table 4: Summary Of Hydrophobic Interactions Of Ha And Na With Prescreened Ligands
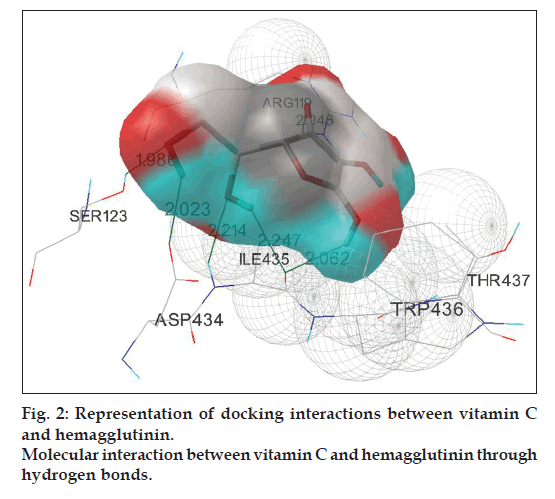
Fifty runs of individual docking simulations of 5 ligands namely oseltamivir, perindopril, andrographolide, eugenol, and vitamin C with HA using the RMSD – tolerance of 2.0 Å resulted in multiple, distinct conformational clusters. Analysis of the positions and orientations of the targets in the best docking models show the formation of 2 different hydrogen bonds, both for perindopril and eugenol, with lowest binding energy -6.13 and -4.77 kcal/mol, respectively. The docking simulation of oseltamivir with HA confirms the formation of three hydrogen atoms with lowest binding energy -3.58 kcal/mol. Interaction of andrographolide with HA is comparatively better than the above three ligands, which is confirmed by the formation of 5 hydrogen bonds with lowest binding energy -6.48 kcal/mol. Maximum number of hydrogen bond formation is observed in the docking complex of vitamin C with HA (fig. 2). The #1 ranked conformer from the cluster-1 of the complex was the favored structure and repeated 26 times out of 50 runs. Out of the six hydrogen bonds formed between vitamin C and HA, three of them run between the backbone oxygen atoms of ligand with ARG119, SER123 and ILE435 residues on HA. The remaining three hydrogen bonds extend from hydrogen atoms of vitamin C with ILE435, ASP434, and ILE434 residues on HA. The best-ranked binding energy of this complex was estimated to be -4.28 kcal/mol with cluster RMSD and reference RMSD 1.93 and 88.79, respectively (Table 4).
A total of 50 docking runs with the RMSD-tolerance of 2.0 Å were carried out in the simulation of various prescreened ligands into NA. Oseltamivir, perindopril, and andrographolide interact with NA through the formation of three hydrogen bonds. The lowest binding energy among the three simulations has been reported by andrographolide -7.04 kcal/mol when compared with perindopril -6.84 kcal/mol and oseltamivir -5.33 kcal/mol. Eugenol is observed to interact through the formation of two hydrogen bonds with the lowest binding energy of -4.31 kcal/mol. However, docking simulation of zanamivir shows no interaction with NA (Table 3). This in silico observation further supports the warning of FDA for zanamivir [30].
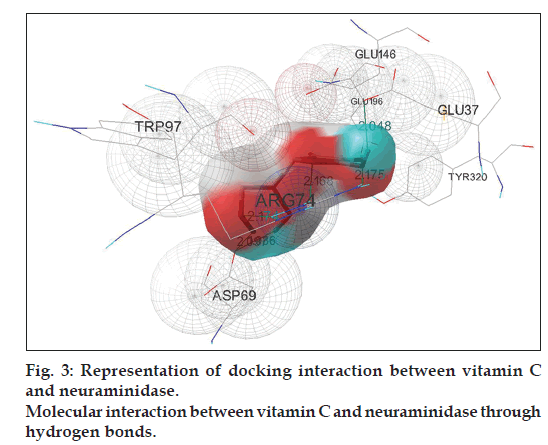
Docking simulation of vitamin C interacts into NA confirms the formation of maximum number of hydrogen bonds, six in total (fig. 3). This is similar to vitamin C–HA simulation complex. Half of the total hydrogen bonds are formed between the backbone oxygen atoms of the ligand with ARG74 of three different chains of NA and the remaining three hydrogen atoms run between the ligand to ASP69 residue on two different chains and GLU37 of NA. Number of distinct conformational clusters reported is 7 out of 50 runs. The favored structure identified is the #1 ranked conformer from the cluster-1 of the complex that repeats 23 times out of 50 runs. The best-ranked binding energy of this complex is estimated to be -4.56 kcal/mol (Table 4) with cluster RMSD and reference RMSD 0.34 and 15.28, respectively.
The efficacy of a ligand can be explained in terms of the number of hydrogen bonds formed between the ligand and protein, as well as with the overall binding energy of the ligand to the receptor protein. The binding energy in the process is expressed as the reduction in free energy due to binding between the ligand and receptor protein. Therefore the efficacy of the ligand is directly related to the number of hydrogen bonds formation with the reduction in free energy for binding. The in silico analysis of the selected ligands with HA and NA confirms the formation of hydrogen bonds with reduction in binding energy by six ligands namely oseltamivir, zanamivir, perindopril, andrographolide, eugenol, and vitamin C (Table 3). Andrographolide shows the lowest binding energy of -6.48 kcal/mol toward HA and -7.04 kcal/mol toward NA with the formation of five and three hydrogen bonds, respectively. In contrast, vitamin C shows the maximum number of hydrogen bond formation (six for both HA and NA) with lower binding energy of -4.28 kcal/mol for HA binding and -4.56 kcal/mol for NA binding.
Vitamin C at sufficiently high doses, 30 000-200 000 mg is accepted by the physicians for its ability to cure many viral diseases such as common cold, flu, hepatitis, viral pneumonia, and even polio [31-33]. However, the daily requirement of vitamin C differs among people (between 2000 and 20 000 mg) because of the differences in genetics and individual biochemistry [34]. Interestingly Pauling [35], and Hoffer and Saul [36] have proven the importance of vitamin C for healthy life by consuming large doses of vitamin C and lived above 90 years of age. Hoffer successfully prescribed vitamin C as antiviral drug for thousands of patients for over 55 years of medical practice [31,32].
Several mechanisms for the antiviral action of vitamin C have been suggested so far [37]. The antioxidant property of vitamin C enhances the defending mechanism of the body against microbes and viruses that propagate in stress conditions [38]. The specific antiviral effect of vitamin C to inactivate DNA of viruses such as bacteriophage deltaA [39], to suppress the replication of human immunodeficiency virus (HIV) [40] and to cure the viral infection by herpes viruses and paramyxoviruses [41] are some of the strong evidences for the antiviral action of vitamin C.
The in silico docking analysis of vitamin C with HA and NA shows strong drug–receptor binding through hydrogen bond formation and reduction in binding energy. The interaction is further confirmed by the hydrophobic interaction between vitamin C with HA and NA (Table 4).
In our study, most ligands passed through the prescreening phase on physio-chemical properties such as Lipinski rule, ADMET and bioavailability. Yet, zanamivir and vitamin E failed to satisfy a few drug likeness parameters and were not included for further docking simulation studies. Our docking results explain that the number of clusters with each prescreened ligands vary in the range from 6 to 24 for HA and from 2 to 10 for NA, indicating that the binding specificity for each ligand varies in both HA and NA. The hydrogen bond makes an important contribution to the interaction between the ligand and the protein. Lack of hydrogen bond formation for gelsemine toward both HA and NA confirms its lack of interaction with both proteins. Number of hydrogen bonds for both the proteins is the same for eugenol and oseltamivir. Interaction of HA with perindopril and eugenol happens through two hydrogen bonds each, whereas andrographolide reports five hydrogen bonds. Likewise, interaction of NA with oseltamivir, perindopril, and andrographolide are through three hydrogen bonds each and that of eugenol is through two hydrogen bonds. Vitamin C reports the maximum number of hydrogen bond formation with both HA and NA, that is, six bonds in each simulation. In this work, we identified that zanamivir, vitamin E, and gelsemine lack potency and efficacy to prevent/ cure swine flu. Further, our in silico docking studies strongly recommend vitamin C to be more effective against H1N1 through (i) maximum number of hydrogen bonds formation, i.e. six for each protein HA and NA (ii) the low binding energy -4.28 and -4.56 kcal/mol for HA and NA, respectively.
References
- WHO Pandemic (H1N1) 2009 - update 86. Global alert and response (GAR). 2010. Availabel from: http://www.who.int/csr/don/2010_02_5/ en/index.html. [Last accessed on 2013 Nov 17].
- Koen JS. A practical method for field diagnosis of swine diseases. J Vet Med 1919;14:468-70.
- Brockwell-Staats C, Webster RG, Webby RJ. Diversity of Influenza Viruses in Swine and the Emergence of a Novel Human Pandemic Influenza A (H1N1). J Proteomics Bioinform 2009;3:207-13.
- Trifonov V, Khiabanian H, Rabadan R. Geographic dependence, surveillance, and origins of the 2009 influenza A (H1N1) virus. N Engl J Med 2009;61:115-9.
- Writing Committee of WHO, Consultation on Clinical Aspects of Pandemic (H1N1) 2009 influenza. N Engl J Med 2010;362:1708-19.
- Zaraket H, Saito R, Sato I, Suzuki Y. Molecular evolution of human influenza a viruses in a local area during eight influenza epidemics from 2000 to 2007. Arch Virol 2009;154:285-329.
- Dhanachandra KS, Karthikeyan M. In silico genome analysis and drug efficacy test of influenza A virus (H1N1) 2009. Indian J Microbiol 2009;49:358-64.
- Kirti K, Pooja S, Rup L. Swine flu virus H1N1: A threat to human health. Indian J Microbiol 2009;49:201.
- Morens D, Taubenberger J, Fauci A. The persistent legacy of the 1918 influenza virus. N Engl J Med 2009;361:225-9.
- Moscona A. Global transmission of oseltamivir-resistant influenza. N Engl J Med 2009;360:953-6.
- Hayden F. Antiviral resistance in influenza viruses–implications for management and pandemic response. N Engl J Med 2006;354:785-6.
- Chen JX, Xue HJ, Ye WC, Fang BH, Liu YH, Yuan SH, et al. Activity of andrographolide and its derivatives against influenza virus in vivo and in vitro. Biol Pharm Bull 2009;32:1385-91.
- Ians. Homeopathy can prevent, cure swine flu. The Times of India, Health and Fitness, 2009. Available from: http://articles.timesofindia. indiatimes.com/2009-08-18/health/28171243_1_swine-flu-homeopathy-homeopathic-treatment. [Last accessed on 2013 Nov 17].
- Tiwari UK. Sacred basil/Tulsi can help prevent, or speed recovery from the H1N1 swine influenza. Alternative-health, 2009. Available from: http://alternative-health.weebly.com/tulsi---sacred-basil---antiviral.html. [Last accessed on 2013 Nov 17].
- Siddhadreams. The cure for swine flu. Swine flu Vs Vitamin C, 2009. Available from: http://siddhadreams.wordpress.com/2009/08/17/swine-flu-vs-vitamin-c/. [Last accessed on 2013 Nov 17].
- Writer NM. How to Avoid The H1N1 Swine Flu With Vitamins And Supplements. eHow, 2009. Available from: http://www.ehow. com/how_4947989_avoid-swine-flu-vitamins-supplements.html. [Last accessed on 2013 Nov 17].
- Faraone SV, Biederman J, Spencer TJ, Aleardi M. Comparing the Efficacy of Medications for ADHD Using Meta-analysis. Med Gen Med 2006;8:4.
- Cipriani A, Furukawa TA, Salanti G, Geddes JR, Higgins JP, Churchill R, et al. Comparative efficacy and acceptability of 12 new-generation antidepressants: A multiple-treatments meta-analysis. Lancet 2009;373:746-58.
- Ekins S, Rose J. In silico ADME/Tox: The state of the art. J Mol Graph Model 2002;20:305-9.
- Miteva MA, Violas S, Montes M, Gomez D, Tuffery P, Villoutreix BO. FAF-drugs: Free adme/tox filtering of compound collections. Nucleic Acids Res 2006;34:W738-44.
- Smith PA, Sorich MJ, Low LS, McKinnon RA, Miners JO. Towards integrated ADME prediction: Past, present and future directions for modelling metabolism by UDPglucuronosyltransferases. J Mol Graph Model 2004;22:507-17.
- Lipinski CA, Lombardo F, Dominy BW, Feeney PJ. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv Drug Deliv Rev 2001;46:3-26.
- Lipinski CA. Drug-like properties and the causes of poor solubility and poor permeability. J Pharmacol Toxicol Methods 2000;44:235.
- Fumiyoshi Y, Mitsuru H. In Silico Approaches for Predicting ADME properties of Drugs. Drug Metab Pharmacokinet 2004;19:327-38.
- Darko B, Matthew DS, Katrina F. Predicting ADME properties In Silico: Methods and models. Drug Discov Today 2002;7:S83-8.
- Goodsell DS, Morris GM, Olson AJ. Automated docking of flexible ligands: Applications of AutoDock. J Mol Recognit 1996;9:1-5.
- Jones G, Willett P, Glen RC, Leach AR, Taylor R. Development and validation of a genetic algorithm for flexible docking. J Mol Biol 1997;267:727-48.
- Rarey M, Kramer B, Lengauer T, Klebe G. A fast flexible docking method using an incremental construction algorithm. J Mol Biol 1996;261:470-89.
- Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comp Chem 1998;19:1639-62.
- Cashin JN. Important drug warning: RELENZA ® (zanamivir) Inhalation Powder Must Not Be Nebulized, 2009. Available from: http://www.fda.gov/downloads/Safety/MedWatch/SafetyInformation/ SafetyAlertsforHumanMedicalProducts/UCM186224.pdf. [Last accessed on 2013 Nov 17].
- Levy TE. Curing the Incurable: Vitamin C, Infectious Diseases, and Toxins. 2002;ISBN-13: 9781401069636.
- Hickey S, Saul AW. Vitamin C: The Real Story, the Remarkable and Controversial Healing Factor 2008;ISBN-13: 9781591202233.
- Klenner FR. The significance of high daily intake of ascorbic acid in preventive medicine. In: Williams R, editor. Physician’s Handbook on Orthomolecular Medicine. 3rd ed. New York: Pergamon Press; 1979. p. 51-9.
- Williams RJ, Deason G. Individuality in vitamin C needs. Proc Natl Acad Sci USA 1967;57:1638-41.
- Pauling L. How to Live Longer And Feel Better. Corvallis, OR: Oregon State University Press; 1986;ISBN-13: 9780870710964.
- Hoffer A, Saul AW. Orthomolecular Medicine for Everyone: Megavitamin Therapeutics for Families and Physicians. Laguna Beach, CA: Basic Health Publication, Inc.; 2009;ISBN-13: 9781591202264.
- Webb AL, Villamor E. Update: Effects of antioxidant and non - antioxidant vitamin supplementation on immune function. Nutr Rev 2007;65:181-217.
- Kastenbauer S, Koedel U, Becker BF, Pfister HW. Oxidative stress in bacterial meningitis in humans. Neurology 2002;58:186-91.
- Murata A, Oyadomari R, Ohashi T, Kitagawa K. Mechanism of inactivation of bacteriophage deltaA containing single-stranded DNA by ascorbic acid. J Nutr Sci Vitaminol 1975;21:261-9.
- Harakeh S, Jariwalla RJ, Pauling L. Suppression of human immunodeficiency virus replication by ascorbate in chronically and acutely infected cells. Proc Natl Acad Sci USA 1990;87:7245-9.
- White LA, Freeman CY, Forrester BD, Chappell WA. In vitro effect of ascorbic acid on infectivity of herpesviruses and paramyxoviruses. J Clin Microbiol 1986;24:527-31.