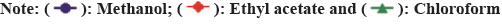- *Corresponding Author:
- N. Shivaiah
Department of Studies and Research in Biochemistry, Tumkur University, Tumkur, Karnataka 572103, India
E-mail: nagarajubiochem@gmail.com
| Date of Received | 02 June 2022 |
| Date of Revision | 17 March 2023 |
| Date of Acceptance | 20 November 2023 |
| Indian J Pharm Sci 2023;85(6):1731-1744 |
This is an open access article distributed under the terms of the Creative
Commons Attribution-NonCommercial-ShareAlike 3.0 License, which
allows others to remix, tweak, and build upon the work non-commercially,
as long as the author is credited and the new creations are licensed under
the identical terms
Abstract
Argyreia imbricata belongs to the family Convolvulaceae and is commonly referred as imbricate wattle. The plant has been known to exhibit in vitro and in vivo anti-diabetic activities. In this study, we have prepared the leaves extract using three different solvents methanol, ethyl acetate, and chloroform separately by the Soxhlet extraction method. The extracts were analyzed for phytoconstituents by gas chromatography-mass spectrometry. The extracts were studied for total antioxidant activity, ferric reducing antioxidant power assay, 1,1-diphenyl-2-picrylhydrazyl radical scavenging activity, 2,2-azino-bis-3-ethylbenzthiazoline-6- sulphonic acid radical cation decolorization assay, hydrogen peroxide scavenging assay and lipid peroxidation activities. Molecular docking was used to further study interaction of the phytoconstituents. L-ascorbic acid is used as a standard antioxidant in all the antioxidant assays. Among the three extracts, methanol extract showed the highest antioxidant activity, as measured by total antioxidant activity of 1.231μg/ml and ferric reducing antioxidant power of 14.9 μg/ml. Furthermore, methanol extract showed significant 1,1-diphenyl-2- picrylhydrazyl radical scavenging activity with an half maximal inhibitory concentration value of 7.06 μg/ml, 2,2-azino-bis-3-ethylbenzthiazoline-6-sulphonic acid radical cation decolorization assay with a value of 13.91 μg/ml, hydrogen peroxide scavenging assay with a value of 19.71 μg/ml and inhibited of lipid peroxidation with half maximal inhibitory concentration value of 128.4 μg/ml. The presence of different phytoconstituents was analyzed by gas chromatography-mass spectrometric analysis. The potential interactions between identified phytoconstituents and the antioxidant target protein were studied using docking study. The phytoconstituents of methanol extract scored the highest interaction with 2HE3. This study focuses on the pharmacological discovery process for preventing cell damage by oxidants.
Keywords
Argyreia imbricata, antioxidant activity, ferric reducing antioxidant power assay, 1,1-diphenyl-2-picrylhydrazyl, hydrogen peroxide scavenging assay, 2,2-azino-bis-3-ethylbenzthiazoline-6-sulphonic acid, molecular docking
Studies from several years have evidenced that Reactive Oxygen Species (ROS) have a significant role in the pathophysiology of a wide range of illnesses. Enzymatic and non-enzymatic ROS defense systems are found in most living organisms. These processes can maintain a reactive equilibrium in healthy environments[1]. It is crucial to highlight that ROS playsan important role in the activation and development of inflammation. Inflammatory responses are triggered by an overabundance of the ROS produced by metabolic processes, including superoxide radical anion (SOR), hydrogen peroxide (H2O2), hydroxyl radical (OH) and nitric oxide (NO)[2].
Antioxidants shield the cellular damage from reactive oxygen species and the most common "antioxidants" include Vitamins A (retinol), C (L-ascorbic acid), E (tocopherol), β-carotene, minerals like selenium, and naturally occurring polyphenols are all antioxidants. In foods such as fruits and vegetables, vitamins and β-carotene include conjugated double bonds and antioxidant functional groups. Antioxidants are available in the diet and in addition people also take supplements containing antioxidants[3]. Czernichow et al. study have shown that antioxidant supplements have adverse effect associated with metabolic syndrome. It is important to investigate plant-based sources with antioxidant and free radicle quenching properties without inducing metabolic syndrome[4].
Medicinal plants, on the other hand, have long been utilizedin traditional medicine and have preventative properties, particularly in developing countries. Several medicinal plants' antioxidant capabilities have been investigated. Natural antioxidants including raw extracts or chemical components are extremely effective at preventing oxidative stress-induced damage[5]. A wide variety of herbal extracts and natural substances have been shown in preclinical studies to reduce oxidative stress. In macrophage cells, Thring et al. looked for anti-inflammatory potential in 30 different plant extracts. Interleukin (IL)-6 and tumor necrosis factor-α were found to be inhibited, while IL-10 production was increased, and the expression of Cyclooxygenase-2 and nitric oxide synthase was decreased. Their anti-inflammatory mechanisms were also identified[6].
With over 220 species, Argyreia is one of the major genera in the Convolvulaceae family, found across Asia, including India. Argyreia imbricata (A. imbricata) is prevalent in southern India at altitudes upto 300 m above mean sea level. The flowering and fruiting season is August-December for this dicotyledonous plant. A huge white woollyclimber, asymmetrical, strigose leaves 8-12 cm long, with an obtusely sharp, rounded, or subcordate base, 3 cm petiole, 5 cm peduncle, small bracteoles and bracts. It has little flowers with a short pedicellatecalyx lobe and a 2 cm long pink corolla and it has 5 mm berry, reddish, thickly hairy[7].
Earlier studies have shown that A. imbricata exhibited anti-diabetic effect in a streptozotocin-induced diabetes model in Wistar albino rats. This property was seen in different solvent extracts such as petroleum ether, chloroform, ethyl acetate and methanol extracts. The in vitro antidiabetic efficacy of all extracts was evaluated using α-amylase and β-glucosidase inhibition studies[8].
The A. imbricata has not been explored for its phytoconstituents and antioxidant, lipid peroxidation properties. In this study, we have prepared the different solvent extracts of the A. imbricata leaves. The extracts were analyzed for bioactive composition by Gas Chromatography-Mass Spectrometric (GC-MS) analysis, antioxidant and lipid peroxidation properties. Further, the molecular docking study was conducted. This is the first report to study the phytoconstituents analysis, antioxidant potentials of A. imbricata leaves solvent extracts and molecular docking studies for identified phytoconstituents.
Materials and Methods
Materials:
Chemicals used in this study were gallic acid, sodium carbonate, folin-ciocalteau reagent, sodium phosphate, sulphuric acid, ammonium molybdate, sodium acetate, ferric chloride, tripyridyl triazine (TPTZ), hydrochloric acid, ascorbic acid, methanol, 1,1-diphenyl-2-picrylhydrazyl (DPPH), 2,2-azino-bis-3-ethylbenzthiazoline-6-sulphonic acid (ABTS), potassium persulfate, hydrogen peroxide, glacial acetic acid, sodium phosphate, monopotassium phosphate, potassium hydroxide, sodium dodecyl sulfate, thiobarbituric acid, trichloroacetic acid, monosodium phosphate, sodium chloride were purchased from Sisco Research Laboratories chemicals and SD Fine chemicals. All other chemicals and solvents used were of analytical grade.
Collection of plant material:
A. imbricata plant was collected from Tiptur local area, Tumkur district, Karnataka, India at a Latitude of 13° 15' 36.00" N Longitude of 76° 28' 48.00" E and during the period of 1st August 2021 to 30th November 2021. The plant was authenticated by Prof. Sharanappa, Department of Studies and Research in Biosciences, Hemagangotri, University of Mysore, Hassan. The plant herbarium was kept at Department of Studies and Research in Botany, Tumkur University, Tumakuru.
Extraction of plant material:
The A. imbricata leaves were collected washed and shade dried. The dried leaves were powdered using a mixer and used for extraction. The leaves extract of A. imbricata was prepared by using three different solvents (methanol, ethyl acetate, and chloroform). The leaves were extracted with different solvents individually by using the Soxhlet extraction method. 20 g of leaves powder was suspended in a solvent and extracted for 9 to 10 h in the Soxhlet apparatus. The extracts were filtered and vacuum-condensed at 45°. The extracts were kept in the refrigerator until they were used.
Determination of antioxidant activities:
Determination of Total Antioxidant Activity (TAA): TAA was done by the phosphomolybdenum technique according to the method of Prieto et al.[9]. In 1 ml of a standard reagent (0.6 M H2SO4, 28 mM sodium phosphate, and 4 mM ammonium molybdate) different quantities of the extracts (100-500 µg/ml) were added. The tubes were capped and placed in a water bath at 95° for 90 min. The absorbance was measured at 695 nm against a blank after cooling to room temperature. The TAA per g of the extract was measured in mg of Gallic Acid Equivalence (GAE).
Ferric Reducing Antioxidant Power assay (FRAP): According to Benzi et al.[10], FRAP assay of extracts was determined. 1.0 ml FRAP reagent was mixed with 1.0 ml extracts (100-500 µg/ml). The FRAP reagent was made by combining 300 mM acetate buffer, 10 ml TPTZ dissolved in 40 mM HCl, and 20 mM FeCl3.6H2O in a 10:1:1 ratio. The tubes were incubated for 30 min at 37°. At 593 nm, absorbance was measured against a blank. The FRAP values were measured in mg of GAE per g of extracts.
DPPH radical scavenging activity: According to Braca et al.[11], 50 µl of extracts (5-25 µg/ml) were added to 300 µl of an ethanolic solution containing 0.5 mM DPPH. The tubes were incubated for 5 min at 37° in a dark environment. At 517 nm, the absorbance was measured against a blank. The extract's radical scavenging activity, reported as percent inhibition, was estimated using the equation below.
Percentage inhibition of DPPH radical=[(absorbance control-absorbance test)/absorbance control]×100
ABTS radical cation decolorization assay: According to Seeram et al.[12], the ABTS radical cation decolorization experiment was done. To make an ABTS solution, water containing 7 mM ABTS was mixed with 2.45 mM potassium persulfate (1:1). For 16 to 20 h, this reaction mixture was kept in a container at room temperature. Using methanol as a diluting agent, we prepared a solution of ABTS that had an absorbance of 0.7 at 734 nm. After 30 min of incubation, various doses of extracts (10 to 50 μg/ml) were added to ABTS solution to make a total volume to 5 ml. The absorbance was read at 734 nm. Percent scavenging of extracts was used to measure the results, which were presented as percentages.
Hydrogen peroxide scavenging assay: Ruch et al. method's was used to determine the effectiveness of extracts in scavenging hydrogen peroxide[13]. Phosphate buffer was used to make a 40 mM hydrogen peroxide solution (50 mM pH 7.4). Hydrogen peroxide was added to extracts (10-50 µg/ml) in a total volume of 2 ml and the absorbance was read at 230 nm. To study the hydrogen peroxide scavenging activity in percentage, the above mentioned equation was used.
Lipid peroxidation: By measuring Thiobarbituric Acid Reactive Substances (TBARS), lipid peroxidation was determined as indicated by Ohkawa et al.[14]. The liver homogenate (10 % in cold phosphate buffered saline, pH 7.4) was treated with different concentrations of the extract (100-500 µg/ml) in water. The acetic acid (20 % v/v, pH 3.5), Sodium Dodecyl Sulfate (SDS) (8 % w/v, 0.2 ml), and thiobarbituric acid (0.8 % w/v, 1.5 ml) tubes were filled with the extracts. The mixture was kept in hot water bath for 45 min. The samples were read at 532 nm, L- ascorbic acid was used as the standard and the TBARS produced were extracted into three milliliters of 1-butanol. Malondialdehyde (MDA) equivalents were quantified in terms of nmol MDA produced per mg protein.
GC-MS analysis: GC-MS analysis was conducted for extracts (methanol, ethyl acetate, and chloroform) (model; QP 2010 plus, Shimadzu, Japan). At a rate of 2°/min, the oven temperature rose from 75 to 312°. Electron impact ionization was used to ionize the material (EI, 70eV). It was adjusted to 280° for the injector and 220°C for the detector. Helium was used as a transportation gas. At 1.21 ml.min-1, the carrier gas flow rate was established. 3 scans/s were used to scan compounds in the 40-60 m/z mass range. In a split injection approach, a Hamilton syringe was used to inject each of the three extracts into the GC-MS. Phytoconstituents were identified by comparing mass spectra and retention time patterns to the program Chemstation[15].
Molecular docking:
Ligand Preparation: Phytocomponents identified from the GC-MS analysis of methanol, ethyl acetate and chloroform extract of A. imbricata leaves. The 3D structure of all the compounds was retrieved from the PubChem database (https://pubchem.ncbi.nlm.nih. gov/) and used in this study. All the compounds were subjected to minimization, the hydrogen atoms were added, followed by a minimization step with the Avogadro software. The ligands were built and energy-minimized using the MMFF94 force field.
Selection oftarget protein: As a receptor molecule for antioxidants, antioxidant target protein was used, the protein was retrieved from Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB) with PDB id: 2he3was found from the literature. The 3D structure of this target protein was retrieved from PDB database (https://www.rcsb.org). The 3D structure of antioxidant target Protein was obtained from PDB database, and its PDB ID is 2he3. For the protein structure, structural waters and ligands were removed[16].
Docking studies: Docking studies for the target protein, PDB ID: 2he3 and phytoconstituents identified from the GC-MS analysis of methanol, ethyl acetate and chloroform leaves extract of A. imbricata were performed using Autodock vina software. The binding compounds were searched 25 times using AutoDock Vina's scoring tool with default settings. Visually analyzing categories with RMSD<2 Å. Docking models regarded receptors rigid while ligands flexible. The lowest-binding energy conformations that replicated key interactions were chosen. The results were also analyzed using Discovery Studio 2021.
Statistical analysis:
The data are presented as the mean±Standard Deviation (SD). Antioxidant experiments were conducted using one-way ANOVA tests to compare the half-maximal Inhibitory Concentration (IC50) values of different solvent extracts. To figure out the IC50 values, Graph Pad Prism 5 was used.
Results and Discussion
Majority of the medicinal plants are known to be associated with antioxidant property[17]. The exhibited property depends on the phytoconstituent composition and concentration of the different phytoconstituents present in the extract. As a result, the antioxidant activity of plant extracts cannot be assessed using a single approach[18]. To explore the various processes responsible for antioxidant activities, solvent extracts of A. imbricata leaves (methanol, ethyl acetate, and chloroform) were tested for antioxidant activities such as TAA, FRAP, DPPH, ABTS and hydrogen peroxide scavenging activity.
An acidic pH resulted in the development of a green phosphate/Mo (V) complex that had a TAA. It measures the overall antioxidant capacity of both water- and fat-soluble antioxidants[19]. TAA is a quantitative measure of the antioxidant capacity of the extracts and it is reported in GAE. The total antioxidant capacity of extracts were shown to be in the following order; methanol extract>ethyl acetate extract>chloroform extract (Table 1). Methanol extracts had the highest overall TAA among the three extracts. The TAA of methanol extract increases steadily as extract concentration increases from 100 to 500 μg/ml. The GAE value was found to be 1.231±0.02, 1.216±0.02 and 1.182±0.01 µg/ml GAE at 500 μg/ml of methanol extract, ethyl acetate extract and chloroform extract respectively. All the three extracts exhibited significant TAA. Earlier study using Garcinia cambogia seed extracts (methanol, ethyl acetate and acetone), the methanol extract showed highest TAA at a concentration of 92.63 µg/ml[20].
| TAA in GAE μg/ml of extract | |||
|---|---|---|---|
| Concentration of extract (µg) | TAA (GAE) | ||
| Methanol | Ethyl acetate | Chloroform | |
| 100 | 1.13±0.01 | 1.14±0.01 | 1.13±0.01 |
| 200 | 1.16±0.01 | 1.17±0.01 | 1.14±0.01 |
| 300 | 1.18±0.01 | 1.18±0.01 | 1.16±0.01 |
| 400 | 1.21±0.02 | 1.20±0.01 | 1.18±0.01 |
| 500 | 1.22±0.02 | 1.21±0.02 | 1.19±0.01 |
Table 1: TAA of Different Solvent Extracts of A. imbricata
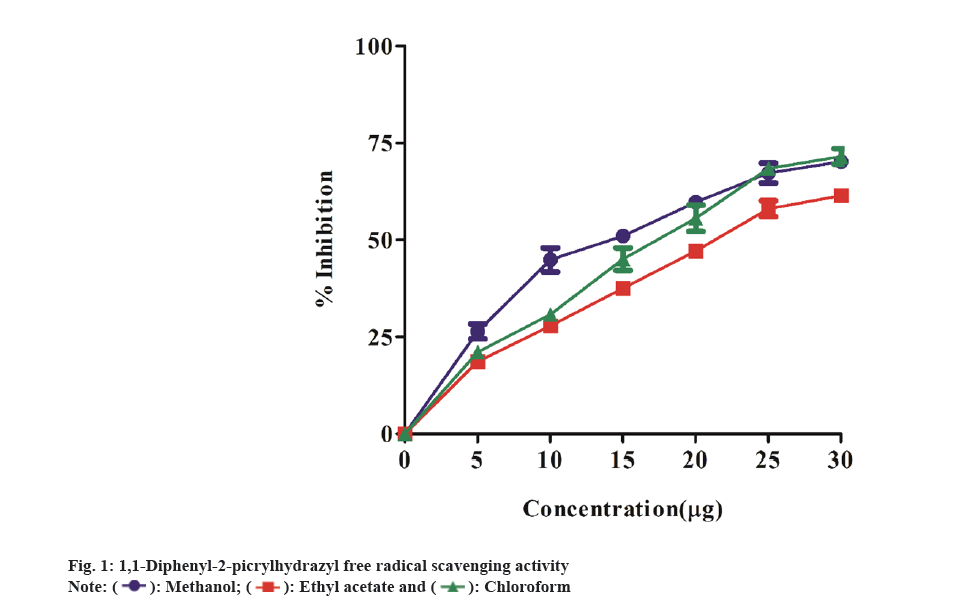
In FRAP assay, antioxidant capacity is measured by the reduction of the ferric ion complex TPTZ. In the presence of antioxidants, the Fe3+/ferricyanide combination can be reduced to Fe2+/ ferrocyanide. As a result, the binding of the ligand to Fe2+ produces a highly vivid navy-blue hue. As a result, the quantity of reduced iron may be measured and associated with the number of antioxidants by detecting the production of Perl's Prussian blue at 700 nm[21]. The results revealed a considerable antioxidant capacity in terms of FRAP measured as GAE of all theextracts, which enhanced steadily with increasing concentrations from 100 to 500 μg/ml of samples. The antioxidant activity of the extracts were observed to decrease in the following order: methanol extract>chloroform extract>ethyl acetate extract (Table 2). Among the extracts, methanol extract showed the highest antioxidant power activity of the three extracts. The GAE value was found to be 14.9±0.03, 17.1±0.04 and 10.1±0.04 µg/ml GAE at 500 μg/ml of methanol extract, ethyl acetate extract, chloroform extract respectively. Likewise, Uddin et al.[22] study have shown the FRAP activity of methanol extract of A. argentea (Roxb) was 150.83±4.26 µmol ascorbic acid/g. The capacity of plant extracts to donate hydrogen atoms was tested by decolorizing a methanol solution of DPPH. DPPH in methanol creates a violet/purple color that fades to yellow in the presence of antioxidants[23]. We investigated the extract's radical scavenging properties by measuring changes in the absorbance of the DPPH at 517 nm. The DPPH radicals were scavenged by all three extracts in a dose-dependent manner. The methanol extract outperformed the other two extracts in terms of radical scavenging activity. The percentage inhibition of free radical (DPPH) scavenging activity at 25 μg/ml of concentration, the methanol extract displayed radical scavenging activity with an IC50 of 7.06 μg/ml, whereas the ethyl acetate and chloroform extracts showed IC50 value of 9.836 μg/ml and 10.03 μg/ml, respectively (fig. 1). Similarly, Mahendra et al.[24] have shown DPPH radical scavenging activity in Argyreia osyrensis Roth plant extract.
| FRAP activity in GAE μg/ml of extract | |||
|---|---|---|---|
| Concentration of extract (µg) | FRAP (GAE) | ||
| Methanol | Ethyl acetate | Chloroform | |
| 100 | 2.1±0.02 | 1.12±0.03 | 1.0±0.02 |
| 200 | 5.4±0.02 | 3.1±0.03 | 1.9±0.02 |
| 300 | 8.4±0.03 | 4.1±0.03 | 5.2±0.02 |
| 400 | 10.1±0.03 | 5.1±0.04 | 8.5±0.04 |
| 500 | 14.9±0.03 | 7.1±0.04 | 10.1±0.04 |
Table 2: FRAP Assay of Different Solvent Extracts of A. imbricata
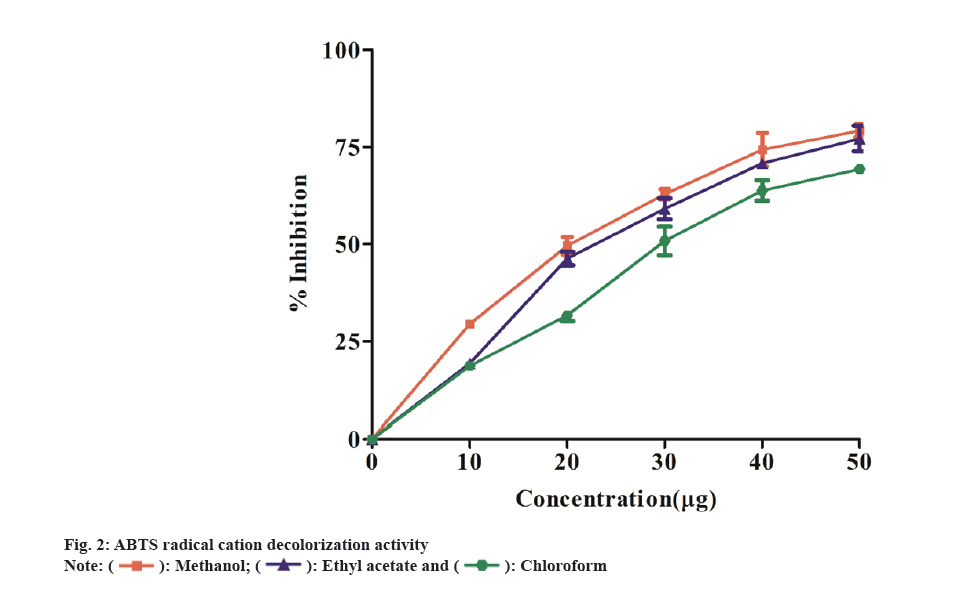
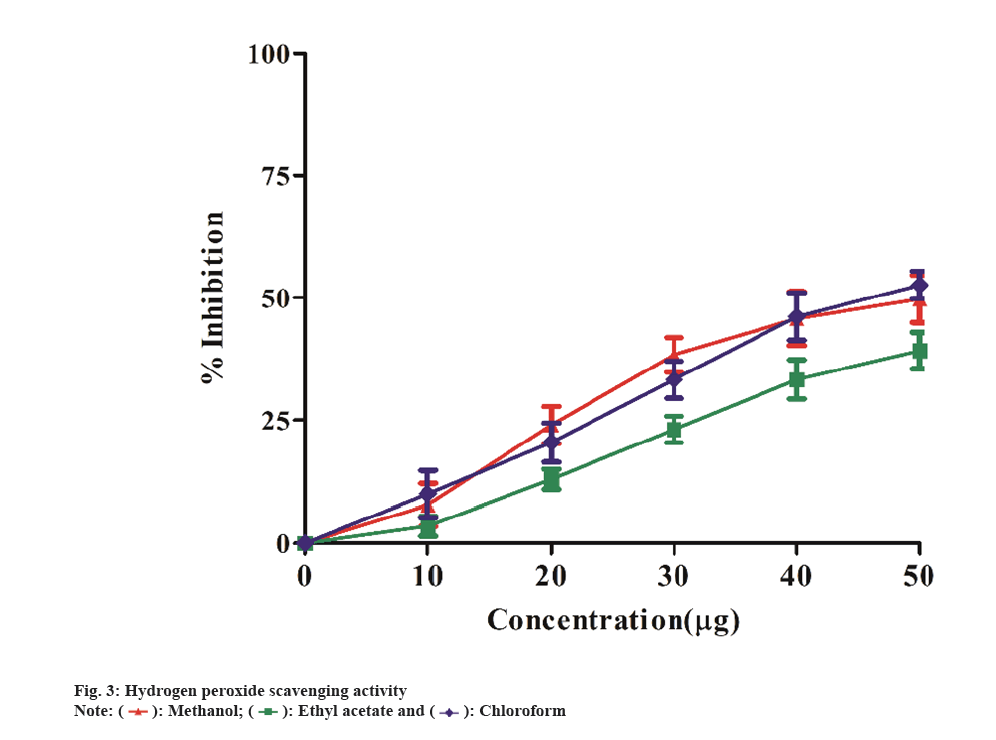
In ABTS method, antioxidants that are capable of donating electrons can convert the blue-green ABTS radical solution into a colorless form. All extracts scavenged the ABTS radical in a concentration-dependent manner (10-50 µg/ml) (fig. 2). The methanol extract exhibited more activity as a radical scavenger than the other two extracts. The methanol extract have shown to scavenge ABTS with an IC50 value of 13.91 μg/ml, while the ethyl acetate and chloroform extracts showed IC50 of 16.52 μg/ml and 18.60 μg/ml, respectively. Similarly, Kekuda et al.[25], showed the maximum ABTS scavenging activity in methanol extract of Argyreia cuneata (willd.) Ker Gawl plant. The leaves methanol extract was more effective in scavenging ABTS radicals with IC50 value of 9.34 μg/ml followed by flower methanol extract (IC50 value 14.98 μg/ml) and stem extract (IC50 value 22.47 μg/ml). The plant extracts ability to scavenge H2O2 is measured spectrophotometrically by the disappearance of H2O2 at 230 nm. H2O2 is found in low concentrations in the natural environment. It decomposes fast into oxygen (O2) and water (H2O), producing hydroxyl radicals (OH), which can cause lipid peroxidation and DNA damage[26]. Fig. 3 depicts the hydrogen peroxide radical scavenging capabilities of the extracts. The methanol extract showed highest hydrogen peroxide radical scavenging activity (percent) with IC50 19.71 μg/ml value compared to the other two extracts, ethyl acetate having IC50 value 25.10 μg/ml and chloroform having IC50 value of 22.36 μg/ml. Likewise, Dintakurthi et al.[27] exhibited H2O2 scavenging activity of Argyreia pilosa Wight and Arn. (Whole Plant). In this study, the methanol and ethyl acetate extracts were found to have the highest percentage of radical scavenging activity.
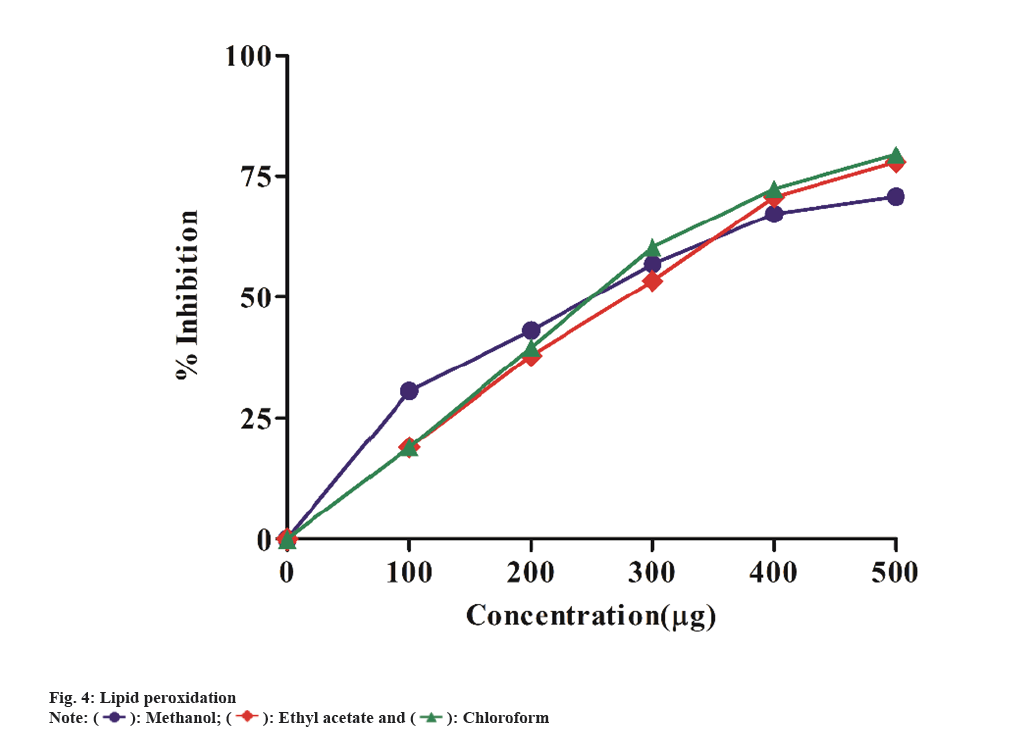
Lipid peroxidation is a chain reaction that causes membrane damage by peroxidizing lipid molecules, particularly polyunsaturated fatty acids. The initial contact produces a second radical, which may react with another macromolecule, resulting in cell malfunction[28]. Lipid peroxidation has the potential to be damaging due to its uncontrolled and self-enhancing nature. This results in the disruption of biomembranes, lipids and other essential cell components. When cells are subjected to oxidative stress, Malonaldehyde (MDA), an end product of lipid peroxidation, increases. Fig. 4 depicts the inhibitory action of extracts in methanol, ethyl acetate and chloroform was determined at concentrations of 100-500 μg/ml. The IC50 values for methanol, ethyl acetate and chloroform extracts were found to be 128.4 μg/ml, 189.4 μg/ml, and 182.6 μg/ml, respectively. The methanol extract significantly showed inhibition of lipid perioxidation when compared to other two extracts. Similar study by Shreedhara et al.[29] exhibited the lipid peroxidation inhibition activity of Argyreia nervosa chloroform extract.
The analysis of the phytoconstituents in the extracts was analyzed by GC-MS method[30]. The potential active compounds responsible for antioxidant activity in A. imbricata leaves extracts was identified using GC-MS analysis. Table 3-Table 5 shows the retention period and peak area percentage of several bioactive principles (fig. 5). The GC-MS analysis of the extracts revealed 26, 46 and 23 different molecules in methanol, ethyl acetate, and chloroform extracts respectively. The identified phytoconstituents belong to alkane, alcohols and ketones group of secondary metabolites. Likewise, Bharati et al.[31] studied the GC-MS analysis of Argyreia Nervosa Burm. F plant. They obtained 26 phytoconstituents belonging to different group of secondary metabolites.
| S No. | Compound Name | Molecular weight | Molecular formula | Retention time | Peak area % |
|---|---|---|---|---|---|
| 1 | 5-O-Methyl-d-gluconic acid dimethyl amide | 237.25 | C9H19NO6 | 8.363 | 3.08 |
| 2 | 1-Octadecyne | 250.5 | C18H34 | 14.936 | 0.65 |
| 3 | 1-Heptadecanol | 256.5 | C17H36O | 17.354 | 0.81 |
| 4 | Phytol | 296.5 | C20H40O | 17.641 | 1.52 |
| 5 | Octasiloxane | 336.68 | O7Si8 | 18.825 | 1.53 |
| 6 | Bis[di(trimethylsiloxy)phenylsiloxy] trimethylsiloxyphenylsiloxane |
793.5 | C33H60O7Si8 | 19.626 | 2.13 |
| 7 | Heptasiloxane | 292.59 | O6Si7 | 19.979 | 2.09 |
| 8 | Cyclononasiloxane | 414.9 | H18O9Si9 | 21.053 | 2.77 |
| 9 | Cyclodecasiloxane | 741.5 | C20H60O10Si10 | 23 | 3.14 |
| 10 | 2,6,10,14,18,22-Tetracosahexaene | 326.6 | C24H38 | 23.24 | 0.99 |
| 11 | 1-Tetracosanol | 354.7 | C24H50O | 23.693 | 1.77 |
| 12 | Heptasiloxane | 292.59 | O6Si7 | 23.864 | 2.66 |
| 13 | Silicic acid | 192.23 | H8O8Sis | 24.527 | 1.33 |
| 14 | Tetracosamethyl-cyclododecasiloxane | 889.8 | C24H72O12Si12 | 24.719 | 2.17 |
| 15 | 1-Triacontanol | 438.8 | C30H62O | 25.083 | 2.29 |
| 16 | Stigmasterol | 412.7 | C29H48O | 27.035 | 3.08 |
| 17 | Cholest-5-en-3-ol | 386.7 | C27H46O | 27.664 | 1.4 |
| 18 | Alpha -Amyrin | 426.7 | C30H50O | 28.271 | 2.46 |
| 19 | 13,14-Epoxyursan-3-ol | 470.7 | C31H50O3 | 28.619 | 5.37 |
| 20 | Lupeol | 426.7 | C30H50O | 28.856 | 2.3 |
| 21 | D:B-Friedo-B':A'-neogammacer-5-en-3-ol | 426.7 | C30H50O | 29.006 | 1.57 |
| 22 | 18,19-Secolupan-3-ol | 430.7 | C30H54O | 29.544 | 5.33 |
| 23 | Tetracosamethyl-cyclododecasiloxane | 889.8 | C24H72O12Si12 | 29.917 | 4.15 |
| 24 | D:A-Friedooleanan-3-ol, | 428.7 | C30H52O | 30.467 | 1.97 |
| 25 | Friedelan-3-one | 426.7 | C30H50O | 30.861 | 17.06 |
| 26 | Tetrapentacontane | 759.4 | C54H11O | 32.052 | 5.39 |
Table 3: Phytocompounds Identified by GC-MS Analysis In Methanol Extract of A. Imbricate
| S No. | Compound name | Molecular weight | Molecular formula | Retention time | Peak area % |
|---|---|---|---|---|---|
| 1 | Isoamyl acetate | 130.18 | C7H14O2 | 3.186 | 0.47 |
| 2 | 5-O-Methyl-D-GluconicAcidDimethylamide | 237.25 | C9H19NO6 | 8.318 | 2.58 |
| 3 | Heneicosane | 296.6 | C21H44 | 11.094 | 0.5 |
| 4 | Heneicosane | 296.6 | C21H44 | 13.561 | 0.62 |
| 5 | 2,5,5,8a-Tetramethyl-4-methylene-6,7,8,8a-tetrahydro-4H,5H-chromen-4a-yl hydroperoxide | 238.32 | C14H22O3 | 14.63 | 0.38 |
| 6 | 3,7,11,15-Tetramethyl-2-hexadecen-1-ol | 296.5 | C20H40O | 14.947 | 1.62 |
| 7 | Tetrapentacontane | 759.4 | C54H110 | 15.41 | 0.76 |
| 8 | Eicosane | 282.5 | C20H42 | 17.366 | 0.9 |
| 9 | 1-Nonadecene | 266.5 | C19H38 | 17.649 | 1.53 |
| 10 | Phytol | 296.5 | C20H40O | 18.959 | 2.97 |
| 11 | Oxirane | 44.05 | C2H4O | 19.583 | 0.41 |
| 12 | Dotriacontane | 450.9 | C32H66 | 20.745 | 0.48 |
| 13 | Dotriacontane | 450.9 | C32H66 | 21.244 | 0.74 |
| 14 | Dotriacontane | 450.9 | C32H66 | 21.498 | 0.65 |
| 15 | Tetrapentacontane | 759.4 | C54H110 | 22.223 | 0.54 |
| 16 | Dotriacontane | 450.9 | C32H66 | 23.247 | 0.58 |
| 17 | Dotriacontane | 450.9 | C32H66 | 23.596 | 0.71 |
| 18 | Squalene | 410.7 | C30H50 | 23.693 | 1.02 |
| 19 | Tetrapentacontane | 759.4 | C54H110 | 23.994 | 1.55 |
| 20 | 1-Pentacosanol | 368.7 | C25H52O | 24.13 | 1.08 |
| 21 | 1,6,10,14,18,22-Tetracosahexaen-3-ol | 426.7 | C30H50O | 24.265 | 3.63 |
| 22 | Triacontane | 422.8 | C30H62 | 24.399 | 2.03 |
| 23 | Tetrapentacontane | 759.4 | C54H110 | 24.62 | 1.65 |
| 24 | Tetrapentacontane | 759.4 | C54H110 | 24.951 | 1.66 |
| 25 | 2-Octadecyl-propane-1,3-diol | 328.6 | C21H44O2 | 25.088 | 1.72 |
| 26 | Tetrapentacontane | 759.4 | C54H110 | 25.599 | 2.03 |
| 27 | 1-Triacontanol | 438.8 | C30H62O | 25.918 | 1.01 |
| 28 | Vitamin E | 430.7 | C29H50O2 | 26.24 | 0.68 |
| 29 | 1-Pentacosanol | 368.7 | C25H52O | 26.623 | 0.85 |
| 30 | Eicosanoic acid | 312.5 | C20H40O2 | 26.743 | 1.55 |
| 31 | Tetrapentacontane | 759.4 | C54H110 | 26.845 | 2.24 |
| 32 | Ergost-5-en-3-ol | 400.7 | C28H48O | 27.046 | 0.55 |
| 33 | 1-Pentacosanol | 368.7 | C25H52O | 27.681 | 0.44 |
| 34 | Stigmasterol | 412.7 | C29H48O | 27.933 | 1.74 |
| 35 | Gamma-Sitosterol | 432.7 | C29H52O2 | 28.291 | 4.68 |
| 36 | 13-Methyl-Z-14-nonacosene | 420.8 | C30H60 | 28.637 | 0.49 |
| 37 | 4,4,6a,6b,8a,11,12,14b-Octamethyl-1,4,4a,5,6,6a,6b,7,8,8a,9,10,11,12,12a,14,14a,14b-octadecahydro-2H-picen-3-one | 424.7 | C30H48O | 28.877 | 3.61 |
| 38 | 9,19-Cyclo-9.beta.-lanostane-3.beta.,25-diol | 444.7 | C30H52O2 | 29.029 | 1.74 |
| 39 | Lup-20(29)-en-3-ol | 426.7 | C30H50O | 29.229 | 16.77 |
| 40 | B-Friedo-B':A'-neogammacer-5-en-3-ol | 426.7 | C30H50O | 29.565 | 6.81 |
| 41 | 3,3,7,11-Tetramethyltricyclo[5.4.0.0(4,11)]undecan-1-ol | 222.37 | C15H26O | 29.848 | 0.3 |
| 42 | 1,4-Dimethyl-8-isopropylidenetricyclo[5.3.0.0(4,10)]decane | 204.35 | C15H24 | 30.09 | 0.4 |
| 43 | Naphthalene | 128.169 | C10H8 | 30.884 | 2.3 |
| 44 | 9,10-Dihydrodeoxynivalenol | 298.33 | C15H22O6 | 31.159 | 0.38 |
| 45 | D:A-Friedooleanan-3-ol | 428.7 | C30H52O | 32.061 | 10 |
| 46 | Friedelan-3-one | 426.7 | C30H50O | 33.434 | 7.09 |
Table 4: Phytocompounds Identified by Gc-Ms Analysis in Ethyl Acetate Extract of A. Imbricata
| S No. | Compound Name | Molecular weight | Molecular formula | Retention time | Peak area % |
|---|---|---|---|---|---|
| 1 | Bacteriochlorophyll-c-stearyl | 841.5 | C52H72MgN4O42 | 18.024 | 1.21 |
| 2 | Tetrapentacontane,1,54-dibromo- | 917.2 | C54H108Br2 | 20.67 | 4.06 |
| 3 | 2,6,10,14,18,22-Tetracosahexaene, 2,6,10,15,19,23-hexamethyl-, (all-E)- | 410.7 | C30H50 | 24.016 | 8.6 |
| 4 | Tetrapentacontane | 759.4 | C54H110 | 24.432 | 1.59 |
| 5 | 2,2-Dimethyl-3-(3,7,16,20-tetramethyl-heneicosa-3,7,11,15,19-pentaenyl)-oxirane | 412.7 | C29H48O | 24.816 | 2.77 |
| 6 | Tetrapentacontane | 759.4 | C54H110 | 26.065 | 3.4 |
| 7 | 1-Triacontanol | 438.8 | C30H62O | 26.181 | 2.16 |
| 8 | Vitamin E | 430.7 | C29H50O2 | 26.78 | 1.01 |
| 9 | Tetrapentacontane | 759.4 | C54H110 | 27.073 | 0.44 |
| 10 | Tetracontane-1,40-diol | 595.1 | C40H82O2 | 27.687 | 0.52 |
| 11 | Tetrapentacontane | 759.4 | C54H110 | 28.261 | 9.2 |
| 12 | 1-Triacontanol | 438.8 | C30H62O | 28.464 | 1.43 |
| 13 | Stigmasterol | 412.7 | C29H48O | 28.566 | 3.1 |
| 14 | Gamma -Sitosterol | 414.7 | C29H50O | 29.374 | 5.25 |
| 15 | Hexacontane | 843.6 | C60H122 | 30.116 | 0.66 |
| 16 | 3.alpha.,7.beta.-Dihydroxy-5.beta.,6.beta.-epoxycholestane | 418.7 | C27H46O3 | 30.842 | 2.72 |
| 17 | Lup-20(29)-en-3-ol | 468.8 | C32H52O2 | 31.053 | 12.63 |
| 18 | D:B-Friedo-B':A'-neogammacer-5-en-3-ol | 426.7 | C30H50O | 31.334 | 5.58 |
| 19 | Tetrapentacontane | 759.4 | C54H110 | 31.694 | 9.41 |
| 20 | Tetracontane-1,40-diol | 595.1 | C40H82O2 | 32.011 | 1.58 |
| 21 | 2-Propenoic acid | 164.16 | C9H8O3 | 32.9 | 1.35 |
| 22 | D:A-Friedooleanan-3-ol, (3.alpha.)- | 428.7 | C30H52O | 33.25 | 11.53 |
| 23 | Friedelan-3-one | 426.7 | C30H50O | 33.415 | 7.21 |
Table 5: Phytocompounds Identified by GC-MS Analysis in Chloroform Extract of A. imbricate
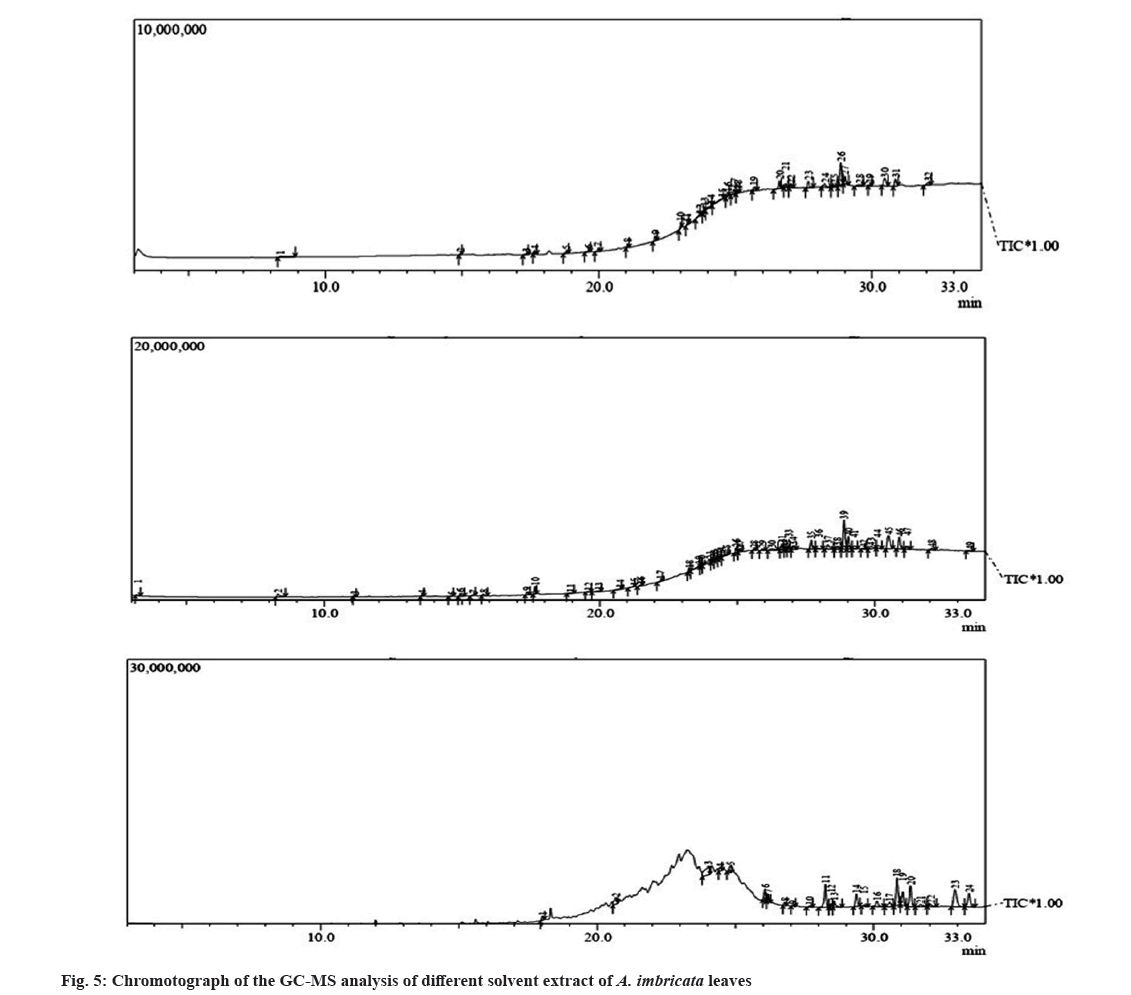
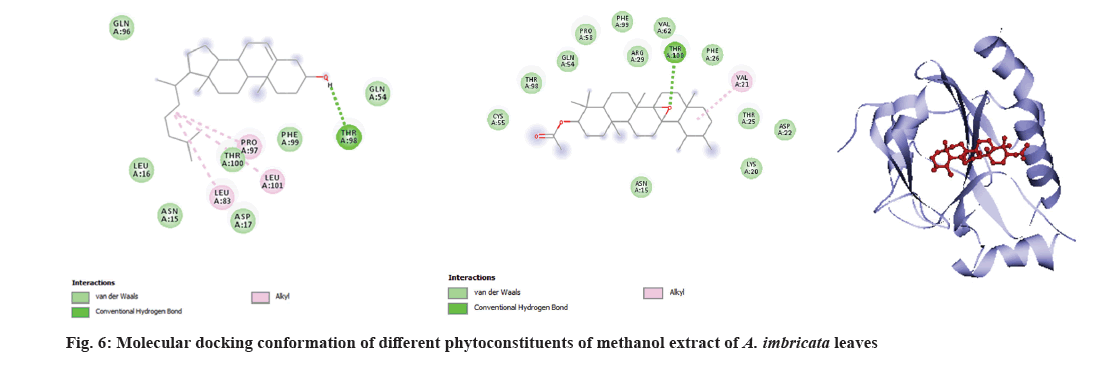
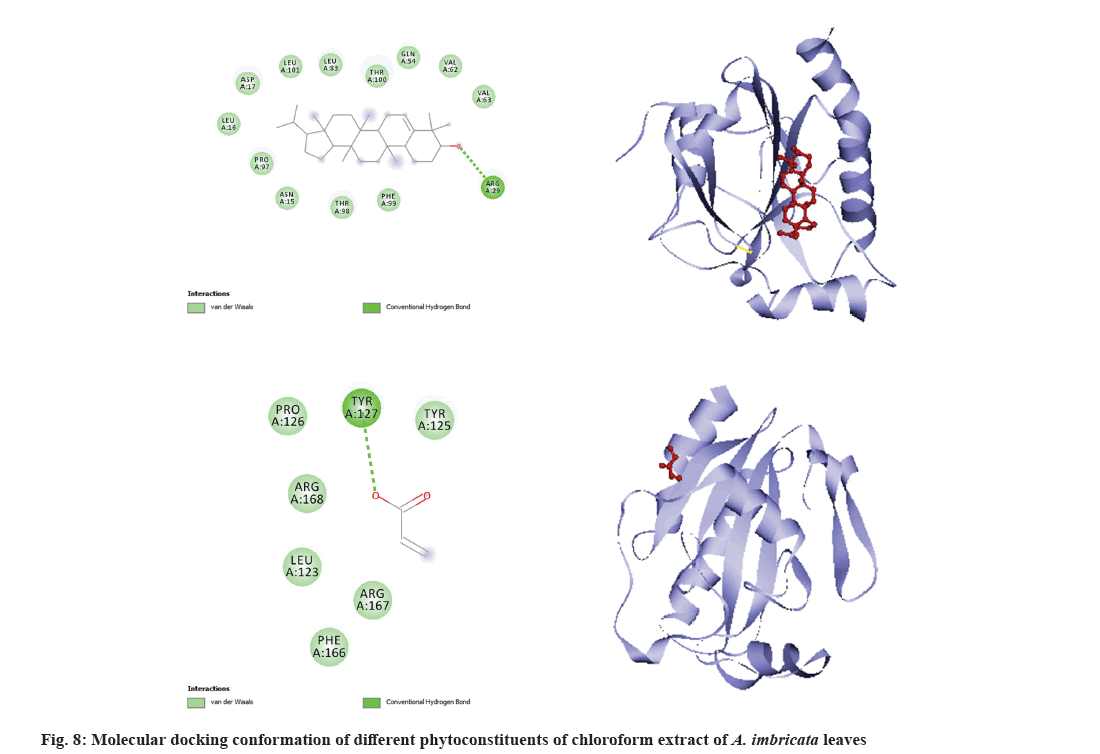
Molecular docking is a very important tool in structural molecular biology, computer-aided drug designing and to analyze the interaction of phytoconstituents with target molecule. The goal of ligand-protein docking is to figure out how a ligand and a protein that has a known 3D structure will interact with each other with mode of binding with the active site of the molecule[15]. Docking analysis was performed using specific pharmacological targets such as antioxidant target protein (PDB id: 2he3) in an attempt to justify the antioxidant activity for the phytoconstituents of all three extracts from GC-MS data. The docking results of antioxidant target protein revealed that the compounds Cholest-5-en-3-ol and 13,14-Epoxyursan-3-ol of methanol extract have a significant binding mode with docking scores of -6.3 and -8.6 Kcal/mol. The compound Cholest-5-en-3-ol formed hydrogen bonding with THR:98 amino acid. 13,14-Epoxyursan-3-ol formed hydrogen bonding with THR:100 amino acid. Further, the interaction was stabilized by alkyl bonds (fig. 6). The 3,7,11,15-Tetramethyl-2-hexadecen-1-ol and 9,10-Dihydrodeoxynivalenol of ethyl acetate extract has substantial binding modes with docking scores of -5 and -5.8 Kcal/mol, respectively (fig. 7). The 3,7,11,15-Tetramethyl-2-hexadecen-1-ol formed hydrogen bonding with LYS:87 amino acid. The 9,10-Dihydrodeoxynivalenol formed hydrogen bonding with LEU:101 and ASN: 12. Further, the interaction was stabilized by alkyl bonds. In chloroform extract D:B-Friedo-B':A'-neogammacer-5-en-3-ol and 2-Propenoic acid indicate substantial binding mechanisms showing -7.7 and -5.3 kcal/mol docking scores. The D:B-Friedo-B':A'-neogammacer-5-en-3-ol formed hydrogen bonding with ARG:29 and 2-Propenoic acid formed hydrogen bonding with TYR:98 (fig. 8). Similar study from Kulkarni et al.[32] showed molecular docking of LC-MS analysis phytoconstituents of petroleum ether, chloroform, ethyl acetate, acetone, methanol and water extracts of Argyreia nervosa (Burm. f) Bojer (Table 6-Table 8).
| Ligand | PubChem id | Energy scores (Kcal/mol) |
|---|---|---|
| Cholest-5-en-3-ol | 304 | -6.5 |
| 13,14-Epoxyursan-3-ol | 605176 | -7.9 |
Table 6: Docking Score (Kcal/mol) of the Phytoconstituents from the Methanol Extract of A. imbricata Leaves Extracts
| Ligand | PubChem id | Energy scores (Kcal/mol) |
|---|---|---|
| 3,7,11,15-Tetramethyl-2-hexadecen-1-ol | 5366244 | -5 |
| 9,10-Dihydrodeoxynivalenol | 573016 | -5.9 |
Table 7: Docking Score (Kcal/Mol) of the Phytoconstituents from the Ethyl Acetate Extract of A. imbricata Leaves
| Ligand | PubChem id | Energy Scores (Kcal/mol) |
|---|---|---|
| D:B-Friedo-B':A'-neogammacer-5-en-3-ol | 623604 | -7.8 |
| 2-Propenoic acid | 637542 | -5.5 |
Table 8: Docking Score (KCAL/MOL) of the Phytoconstituents From The Chloroform Extract of A. Imbricata Leaves
In conclusion, the three different extracts of A. imbricata leaves were analyzed for antioxidant activities by different assay methods, in which all the extracts exhibited significant antioxidant capacities. In addition, all the three extracts inhibited lipid peroxidation activity. GC-MS was used to identify a variety of secondary metabolites from methanol extract, Cholest-5-en-3-ol and 13,14-epoxyursan-3-ol showed highest binding energy with antioxidant target protein by in silico. THR: 98, THR:100, LYS:87, LEU:101 ASN:15, ARG:29 and TYR:127 were the essential amino acid residue involved in the intermolecular interactions. This study focused on the ethnopharmacological phytoconstituents in preventing cell damage by oxidants. Further in vivo studies will give a better approach towards the antioxidant capacities of the molecules identified in the extracts.
Conflict of interests:
The authors declared no conflict of interests.
References
- Pietta P, Simonetti P, Mauri P. Antioxidant activity of selected medicinal plants. J Agri Food Chem 1998;46(11):4487-90.
- Souza NC, de Oliveira JM, Morrone MD, Albanus RD, Amarante MD, Camillo CD, et al. Antioxidant and anti-inflammatory properties of Anacardium occidentale leaf extract. Evid Based Complement Alternat Med 2017;2017:2787308.
[Crossref] [Google Scholar] [PubMed]
- Lee YY, Saba E, Kim M, Rhee MH, Kim HK. Antioxidant and anti-inflammatory properties of raw and processed fruits and vegetables. Biomed Sci Lett 2018;24(3):196-205.
- Czernichow S, Vergnaud AC, Galan P, Arnaud J, Favier A, Faure H, et al. Effects of long-term antioxidant supplementation and association of serum antioxidant concentrations with risk of metabolic syndrome in adults. Am J Clin Nutr 2009;90(2):329-35.
[Crossref] [Google Scholar] [PubMed]
- Saeed N, Khan MR, Shabbir M. Antioxidant activity, total phenolic and total flavonoid contents of whole plant extracts Torilis leptophylla L. BMC Complement Altern Med 2012;12:1-2.
[Crossref] [Google Scholar] [PubMed]
- Thring TS, Hili P, Naughton DP. Antioxidant and potential anti-inflammatory activity of extracts and formulations of white tea, rose, and witch hazel on primary human dermal fibroblast cells. J Inflamm 2011;8(1):1-7.
[Crossref] [Google Scholar] [PubMed]
- Sebastin V, Gopalakrishnan G, Sreejith M, Anoob Kumar KI. Preliminary phytochemical evaluation and spectral characterization of active constituents in the dried extracts of the whole plant Argyreia imbricata (Rox) Sant and Patel. Asian J Pharm Clin Res 2019;12(11):56-60.
- Sebastin V, Gopalakrishnan G, Sreejith M, Kumar KA. In vitro and In vivo antidiabetic evaluation of whole plant extracts of Argyreia imbricata (Roth) Sant. and Patel. Pharmacogn J 2021;13(1):30–6.
- Prieto P, Pineda M, Aguilar M. Spectrophotometric quantitation of antioxidant capacity through the formation of a phosphomolybdenum complex: Specific application to the determination of vitamin E. Anal Biochem 1999;269(2):337-41.
[Crossref] [Google Scholar] [PubMed]
- Benzie IF, Strain JJ. The ferric reducing ability of plasma (FRAP) as a measure of “antioxidant power”: the FRAP assay. Anal Biochem 1996;239(1):70-6.
[Crossref] [Google Scholar] [PubMed]
- Braca A, Sortino C, Politi M, Morelli I, Mendez J. Antioxidant activity of flavonoids from Licania licaniaeflora. J Ethnopharmacol 2002;79(3):379-81.
[Crossref] [Google Scholar] [PubMed]
- Seeram NP, Henning SM, Niu Y, Lee R, Scheuller HS, Heber D. Catechin and caffeine content of green tea dietary supplements and correlation with antioxidant capacity. J Agric Food Chem 2006;54(5):1599-603.
[Crossref] [Google Scholar] [PubMed]
- Ruch RJ, Cheng SJ, Klaunig JE. Prevention of cytotoxicity and inhibition of intercellular communication by antioxidant catechins isolated from Chinese green tea. Carcinogenesis 1989;10(6):1003-8.
[Crossref] [Google Scholar] [PubMed]
- Ohkawa H, Ohishi N, Yagi K. Assay for lipid peroxides in animal tissues by thiobarbituric acid reaction. Anal Biochem 1979;95(2):351-8.
[Crossref] [Google Scholar] [PubMed]
- Huey R, Morris GM, Olson AJ, Goodsell DS, Lindstrom W, Sanner MF, et al. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility
[Crossref] [Google Scholar] [PubMed]
- Kelley LA, Mezulis S, Yates CM, Wass MN, Sternberg MJ. The Phyre2 web portal for protein modeling, prediction and analysis. Nat Protoc 2015;10(6):845-58.
[Crossref] [Google Scholar] [PubMed]
- Salar RK, Seasotiya L. Free radical scavenging activity, phenolic contents and phytochemical evaluation of different extracts of stem bark of Butea monosperma (Lam.) Kuntze. Front Life Sci 2011;5(3-4):107-16.
- Kumar V, Lemos M, Sharma M, Shriram V. Antioxidant and DNA damage protecting activities of Eulophia nuda Lindl. Free Radic Antioxid 2013;3(2):55-60.
- Aliyu AB, Ibrahim MA, Musa AM, Musa AO, Kiplimo JJ, Oyewale AO. Free radical scavenging and total antioxidant capacity of root extracts of Anchomanes difformis Engl (Araceae). Acta Pol Pharm 2013 Jan 1;70(1):115-21.
[Google Scholar] [PubMed]
- Sumachirayu CK, Kusuma CG, Gubbiveeranna V, Bhavana S, Nagaraju S. Angiotensin converting enzyme inhibition, antioxidant activity and phytochemical evaluation of Garcinia cambogia seeds. Int J Res Eng Sci Manag 2020;9(7): 1532-1548.
- Al-Salahi R, Anouar EH, Marzouk M, Taie HA, Abuelizz HA. Screening and evaluation of antioxidant activity of some 1, 2, 4-triazolo [1, 5-a] quinazoline derivatives. Future Med Chem 2018;10(4):379-90.
[Crossref] [Google Scholar] [PubMed]
- Uddin MN. Network pharmacology of arbutin view project association of stromal MMP2 with immune inhibition. J Ethnopharmacol 2014;235:227-42.
- Rahman MM, Islam MB, Biswas M, Khurshid Alam AH. In vitro antioxidant and free radical scavenging activity of different parts of Tabebuia pallida growing in Bangladesh. BMC Res Notes 2015;8(1):1-9.
[Crossref] [Google Scholar] [PubMed]
- Mahendra C, Manasa G, Murali M, Amruthesh KN, Sudarshana MS, Lingaraju DP. Antibacterial and antioxidant properties of Argyreia osyrensis Roth. Ann Phytomed 2016;5(1):110-5.
- Kekuda TP, Bharadwaj NA, Sachin MB, Sahana BK, Priyanka GS. In vitro antimicrobial and antioxidant activity of Argyreia cuneata (Willd.) Ker Gawl.(Convolvulaceae). J Drug Deliv Ther 2018;8(6):22-7.
- Bhatti MZ, Ali A, Ahmad A, Saeed A, Malik SA. Antioxidant and phytochemical analysis of Ranunculus arvensis L. extracts. BMC Res Notes 2015;8:1-8.
[Crossref] [Google Scholar] [PubMed]
- Dintakurthi P, Prasanth DSNBK, Rao AS, Yejella RP. Phytochemistry view project phytochemical, in vitro antioxidant and antibacterial activities of Argyreia pilosa wight and arn. (Whole plant). Int J Pharmacogn 2017;4(4):109-17. [Crossref] [Google Scholar] [PubMed]
- Khan MA, Rahman AA, Islam S, Khandokhar P, Parvin S, Islam MB, et al. A comparative study on the antioxidant activity of methanolic extracts from different parts of Morus alba L.(Moraceae). BMC Res Notes 2013;6:1-9.
[Crossref] [Google Scholar] [PubMed]
- Shreedhara CS, Ram HA, Zanwar SB, Falguni GP. Free radical scavenging activity of aqueous root extract of Argyreia nervosa (Burm. f.) Boj. (Convolvulaceae). J Nat Remedies 2009;9(2):216-23.
- Garg K, Shrivastava B, Bhargava A. GC-MS analysis of methanol and ethyl acetate extract of fruits of Sphaeranthus indicus. J Drug Deliv Ther 2019;9(2):28-30.
- Bharati AJ, Bansal YK. In vitro propagation and GC-MS study of Argyreia nervosa Burm. F.: An endangered ornamental and medicinal plant. Anal Chem Lett 2015;5(6):385-98.
- Kulkarni AA, Kamble AD. Flavonoids from Argyreia nervosa (Burm. f.) bojer: A ready arsenal against pests as well as diabetes. Biol Life Sci Forum 2021;4(1):56.