- *Corresponding Author:
- T. R. Saini
Industrial Pharmacy Research Lab, Department of Pharmacy, Shri G. S. Institute of Technology and Science, Indore-452 003, India
E-mail: tsaini@sgsits.ac.in
| Date of Submission | 23 January 2017 |
| Date of Revision | 23 November 2017 |
| Date of Acceptance | 18 June 2018 |
| Indian J Pharm Sci 2018;80(4):694-701 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
In the present investigation a novel, scientific, simple, and reliable nail safety screening test based on Fourier-transform infrared spectroscopy was developed for the screening of nail-friendly transungual drug permeation enhancers. In this test, nail clippings were treated with the permeation enhancer and their FTIR spectra were recorded and compared with FTIR spectrum of untreated nail clipping. With the help of three spectral parameters namely, peak purity index, peak intensity ratio, and peak-fitting analysis any type of structural alteration brought out by the permeation enhancer in the matrix of nail plate could be determined and nail safety could be readily assessed. The validity of the proposed method was established by studying the effect of eight different types of permeation enhancers, thioglycolic acid, salicylic acid, n-acetyl-l-cysteine, sodium sulfite, resorcinol, sodium hydroxide, oxalic acid, and hydroxypropyl-β-cyclodextrin on the nail clippings. It was observed that thioglycolic acid, salicylic acid, sodium sulfite, resorcinol, sodium hydroxide, oxalic acid, and n-acetyl-l-cysteine severely damaged the nail plate keratin as evidenced by the significantly altered values of peak purity index, peak intensity ratio, and peak-fitting analysis, whereas hydroxylpropyl-β-cyclodextrin was found to be innocuous as the above three spectral parameters remained almost unaltered. This technique could be easily employed for screening nail-friendly drug permeation enhancers from the large pool of available chemical permeation enhancers during the preformulation stage of development of transungual drug delivery systems.
Keywords
FTIR, transungual permeation enhancers, preformulation, peak purity index, peak-fitting analysis, hydroxypropyl-β-cyclodextrin
For years, the treatment of onychomycosis was principally based on oral antifungal drug therapy but due to high incidences of nephrotoxicity and hepatotoxicity of orally administered antifungal drugs [1,2], the oral drug delivery is being gradually replaced by transungual drug delivery through topical application of nail lacquer formulations. Due to selective drug targeting to the nail unit in this therapy, very low incidences of drug side effects are observed because of lower systemic absorption of administered drugs by this route. These transungual formulations are, however, associated with the inherent problem of poor transungual drug permeability due to highly impermeable nature of keratinized nail plate [2]. The research findings have shown that the drug permeation across nail plate can be significantly enhanced in presence of some keratolytic agents, which can disrupt keratin network of nail plate and thus weaken its barrier property resulting into enhancement of drug permeation [3-5]. Keratolytic agents, urea and salicylic acid destabilize hydrogen bonds, while sulfhydryl compounds such as 2-mercaptoethanol cleave the disulphide linkages of nail keratin and enhance permeability of nail plates [6]. As these types of permeation enhancers irreversibly damage the microstructure of nail plate, these cannot be considered as safe and nail-friendly permeation enhancers for increasing the transungual permeability of drugs [7]. The structural alterations might turnout to be permanent in nature since transungual formulations are applied for a long period of time ranging from 6-12 mo [6]. Therefore, screening of transungual permeation enhancers for their nail safety is necessary at preformulation stage of product development. In the present investigations a nail safety test technique for evaluation of permeation enhancers was evolved. The test is based on Fourier-transform infrared (FTIR) spectroscopic analysis of nail plate.
FTIR spectroscopy is a widely used analytical technique for the identification and characterization of secondary structure of proteins. Chemically the nail plate is composed of ~80 % w/w hard α-keratin protein with ~10 % w/w content of cystine [3,8,9]. Quantitative information about the secondary structure of a keratin protein could be obtained by analysing the amide-I band observed in the FTIR spectrum (1700-1600 cm-1) of protein [10,11]. The amide-I band is originated from the C=O stretching vibration of the peptide group, and its frequency depends upon the hydrogen-bonding and coupling along the protein chain and therefore, appearance of this band in the FTIR spectra is dependent on the conformation of the secondary structure of the protein [10].
When permeation enhancers are applied to the nail plate and if these damage the secondary structure of keratin, it would be easily identified by the FTIR spectral analysis and similarly if the permeation enhancer did not alter the secondary structure of nail plate protein it could also be detected and confirmed. Working on this theory, a hypothesis was deduced that any change in the structure of nail plate protein upon application of a permeation enhancer would be predicted by an altered FTIR spectrum. This would certainly confirm that the applied permeation enhancer has adversely affected the chemical integrity of nail plate and therefore it would not be safe or nail-friendly. If the permeation enhancer did not alter or marginally alter the FTIR spectra of nail plate it would indicate that the permeation enhancers is a safe and nail-friendly agent. Thus the proposed FTIR spectroscopic technique could be successfully employed to screen the safety of transungual permeation enhancers at the preformulation stage of product development.
The validity of the above hypothesis was experimentally tested by exposing human nail clippings with different types of permeation enhancers, which were reported to have a good transungual drug permeation enhancement property. The FTIR spectra of nail clippings were recorded before and after treatment with these permeation enhancers and compared for the changes in the secondary structure of nail keratin protein due to their effect. Three spectral screening parameters namely, peak purity index, peak intensity ratio, and peak-fitting analysis were evolved and employed for this study. The proposed technique is a new, innovative, and non-invasive approach to screen the given permeation enhancer as safe or unsafe for transungual drug delivery applications.
Materials and Methods
Analytical grade thioglycolic acid, salicylic acid, n-acetyl-l-cysteine, sodium sulphite, resorcinol, sodium hydroxide, and oxalic acid were purchased from Loba Chemie Pvt. Ltd., India. Hydroxypropyl-β-cyclodextrin (Kleptose®, HPB) was obtained as generous gift from Roquette Corporate (Lestrem Cedex, France). Spectroscopy grade potassium bromide was purchased from Merck, India. All solutions were prepared in purified water (Direct-Q 3R® water purification system, Millipore, India).
Nail clippings
Nail clippings were obtained from healthy human volunteers (male and female, age 25-50 y) using nail clippers. The middle, index, and ring finger nails were used for the study. Nail clippings were cleaned with purified water and wiped with tissue paper. Washed nail clippings were dried in an oven at 45° for 12 h before use [7].
Selection of permeation enhancers
Eight permeation enhancers, which were found to have prominent drug uptake enhancement factor (UEF24) in earlier studies [7] were selected as the model permeation enhancers for the present nail safety screening test (Table 1). The concentrations of permeation enhancers were selected on the basis of previously published reports [3,12].
| Permeation enhancer | Concentration (% w/v) |
|---|---|
| Hydroxypropyl-β-cyclodextrin (HP-β-CD) | 5.00 |
| N-acetyl-l-cysteine | 1.00 |
| Oxalic acid | 5.00 |
| Resorcinol | 5.00 |
| Salicylic acid | 1.00 |
| Sodium hydroxide | 0.08 |
| Sodium sulphite | 5.00 |
| Thioglycolic acid | 5.00 |
Table 1: Transungual Permeation Enhancers Selected for FTIR-based Nail Safety Screening Test
Preparation of permeation enhancer test solutions
Test solutions of permeation enhancers in the desired concentrations as shown in Table 1 were prepared in purified water. All permeation enhancers except salicylic acid formed a clear solution. The solution of salicylic acid was turbid in appearance.
Treatment of nail clippings with permeation enhancer
Nail clippings were placed in a glass vial filled with 1 ml test solution of each permeation enhancer. Simultaneously, a control was run by placing a nail clipping in 1 ml purified water in another glass vial. The glass vials were sealed and incubated at room temperature for 24 h [7]. After that the nail clippings were removed from the respective vial and washed three times with purified water and three times with methanol to remove any traces of permeation enhancers on the surface. The washed nail clippings were then socked in 10 ml purified water for 1 h to further remove the traces of permeation enhancer left. The nail clippings were then wiped with tissue paper and dried for 12 h in an oven at 45°. The dried nail clippings were stored in sealed glass vials at 2-8° until FTIR studies performed.
FTIR analysis
The nail clippings that were exposed to permeation enhancers were rubbed against a metallic nail file. Sufficient amount of nail powder produced was mixed with spectroscopy grade potassium bromide in the ratio of 1:4 and ground further. The mixture was placed in the small sample holder of diffused reflectance spectroscopy assembly of a FTIR spectrophotometer (IR Affiniti-1, Shimadzu, Japan) and the FTIR spectrum was recorded in the range of 400-4000 cm-1.
Peak purity index
The peak purity index (P) signifies the degree and extent of matching of two spectra. It is obtained by the least-squares-fit coefficient calculated for every intensity pair of the two spectra being compared. The purity index, lies between 0 and 1. Zero indicates absolute dissimilarity between two spectra, and 1 indicates that the two spectra are identical [13]. In the present study the peak purity index was determined by comparing the FTIR spectrum of the control nail plate with the spectrum of each permeation enhancer-treated nail plate using IR solution-1.40 software (Shimadzu, Japan). The value of peak purity index was calculated by applying normalization, which automatically corrected the differences between the two spectra. In the process of normalization the baseline of the spectrum was first corrected to zero followed by determining the peak of highest intensity and the scaling factor (f=1/ maximum intensity). The scaling factor, f, was then multiplied with the spectrum and the peak purity index was obtained using the Eqn. 1, P = Σin(si–S)(ri–R)/√Σin(si–S)2Σin(ri–R)2, where, P was the purity index; si was the individual peak intensity from spectrum of permeation enhancer-treated nail clipping; ri, the individual peak intensity from spectrum of control nail clipping; S was the average intensity from spectrum of permeation enhancer-treated nail clipping; R was the average intensity from spectrum of control nail clipping and n was the number of peaks.
Peak intensity ratio
Close observation of the FTIR spectra of permeation enhancer-treated nail clippings demonstrated that the entire region of spectra was affected by all the permeation enhancers except HP-β-CD. To know the effect of permeation enhancer more precisely the parameter of peak intensity ratio was evolved through which the effect of each permeation enhancer on five major characteristic spectral peaks of nail protein in the FTIR spectra of control nail clippings was observed. The peak intensity ratio of each of the five peaks was calculated using Eqn. 2: peak intensity ratio = Itreated/ Icontrol, where, Itreated is the intensity of a characteristic spectral peak in FTIR spectrum of permeation enhancer treated nail clippings; and Icontrol is the intensity of a characteristic spectral peak in FTIR spectrum of control nail clippings.
Peak-fitting analysis
As the integrity of nail plate is dependent on the stability of secondary structure of nail plate protein, therefore, determination of secondary structure of nail plate protein could give more accurate information about the exact effect of permeation enhancers on integrity of nail plate [11]. In the present study, changes in the secondary structure of nail keratin protein produced by permeation enhancers was measured by the peakfitting analysis. The region of amide-I band (1750- 1550 cm-1) of the FTIR spectrum of control nail clippings and permeation enhancer-treated nail clippings was selected for the analysis as this is specifically contributed by the secondary structure of protein and produced by additive effect of different bands assigned to the α-helix structures, β-sheet component, and turns of secondary protein structure [14]. To resolve these bands the peak-fitting analysis was done using Peakfit-4.12 software (Systat software Inc., CA, USA) as a linear combination of Gaussian components. First, the flat baseline of the amide-I band at 1750-1550 cm-1 was obtained by adjusting the subtraction factor, then the Savitsky-Golay smoothing was applied (3rd grade polynomial, 5 smoothing points) to the spectrum to obtain a second-derivative spectrum. During the peakfitting, a number of Gaussian bandwidths and initial values of their peak positions were obtained from the second derivative spectrum. Taking the above values as initial parameters, the curve fitting of the amide-I band was done by leaving the parameters free to adjust iteratively except the peak wave-numbers, which were allowed to vary only within a range of ± 2 cm−1. Finally, the area corresponding to each secondary structure was calculated and expressed as the percent area of the amide-I band [11].
Results and Discussion
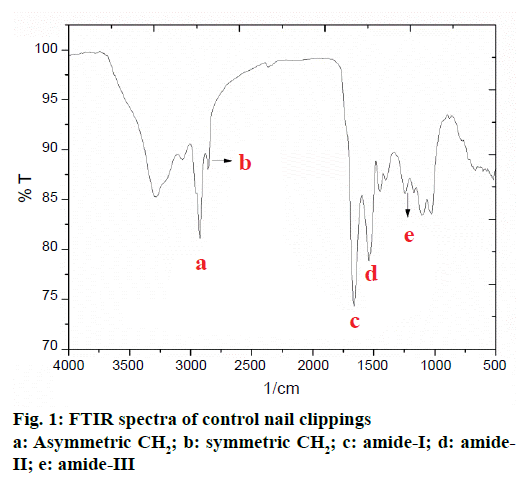
The FTIR spectrum of human nail plate is presented in Figure 1. For the purpose of interpretation, the FTIR spectrum of human nail clipping was divided into three basic regions; first region falling between 4000- 3000 cm-1 consisted of bands of water and hydroxyl group present in the nail plate and signifies the nature of hydrogen bonding, the second region located between 3000-1500 cm-1 described the bands for functional groups. The third region lying between 1500-200 cm-1 indicated peaks due to some salts and organic molecules present in the nail. In the first two regions the major FTIR absorption peaks of chemical contents of nail plate have appeared. In the second and third regions the amide-I, II, and III bands are present (Table 2). Majority of these bands belong to protein structure of nail [15]. The amide-I band (1643.55 cm-1) originated mainly from the C=O stretching vibrations of the amide groups weakly coupled to the in-plane N-H bending and C-N stretching modes. The exact frequency of amide-I band depends on the nature of hydrogen bonding formed due to the C=O and N-H groups, and any change in secondary structure of protein would reflect into change in the nature of hydrogen bonding. Therefore, the frequency of the amide-I band has been used to identify the alterations in secondary structure of proteins and polypeptides in biological samples [14].
| Peak position wave number (cm-1; mean ± SD) |
Intensity (%; mean ± SD) |
Peak name assigned[14,18] |
|---|---|---|
| 1643.35 ± 0.5 | 76.79 ± 2.1 | Amide-I |
| 1546.91 ± 0.7 | 78.57 ± 1.2 | Amide-II |
| 1242.16 ± 0.3 | 85.43 ± 2.4 | Amide-III |
| 2924.09 ± 0.4 | 80.61 ± 2.6 | Asymmetric CH2 |
| 2852.72 ± 0.3 | 87.33 ± 3.5 | Symmetric CH2 |
Table 2: Identification of Peak Positions in the FTIR Spectrum of Control Nail Clippings
The amide-II band (1546.91 cm-1) appeared with weaker intensity due to the N–H bending and the C–N stretching vibrations of protein. The amide-III band (1242.16 cm−1) is due to the N–H bending, C–C and C–N stretching vibrations. The amide-III band contributes to the vibrations due to the backbone and side-chain of protein molecule [15]. Presence of combination of amide-I and amide-III bands indicated that the keratin protein present in nail plates are mostly in the α-helix conformation [16,17].
The presence of lipid ester carbonyl peak at 1745 cm-1 and the lipid methylene stretching vibrations at 2924.09 cm-1 (asymmetric CH2) and 2852.72 cm-1 (symmetric CH2) are attributed to unordered lipid acyl chains (Table 2) and this confirms the presence of lipid in the nail plate [18-20].
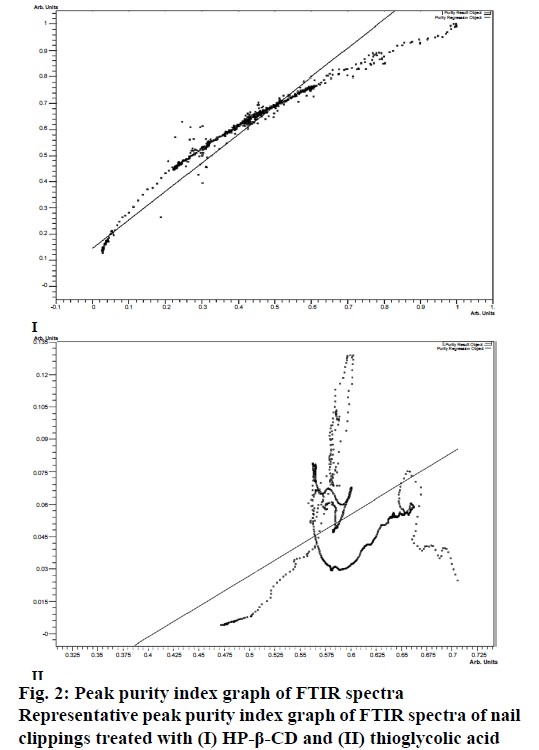
To evaluate the effect of permeation enhancers on the predominantly protein-based structure; the nail plates were immersed in the aqueous solutions of different permeation enhancers for 24 h (Table 1). The FTIR analysis of these nail plates was then performed and FTIR spectra of control and permeation enhancerstreated nail plates were compared. The purity index (Figure 2) of the spectra was calculated using Eqn. 1. As shown in Table 3, the highest value of peak purity index was obtained from the nail clippings exposed to HP-β- CD (0.976 ± 0.07) and the least value in the nail clippings treated with thioglycolic acid (0.751 ± 0.04). The values of peak purity index confirmed that all the seven permeation enhancers except HP-β-CD significantly altered the FTIR spectra of the nail clippings due to their effect on the keratin structure of nail clippings. On the basis of peak purity index the structural alteration effect of different permeation enhancers was in the order of thioglycolic acid˃salicylic acid˃nacetyl- l-cysteine˃resorcinol=sodium sulphite˃sodium hydroxide ˃oxalic acid˃HP-β-CD.
| Permeation enhancer | Peak purity index (mean ± SD) |
|---|---|
| HP-β-CD | 0.976 ± 0.07 |
| N-acetyl-l-cysteine | 0.893 ± 0.06 |
| Oxalic acid | 0.923 ± 0.03 |
| Resorcinol | 0.905 ± 0.02 |
| Salicylic acid | 0.769 ± 0.01 |
| Sodium hydroxide | 0.919 ± 0.06 |
| Sodium sulphite | 0.905 ± 0.06 |
| Thioglycolic acid | 0.751 ± 0.04 |
Table 3: Peak Purity Index of FTIR Spectrum of Nail Plates Treated with Different Permeation Enhancers
The peak intensity ratio determined by Eqn. 2 signifies the extent of nail protein alteration due to effect of permeation enhancers as these peaks are very specifically appeared in FTIR spectra of nail clippings and collectively confirm the presence of protein in nail plate. If permeation enhancer alters the structure of nail protein it would reduce its spectral intensity. As the peak intensity is a quantitative parameter and its magnitude is resultant of the concentration of a functional group appeared in a sample, therefore, this parameter was used for quantitative measurement of change in nail protein composition. If the intensity of a characteristic peak in permeation enhancer-treated nail clipping is reduced in comparison with the intensity of same peak in control nail clipping it would lower the intensity ratio, which in turn would signify the proportional reduction in protein content of nail plate. The peak intensity ratio of each of the five characteristic spectral peaks obtained from permeation enhancers-treated nail clippings was calculated and the results are recorded in Table 4.
| Permeation enhancer | Peak intensity ratio | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Characteristic spectral peak | ||||||||||
| Amide-I | Amide-II | Amide-III | Asymmetric CH2 | Symmetric CH2 | ||||||
| Peak intensity | Peak intensity ratio | Peak intensity | Peak intensity ratio | Peak intensity | Peak intensity ratio | Peak intensity | Peak intensity ratio | Peak intensity | Peak intensity ratio | |
| Control | 76.79 | NA | 78.57 | NA | 85.43 | NA | 80.61 | NA | 87.33 | NA |
| HP-β-CD | 75.25 | 0.98 | 74.64 | 0.95 | 76.03 | 0.89 | 78.19 | 0.97 | 85.58 | 0.98 |
| N-acetyl-l-cysteine | 39.97 | 0.52 | 41.30 | 0.53 | 42.39 | 0.50 | 46.58 | 0.58 | 50.05 | 0.57 |
| Oxalic acid | 25.38 | 0.33 | 27.67 | 0.35 | 32.15 | 0.38 | 35.68 | 0.44 | 38.72 | 0.44 |
| Resorcinol | 19.63 | 0.26 | 21.85 | 0.28 | 25.77 | 0.30 | 30.84 | 0.38 | 34.72 | 0.40 |
| Salicylic acid | 28.01 | 0.36 | 30.35 | 0.39 | 29.35 | 0.34 | 31.38 | 0.39 | 31.96 | 0.37 |
| Sodium hydroxide | 48.63 | 0.63 | 51.86 | 0.66 | 57.42 | 0.67 | 58.91 | 0.73 | 65.13 | 0.75 |
| Sodium sulfite | 23.22 | 0.30 | 24.26 | 0.31 | 28.59 | 0.33 | 31.67 | 0.39 | 34.61 | 0.40 |
| Thioglycolic acid | 25.00 | 0.33 | 25.79 | 0.33 | 25.10 | 0.29 | 27.11 | 0.34 | * | * |
Table 4: Peak Intensity Ratio of Characteristic Spectral Peaks in the FTIR Spectra of Permeation Enhancer-Treated Nail Clippings
The peak intensity ratio clearly depicted the extent of effect of permeation enhancer on nail structure. Maximum structural alteration was produced by thioglycolic acid followed by salicylic acid, sodium sulphite, oxalic acid, resorcinol, n-acetyl-l-cysteine, and sodium hydroxide, respectively as in the nail clippings treated by these permeation enhancers, the peak intensity ratio was significantly reduced. The effect of HP-β-CD was negligible as peak intensity ratio value was close to 1. On the basis of peak intensity ratio the structural alteration effect of different permeation enhancers was found to be in the order of thioglycolic acid˃resorcinol˃sodium sulphite˃oxalic acid˃salicylic acid˃n-acetyl-l-cysteine˃sodium hydroxide˃HP-β- CD.
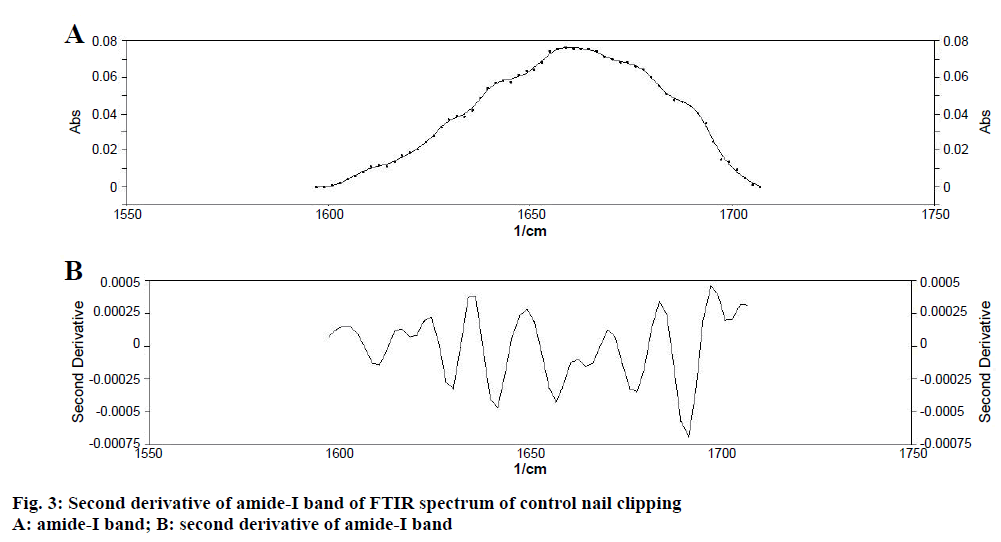
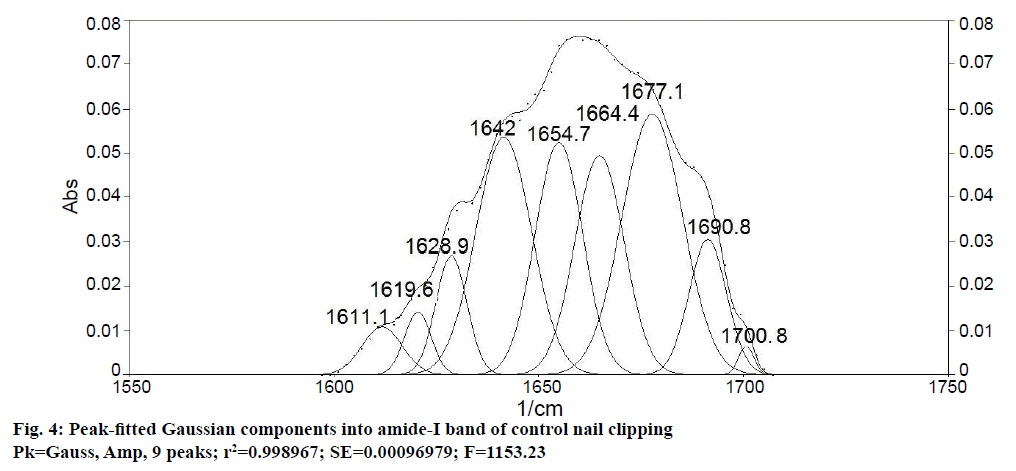
Determination of secondary structure of nail plate protein could give confirmatory information about the effect of permeation enhancers on integrity of nail plate, as it is closely related with the stability of secondary structure of protein [11]. To obtain quantitative information about secondary structures of nail plate protein, peak-fitting analysis of the amide-I absorption band of untreated (control) nail clippings was carried out. It revealed nine hidden peaks in the amide-I band. To establish the validity of the peak-fitting analysis, correlation coefficient between peak positions of all nine peaks in the second derivative spectrum of amide-I band (Figure 3) and their peak-fitted spectrum (Gaussian components) was calculated (Figure 4). An excellent agreement was observed (r2=0.999) for the band positions of the best-fit Gaussian components with those determined from the second derivative spectrum. These results supported the reliability of the peak-fitting procedure and the predicted secondary structure of the nail plate protein. All the nine peaks obtained in peak-fitting analysis corresponded to the specific secondary structure of nail plate protein. All identical components were clubbed to obtain the percent composition of nail plate protein. By this calculation the secondary structure of the control nail clipping was found to be comprised of 19.08 % α-helix (peak no. 5), 27.54 % β-sheets (peak no. 2, 7-9), 37.24 % β-turns (peak no. 1, 3, and 4), and 16.14 % random coils (peak no. 6) as shown in Table 5.
| Peak No. | Component band position from second derivative of amide-I band (cm−1) |
Secondary structure of nail protein (Best-fit Gaussian components of amide-I band) |
Assigned name of components[10,15] | |
|---|---|---|---|---|
| Band position (cm-1) | Relative area (%) |
|||
| 1 | 1703.1 ± 0.5 | 1700.8 ± 0.2 | 1.14 ± 0.08 | β-turn |
| 2 | 1691.6 ± 0.8 | 1691.3 ± 0.3 | 12.45 ± 0.12 | β-sheet |
| 3 | 1678.1 ± 0.6 | 1677.1 ± 0.4 | 17.57 ± 0.58 | β-turn |
| 4 | 1666.5 ± 0.5 | 1664.4 ± 0.7 | 18.52 ± 1.21 | β-turn |
| 5 | 1654.9 ± 0.3 | 1654.7 ± 0.4 | 19.08 ± 1.82 | α-helix |
| 6 | 1641.4 ± 0.4 | 1642.0 ± 0.3 | 16.14 ± 1.48 | Random coil |
| 7 | 1629.9 ± 0.2 | 1628.9 ± 0.5 | 9.47 ± 1.07 | β-sheet |
| 8 | 1618.3 ± 0.5 | 1619.6 ± 0.4 | 3.07 ± 0.25 | β-sheet |
| 9 | 1608.6 ± 0.2 | 1611.1 ± 0.7 | 2.55 ± 0.45 | β-sheet |
Table 5: Secondary Structure of Nail Protein of Control Nail Clipping Revealed by Peak-Fitting Analysis
As the nail plate proteins are mainly composed of hard α-keratin and are known to have abundant α-helix, which contributes to the stabilization of the nail protein structure [21], therefore, a decrease of α-helix component would result in reduction of the integrity of nail proteins. The results of peak-fitting analysis of permeation enhancers-treated nail clippings (Table 6) showed a significant loss of α-helix and increase of β-sheet and β-turns in the nail protein structure when treated with all the tested permeation enhancers except the HP-β-CD, which did not significantly alter these secondary structures of nail protein. Majority of the tested permeation enhancers, such as oxalic acid, resorcinol, salicylic acid, sodium hydroxide, sodium sulphite, and thioglycolic acid were found to be very harsh on nail proteins as the α-helix component was completely disappeared from their peak-fitted graphs. When the nail clippings were treated with n-acetyl-l-cysteine the α-helix component though was reduced less (18.95 ± 0.45 %) as compared to the control nail clippings (19.08 ± 0.24 %) but the other secondary components, i.e. β-turn, β-sheet, and random coils were significantly changed. Therefore, the n-acetyl-l-cysteine also could not be considered as safe permeation enhancer for transungual applications. Hence, the HP-β-CD was finally emerged as the safest permeation enhancer among all the eight permeation enhancers tested.
| Permeation enhancer | Secondary structure of nail protein (%) (Mean ± SD) |
|||
|---|---|---|---|---|
| β-turn | β-sheet | Random coil | α-helix | |
| Control | 37.24 ± 1.24 | 27.54 ± 1.45 | 16.14 ± 0.64 | 19.08 ± 0.24 |
| HP-β-CD | 36.84 ± 1.17 | 26.14 ± 1.28 | 15.97 ± 0.57 | 19.12 ± 0.17 |
| N-acetyl-l-cysteine | 27.84 ± 0.96 | 34.81 ± 1.72 | 18.39 ± 0.64 | 18.95 ± 0.45 |
| Oxalic acid | 46.77 ± 1.54 | 35.08 ± 1.95 | 18.14 ± 0.15 | 0 |
| Resorcinol | 49.41 ± 1.63 | 31.44 ± 1.24 | 19.15 ± 0.12 | 0 |
| Salicylic acid | 56.13 ± 2.35 | 34.65 ± 1.42 | 9.22 ± 0.12 | 0 |
| Sodium hydroxide | 40.29 ± 1.57 | 36.37 ± 1.25 | 23.34 ± 0.98 | 0 |
| Sodium sulphite | 44.63 ± 1.25 | 37.49 ± 1.86 | 17.88 ± 0.12 | 0 |
| Thioglycolic acid | 36.34 ± 1.27 | 33.94 ± 1.35 | 29.71 ± 0.78 | 0 |
Table 6: Effect of Permeation Enhancers on Secondary Structure of Nail Protein
The effect of eight permeation enhancers on integrity of nail plate was determined by measuring the changes brought about by these agents on the secondary structure of keratin protein of nail plate. The permeation enhancer, which altered the secondary structure of nail plate protein and hence damaged the integrity and physical stability of the nail plate, could be identified by the safety screening test designed in the present investigations by measuring changes in the FTIR spectra of nail protein by employing three spectral parameters namely, peak purity index, peak intensity ratio, and peak-fitting analysis. Thioglycolic acid, salicylic acid, sodium sulphite, oxalic acid, resorcinol, and n-acetyl-l-cysteine substantially affected the secondary structure of keratin protein of nail plate as evinced by changes in the values of peak purity index, peak intensity ratio, and peak-fitting analysis as compared to control nail clippings. On the other hand HP-β-CD showed minimal and insignificant effect on the values of peak purity index, peak intensity ratio, and peak-fitting analysis as compared to control nail clippings and therefore it was found to be the safest amongst all the tested permeation enhancers. The present FTIR-based nail safety screening test appears to be a novel, simple, scientific, accurate, and non-invasive technique of testing the safety of transungual permeation enhancers. This technique could be effectively employed for screening of nailfriendly permeation enhancers from a large pool of available chemical agents during preformulation stage of transungual product development.
Acknowledgements
Authors would like to acknowledge Roquette Corporate (Lestrem Cedex, France), for generous supply of gift sample of hydroxypropyl-β-cyclodextrin (Kleptose® HPB) for present studies.
Conflict of interest
The authors declare that they have no conflict of interest.
References
- Gupta AK, Versteeg SG, Shear NH. Onychomycosis in the 21st Century: An Update on Diagnosis, Epidemiology, and Treatment. J Cutan Med Surg 2017;21(6):1-15.
- Hao J, Smith KA, Li SK. Iontophoretically enhanced ciclopirox delivery into and across human nail plate. J Pharm Sci 2009;98:3608-16.
- Hao J, Smith KA, Li SK. Chemical method to enhance transungual transport and iontophoresis efficiency. Int J Pharm 2008;357:61-9.
- Brown MB, Khengar RH, Turner RB, Forbes B, Traynor MJ, Evans CR, et al. Overcoming the nail barrier: A systematic investigation of ungual chemical penetration enhancement. Int J Pharm 2009;370:61-7.
- Mohorčič M, Torkar A, Friedrich J, Kristl J, Murdan S. An investigation into keratinolytic enzymes to enhance ungual drug delivery. Int J Pharm 2007;332:196-201.
- Murdan S. Drug delivery to the nail following topical application. Int J Pharm 2002;236:1-26.
- Chouhan P, Saini TR. Hydration of nail plate: a novel screening model for transungual drug permeation enhancers. Int J Pharm 2012;436:179-82.
- Barba C, Méndez S, Martí M, Parra JL, Coderch L. Water content of hair and nails. Thermochimica Acta 2009;494:136-40.
- Egawa M, Fukuhara T, Takahashi M, Ozaki Y. Determining water content in human nails with a portable near-infrared spectrometer. Appl Spectrosc 2003;57:473-8.
- Natalello A, Ami D, Brocca S, Lotti M, Doglia SM. Secondary structure, conformational stability and glycosylation of a recombinant Candida rugosa lipase studied by Fourier-transform infrared spectroscopy. Biochem J 2005;385:511-7.
- Sakudo A, Kuratsune H, Kato YH, Ikuta K. Secondary structural changes of proteins in fingernails of chronic fatigue syndrome patients from Fourier-transform infrared spectra. Clin Chim Acta 2009;402:75-8.
- Murthy SN, Vaka SR, Sammeta SM, Nair AB. TranScreen-N: Method for rapid screening of trans-ungual drug delivery enhancers. J Pharm Sci 2009;98:4264-71.
- Li J, Hibbert DB, Fuller S, Vaughn G. A comparative study of point-to-point algorithms for matching spectra. Chemometr Intell Lab Syst 2006;82:50-8.
- Haris PI, Chapman D. Analysis of polypeptide and protein structures using Fourier transform infrared spectroscopy. In: Jones C, Mulloy B, Thomas AH, editors. Methods in Molecular Biology: Microscopy, Optical Spectroscopy, and Macroscopic Techniques. Totowa, New Jersey: Humana Press; 1994. p. 183-202.
- Barth A, Zscherp C. What vibrations tell us about proteins. Q Rev Biophys 2002;35:369-430.
- Kakkar P, Madhan B, Shanmugam G. Extraction and characterization of keratin from bovine hoof: A potential material for biomedical applications. Springerplus 2014;3:596.
- Silva R, Singh R, Sarker B, Papageorgiou DG, Juhasz JA, Roether JA, et al. Hybrid hydrogels based on keratin and alginate for tissue engineering. J Mater Chem B 2014;2:5441-51.
- Sowa MG, Wang J, Schultz CP, Ahmed MK, Mantsch HH. Infrared spectroscopic investigation of in vivo and ex vivo human nails. Vib Spectrosc 1995;10:49-56.
- Gniadecka M, Faurskov NO, Christensen DH, Wulf HC. Structure of water, proteins, and lipids in intact human skin, hair and nail. J Invest Dermatol 1998;110:393-98.
- Wang HY, Lü Y, Wang F, Ma XD, Jiang SP, Wang W, et al. Study on FTIR spectra of finger nails of normal people and patients of esophagus cancer. Guang Pu Xue Yu Guang Pu Fen Xi 2008;8:331-34.
- Marshall RC, Orwin DF, Gillespie JM. Structure and biochemistry of mammalian hard keratin. Electron Microsc Rev 1991;4:47-83.