- *Corresponding Author:
- S. Narayanan
Department of Biotechnology, Center for Post Graduate Studies, Jain University, Bangalore-560 011, India
E-mail: sam19narayan@yahoo.co.in
| Date of Submission | 31 October 2016 |
| Date of Revision | 29 January 2017 |
| Date of Acceptance | 31 May 2017 |
| Indian J Pharm Sci 2017; 79(4): 625-632 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
Presence of multidrug resistant strains makes treatment of tuberculosis difficult. Conventional frontline antituberculosis therapy has high incidence of treatment failure due to non-adherence and severe side effects, and therefore warrants alternative approaches using safe, natural compounds. (-) Epigallocatechin gallate, a green tea catechin with significant antimicrobial properties, and its derivatives have been studied for their inhibitory effects on Mycobacterium smegmatis. Enoyl reductase (Inh A, PDB ID: 2NV6), an important drug target inhibited by the antituberculosis drug, isoniazid, was docked with geometrically optimized conformation of epigallocatechin gallate and two of its derivatives using AutoDock software. Perpropionate and permethyl derivatives of epigallocatechin gallate were synthesized and, along with epigallocatechin gallate, evaluated for their inhibitory and cytotoxic properties against M. smegmatis using microplate dilution method and alamarBlue® assay, respectively. Ames test was performed to assess the mutagenic potential of epigallocatechin gallate. It was docked successfully onto the InhA with docking energy of –9.38 kcal mol-1. Analysis of neighboring groups revealed 11 amino acid residues common to both isoniazid and epigallocatechin gallate. The minimum inhibitory concentrations for epigallocatechin gallate, perpropionate and permethyl were found to be 128 (58.2% inhibition), 8 (32.9%) and 4 (12.5%) µg/ml, respectively. Epigallocatechin gallate showed a cytotoxicity of 18.6% at a concentration of 8 µg/ ml, while the two derivatives did not show any toxicity, even at 64 µg/ml. Ames test revealed the absence of mutagenicity for epigallocatechin gallate, in presence and absence of metabolic activation. Non-mutagenic epigallocatechin gallate inhibits the growth of M. smegmatis and warrants further investigation as an adjunct therapy for pathogenic mycobacteria.
Keywords
Tuberculosis, Mycobacterium, mutagenicity, EGCG, green tea polyphenol, enoyl reductase
Tuberculosis (TB) is one of the most serious diseases and has been the scourge of the mankind since ancient times. As per the WHO estimate of 2015, TB has been responsible for over 1.8 million deaths worldwide annually [1]. This coupled with the occurrence of multidrug resistant (MDR) strains of the pathogen and co-infection with human immunodeficiency virus (HIV) has rendered TB patients highly susceptible to the disease [2]. In addition, conventional frontline antituberculosis (antiTB) therapy is known to have high incidence of treatment failure if the prescribed regimen is not followed [3]. The drugs used to treat TB themselves possess marked side effects, such as blurring or loss of vision and neuritis caused by ethambutol, and abdominal pain, bleeding gums, fever and headache by rifampin [4,5]. Isoniazid is reported to cause nausea, vomiting, stomach effects and allergic reactions [6]. Consequently, there is an urgent need to explore alternative approaches to treatment of TB using compounds derived from natural substances with almost negligible side effects and their possible use in combination therapy.
Mycobacteria are tenacious microorganisms resistant to common disinfectants due to the presence of a lipid coat of mycolic acid. Mycolic acid is synthesized through an elaborate biosynthetic pathway regulated by enzymes of the fatty acid synthase (FAS) complex. One of the key enzymes in this pathway is enoyl reductase, also known as InhA. This enzyme has been regarded as an important drug target and is inhibited by the leading antiTB drug, isoniazid [7]. This has prompted us to examine inhibition of InhA by Epigallocatechin gallate (EGCG) using in silico studies followed by in vitro studies.
EGCG is one of the poly phenolic catechins found in green tea. Green tea has been reported to have significant antimicrobial properties [8]. EGCG has not been evaluated in detail for the inhibition of Mycobacterium tuberculosis (MT), although few reports have mentioned some antimycobacterial activity based on enzyme inhibition studies in combination with compounds, such as triclosan [9]. They have also reported that EGCG, inhibits InhA, the enoyl-ACP reductase of MT and interferes with the binding of NADH to InhA. Kohda et al. have reported significant antimicrobial activity of ECGC against Listeria monocytogenes, another intracellular bacterium like MT [10]. Hu et al. have observed a potent synergy between EGCG and Ampicillin/Sulbactam in the treatment of Methicillinresistant Staphylococcus aureus infection [11]. Cho et al. have reported that EGCG exhibited synergistic activity with the antibiotic Imipenem in treatment of the pathogenic bacteria, Klebsiella pneumoniae implicated in pneumonia infections [12]. Novy et al. have also reported the synergistic and additive effects of EGCG in combination with antibiotics such as oxytetracycline against S. aureus strains [13].
EGCG and three of its derivatives have been synthesized and evaluated for their inhibitory properties against M. smegmatis a non-pathogenic bacterium, genetically similar to MT, using the microplate dilution method. AlamarBlue® assay, a rapid and highly sensitive method to measure cell proliferation and cytotoxicity in various bacteria, fungi and cell lines has also been used to determine the cytotoxicity of EGCG and its derivatives [14]. In the long and often tortuous path of drug discovery, a candidate drug needs to be checked for its mutagenic potential before it is taken up for therapeutic evaluation in a clinical setting, since both cancer and mutagenicity are often linked together. EGCG and its derivatives have also been evaluated for mutagenicity using the Ames test (bacterial reverse mutation test) [15].
Materials and Methods
EGCG was purchased from Carbosynth (Berkshire, UK). Three derivatives of EGCG, viz. EGCG perpropionate, permethyl EGCG and pentamethyl EGC were synthesized from EGCG based on the method of Utenova et al. [16]. In the case of pentamethyl EGC, a modified procedure was used, where the permethyl EGCG was hydrolysed with lithium hydroxide (2 equivalent) in a mixture of tetrahydrofuran:methanol:water (2:2:1) at room temperature to yield pentamethyl EGC.
In silico studies
EGCG structure was drawn using Gauss View package (Gaussian Inc., CT. USA) and subsequently optimized using Gaussian package [17]. Hartree-Fock theory with “6-31++g (d, p)” as the basis set was used for optimizing the structures. Standard orientation of EGCG after convergence to its global energy minima was visualized using Argus package (Argus Lab, WA USA), and saved in PDB format. Geometrically optimized EGCG was later docked onto the crystal structure of the enoyl reductase receptor protein (PDB ID:2NV6) using the AutoDock ver.4.0 package (The Scripps Research Institute, La Jolla, CA, USA) described elsewhere [18]. Docking energy was calculated based on the shape and electrostatics using default grid spacing of 0.375 Å. The docked structure was analysed using PyMOL while the neighbouring residues of the docked structure were identified using Swiss PDB Viewer [19].
Microplate dilution method
The minimum inhibitory concentration (MIC) of EGCG and its derivatives were determined using the microplate dilution method. Briefly, broth culture of M. smegmatis was grown using Muller Hinton (MH) broth. The compounds were serially diluted in the medium (MH broth) and 100 μl of each solution was added into the wells of a 96-well plate. The following concentrations were used-EGCG (1, 2, 4, 8, 16, 32, 64 μg/ml), EGCG perpropionate and permethyl EGCG (0.25. 0.5, 1, 2, 4, 8, 16, 32, 64 μg/ml), ciprofloxacin (0.25, 0.5, 1, 2, 4, 8, 16, 32, 64 μg/ml), rifampicin (0.29, 0.6, 1.2, 3.1, 6.3, 12.5, 25.0, 50.0, 64 μg/ml), amikacin (0.98, 2, 4, 8, 16, 32, 64, 128 μg/ml). The cells were matched with the McFarland standard 0.5, which represents 108 cells and then were diluted to 105 cells in the medium. Hundred microliters of this cell suspension was added into each well. The plate was incubated at 37° for 48 h and the optical density was measured at 600 nm on a Power Wave X Microplate Reader (BioTek, Vinooski, VT, USA). The percent inhibition of cell growth by the given compound was calculated using the Eqn., percent inhibition=(Av.OD600 (control)–OD600 (test))/ (Av.OD600 (control)×100.
AlamarBlue® assay
This dye reduction assay was performed in a 96-well plate, using ciprofloxacin and amikacin as positive controls. Briefly, broth culture of M. smegmatis was grown using MH broth [14]. The test compounds and controls were also serially diluted in MH broth. Hundred microliters each of the diluted solutions was added into the wells. The following concentrations were used-EGCG (8, 16, 32, 64 μg/ml), EGCG perpropionate, permethyl EGCG and pentamethyl EGCG (32, 64, 128), ciprofloxacin (0.03, 0.06, 0.12, 0.25, 0.50, 1.0 μg/ml) and amikacin (0.045, 0.09, 0.18, 0.37, 0.75, 1.50 μg/ml). The cells were matched with the McFarland standard 0.5, which represents 108 cells. The cells were then diluted to 105 cells and 100 μl of the diluted cells were added into each well. Ten microliters of alamarBlue dye was then added to the wells. The plate was incubated at 37° for 48 h. The dye reduction was visualized as change of colour from blue (oxidized) to pink (reduced) indicating viability of cells. The fluorescence intensity (FI) was read on a Power Wave X Microplate Reader under excitation at 560 nm and emission at 590 nm. The percent reduction of alamarBlue (=%cell viability) was calculated using the formula, % reduction=(FI590 (test)–FI590 (untreated control))/(FI590 (100% reduced AB)–FI590 (untreated control))×100; percent cytotoxicity=100– percent reduction.
Mutagenicity assay
Ames assay was performed to assess mutagenicity using the following seven bacterial strains of Salmonella typhimurium – TA 98, TA 100, TA 1535, TA1537, TA97a, TA102; Escherichia coli – WP2 uvrA (pKM101) [15]. Seven wells (one each for positive and vehicle control, 5 test compound concentrations) in three replicates were taken for each compound. Maximum concentration taken was up to 1000 μg/ plate. Bacteria were exposed to the test item both in the presence and absence of metabolic activation (by the S9 rat liver homogenate). Plating was done in a six well Petri plate. A volume of 0.5 ml top agar containing concentrations of test item/vehicle/positive control (20 μl), bacterial culture suspension (20 μl) and S9 mix/PBS (100 μl) were plated on to 4-6 ml basal agar (Vogel Bonner agar). The plates were incubated from 48 to 72 h and observed for the following parameters: precipitation, bacterial background lawn and the reverted mutants were counted. The test item EGCG would be considered mutagenic in this assay if, a concentration related increase in revertant numbers of ≥2‑fold of the concurrent vehicle control values was observed.
Cytotoxicity and precipitation
The presence or absence of cytotoxicity was determined as mentioned in OECD Guideline 471, that is, by a reduction in the number of revertant colonies, a clearing or diminution of the background lawn, or the degree of survival of treated cultures, in presence and absence of metabolic activation. The precipitation was also assessed for each concentration of EGCG as mentioned in the above guidelines.
Results and Discussion
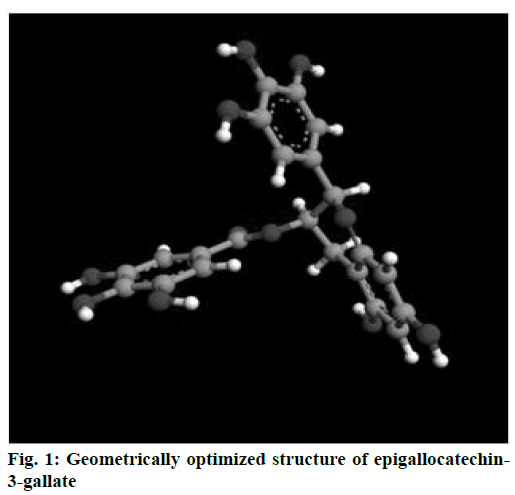
Energy minimized structure of EGCG after Gaussian optimization represented in Figure 1. Characteristic parameters of the optimized structure are shown in Table 1. Lowest docking energy was –9.38 kcal mol-1 observed in Run 8 of the clustering histogram shown in Table 2. The docked structure of EGCG with 2NV6 is shown in Figure 2. The neighbouring residues for EGCG are depicted in Figure 3a and the 11 amino acid residues marked with an asterisk are common to both EGCG and isoniazid (Zid). These residues are Zid300*, Ile122*, Phe97*, Gly96*, Ile95*, Ala94, Gln66*, Val65*, Leu63*, Ile47, Leu44, Arg43, Asp42, Phe41*, Gly40, Ile16*, Gly14*.
| Parameter | Value | Threshold | Convergence |
|---|---|---|---|
| Maximum force | 0.000025 | 0.000450 | Yes |
| RMS force | 0.000005 | 0.000300 | Yes |
| Maximum displacement | 0.001478 | 0.001800 | Yes |
| RMS displacement | 0.000334 | 0.001200 | Yes |
| Predicted change in energy | –1.895627×10-8 | ||
| Optimization | Completed | ||
| Stationary point | Found | ||
Table 1: Energy Minimization Parameters of EGCG
| Cluster rank | Lowest binding energy | Run | Mean binding energy | Number in clusters |
|---|---|---|---|---|
| 1 | –9.38 | 8 | –8.24 | 7 |
| 2 | –8.74 | 11 | –8.74 | 1 |
| 3 | –8.52 | 22 | –8.52 | 1 |
| 4 | –8.47 | 5 | –8.47 | 1 |
| 5 | –8.41 | 7 | –8.41 | 1 |
| 6 | –8.32 | 18 | –8.21 | 3 |
| 7 | –7.65 | 6 | –7.39 | 2 |
| 8 | –7.63 | 1 | –6.88 | 3 |
| 9 | –7.29 | 21 | –7.29 | 1 |
| 10 | –7.27 | 2 | –7.27 | 1 |
| 11 | –7.22 | 23 | –7.22 | 1 |
| 12 | –6.83 | 14 | –6.83 | 1 |
| 13 | –6.61 | 12 | –6.61 | 1 |
| 14 | –6.47 | 17 | –6.47 | 1 |
Table 2: Clustering Histogram: Egcg Docked with 2NV6
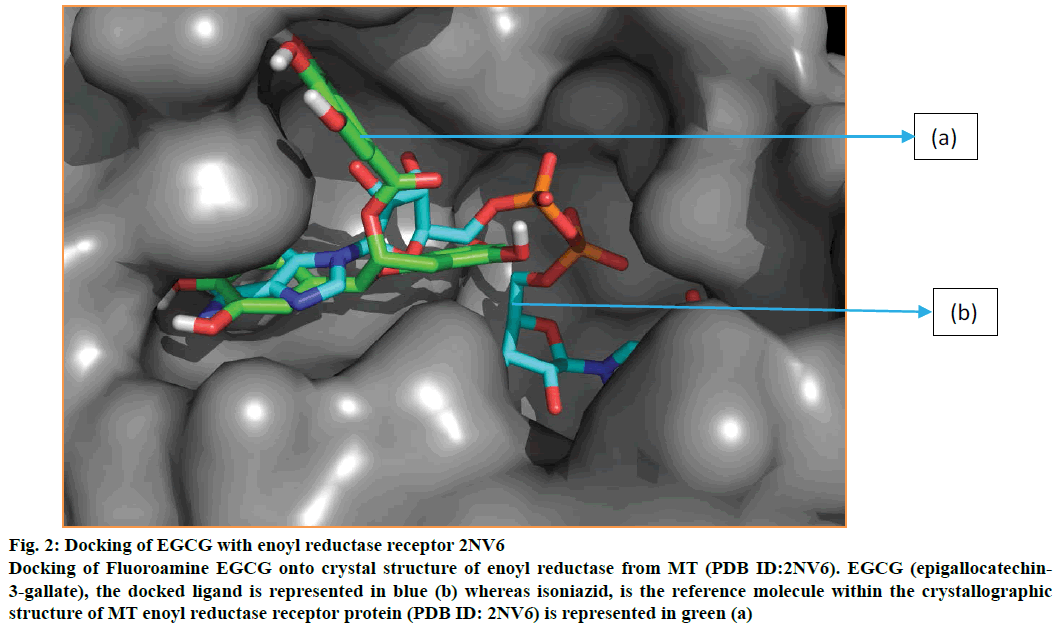
Figure 2: Docking of EGCG with enoyl reductase receptor 2NV6
Docking of Fluoroamine EGCG onto crystal structure of enoyl reductase from MT (PDB ID:2NV6). EGCG (epigallocatechin-3-gallate), the docked ligand is represented in blue (b) whereas isoniazid, is the reference molecule within the crystallographic structure of MT enoyl reductase receptor protein (PDB ID: 2NV6) is represented in green (a)
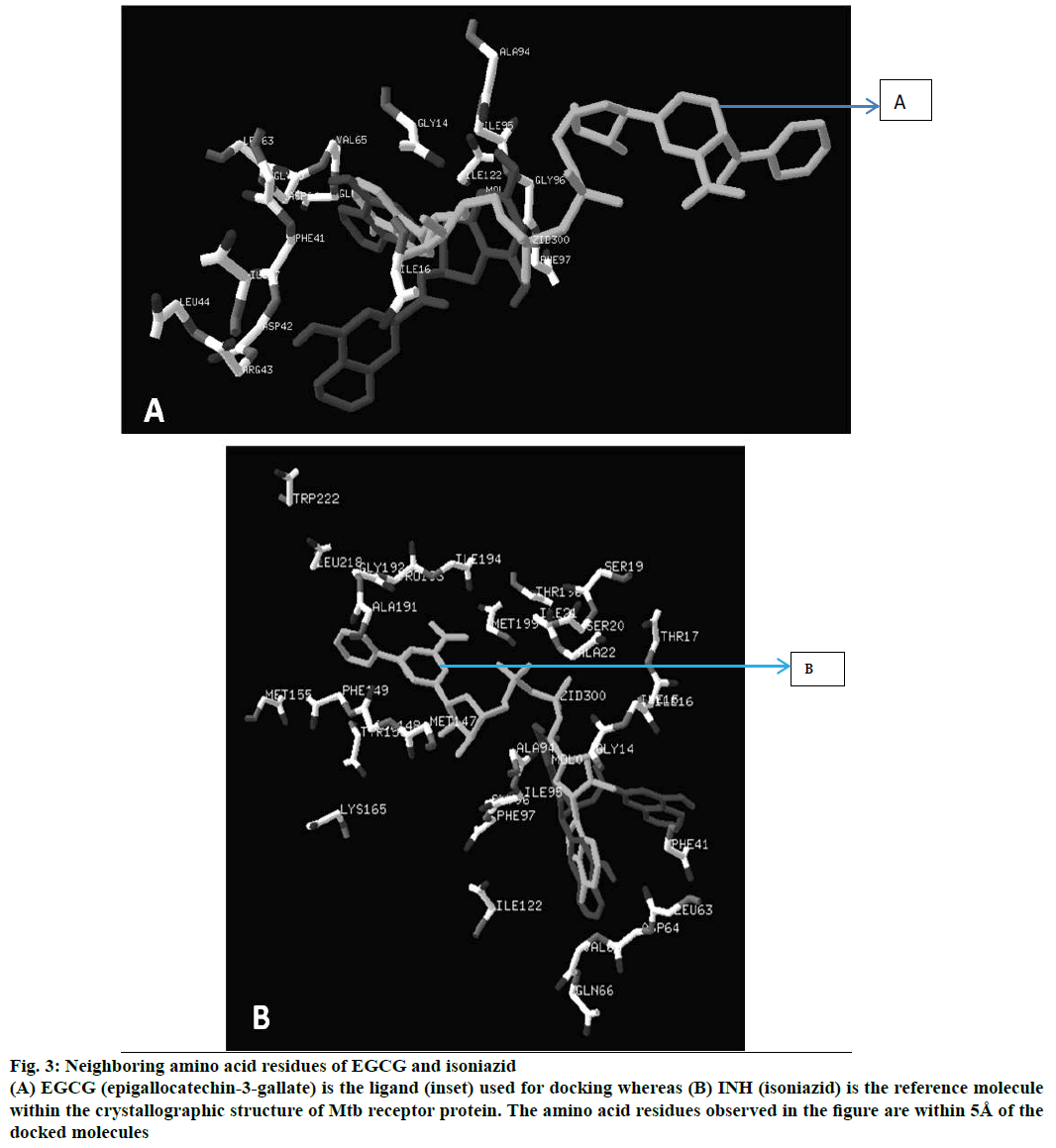
Figure 3: Neighboring amino acid residues of EGCG and isoniazid
(A) EGCG (epigallocatechin-3-gallate) is the ligand used for docking whereas (B) INH (isoniazid) is the reference molecule within the crystallographic structure of Mtb receptor protein. The amino acid residues observed in the figure are within 5Å of the docked molecules
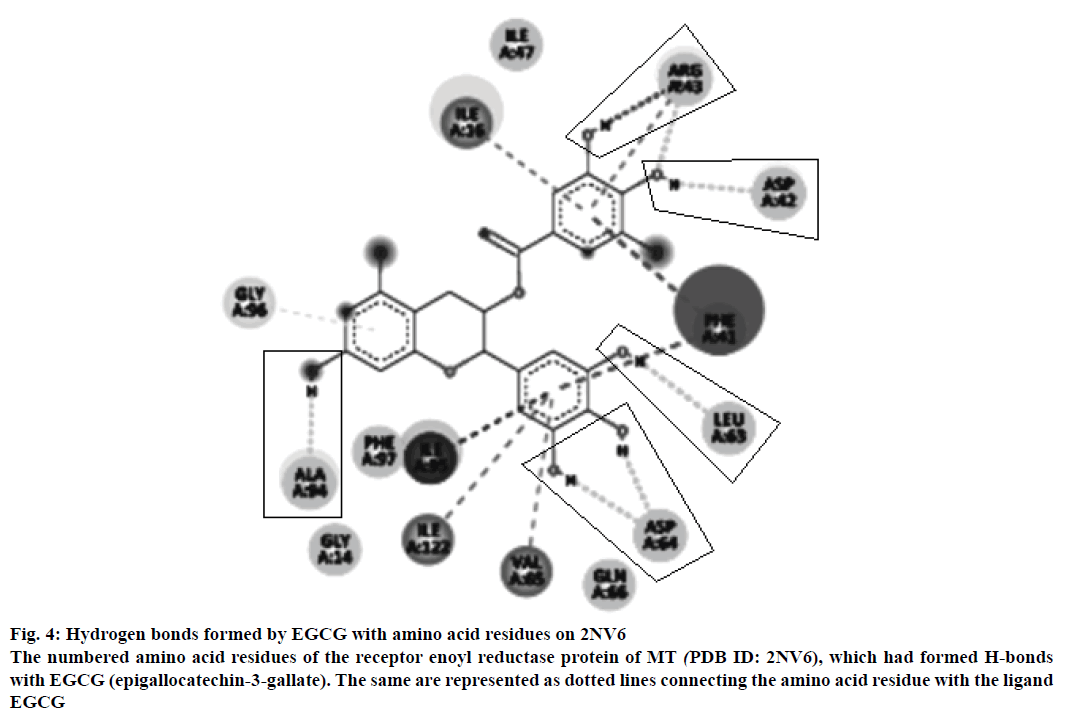
The neighbouring residues for isoniazid are depicted in Figure 3b and eight residues marked with an asterisk are common to both Zid and EGCG (mol): MolA0*, Trp222, Leu218, Met199, Thr196, Ile194, Pro193, Gly192, Ala191, Lys165, Tyr158, Met155, Phe149, Asp148, Met147, Ile122*, Phe97*, Gly96*, Ile95, Ala94*, Gln66, Val65*, Asp64*, Leu63*, Phe41, Ala22, Ile21, Ser20, Ser19, Thr17, Ile16*, Ile15, Gly14*. In all, there are 11 neighbouring residues common to both these ligands. There are five hydrogen bonds observed in the docked structure as shown in Figure 4. These are located between Ala94 --- O; Asp64 --- OD1; Leu63 --- O; Asp42 --- OD2; Arg43 --- N.
Figure 4: Hydrogen bonds formed by EGCG with amino acid residues on 2NV6
The numbered amino acid residues of the receptor enoyl reductase protein of MT (PDB ID: 2NV6), which had formed H-bonds with EGCG (epigallocatechin-3-gallate). The same are represented as dotted lines connecting the amino acid residue with the ligand EGCG
The extent of inhibition and cytotoxicity of EGCG and its synthesized derivatives, determined by the microplate dilution assay and alamarBlue assay, respectively, are shown in Table 3. Microplate dilution assay results indicate that the antibiotics viz. ciprofloxacin, rifampicin and amikacin showed an MIC50 at 0.25, 0.29 and 0.98 μg/ml, respectively, in accordance with the expectation. EGCG showed the MIC at 128 μg/ml (58.2% inhibition), however, inhibition of bacterial growth could be observed even at 64 μg/ml also (28.3% inhibition). EGCG perpropionate showed a slight inhibition of 18.8% at concentration of 1 μg/ml and 32.9% at 8 μg/ml, and no inhibition thereafter at any other concentration tested. Permethyl EGCG showed slight inhibition of 12.5% at a concentration of 4 μg/ml and no inhibition at other tested concentrations. Pentamethyl EGC did not show any inhibition at the concentrations tested.
| Compound | Microplate inhibition assay | AlamarBlue assay | ||
|---|---|---|---|---|
| Inhibition (%) | Concentration | Cytotoxicity (%) | Concentration | |
| (µg/ml) | (µg/ml) | |||
| EGCG | 58.2 | 128α | 82.6 | 64α |
| 28.3 | 64α | 85.3 | 32α | |
| 74.1 | 16α | |||
| 18.6 | 8α | |||
| EGCG Per propionate | 32.9 | 8α | Not observed | 32 to 128 |
| 18.8 | 1α | |||
| Per methyl EGCG | 12.5 | 4α | Not observed | 32 to 128 |
| Penta methyl EGC | ND | ND | Not observed | 32 to 128 |
Table 3: Microplate Inhibition and Alamar Blue Assays
In alamarBlue assay, the growth of M. smegmatis was inhibited by ciprofloxacin (72.5%) at 0.031 μg/ml, amikacin (35.8%) at 0.045 μg/ml. In the case of EGCG, the lowest concentration tested (8 μg/ml) exhibited 18.5% inhibition, which increased gradually to 74.1% at 16 μg/ml. At further higher concentrations, viz. 32 and 64 μg/ml, the effect saturated in the range of 82.6-85.3%, as shown in Table 3. EGCG perpropionate, permethyl ECGG, and pentamethyl EGC did not show any cytotoxicity towards M. smegmatis.
The viable cell counts were within the required range of 1-2×109 for the cultures. Data from control treatments confirmed that the revertant counts were within the historical as well as in-house spontaneous counts for each of the strains tested. The correctness of strain, functioning of the assay, and the data were accepted as valid since genotypic characterizations had been already verified earlier [15]. Following treatment with EGCG, no significant increase in revertant colonies (at least 2-fold increase expected) were observed for any of the tested strains under any condition of metabolic activation. The positive controls induced 5 to 25-fold increases in revertant numbers for different strains when compared to the concurrent vehicle controls.
EGCG did not precipitate on the plates at any concentration tested, indicating no cytotoxicity in the preliminary test. Absence of cytotoxicity was determined as mentioned in OECD Guideline 471, that is there was no reduction in the number of revertant colonies, no clearing or diminution of the background lawn, no decrease in degree of survival of treated cultures, in presence and absence of metabolic activation. Taken together, the results of this bacterial reverse mutation assay establish that EGCG does not cause a positive mutagenic response with any of the tester strains, either the presence or absence of metabolic activation, under the conditions of the assay.
Presently, TB continues to be a rampaging global disease with high morbidity and significant mortality, especially in undernourished populations or those already infected with HIV and other concurrent infections, which reduce the immunity of the patient [1]. Emergence of MDR strains has added to the difficulties of treating this dreaded disease with the existing directly observed treatment short course (DOTS) regimen, which calls for a high degree of compliance [2]. Further, many of the frontline drugs against TB are known to possess severe side effects. This necessitates a rethink in our approach to the treatment of this insidious disease. EGCG, a polyphenol found in green tea has been reported to have wide antimicrobial properties [8]. However, this compound has not been evaluated, in detail, for the inhibition of MT, although few reports have mentioned some antimycobacterial activity based on enzyme inhibition studies in combination with compounds, such as triclosan [9]. It was therefore decided to evaluate EGCG and few derivatives using a combination of approaches comprising in silico and in vitro studies. The results of in silico docking studies indicated good docking energy of the geometrically optimized ligand EGCG with InhA receptor 2NV6, an important link in the FAS enzyme complex responsible for synthesis of mycolic acid in Mycobacteria. Earlier, Ramesh et al. have docked epigallocatechin (EGC) to a homology-modelled domain of ketoacyl-ACP reductase (KR) protein of MT [20]. In the present study, EGCG, the gallate ester of EGC was able to be successfully docked on to enoyl reductase receptor 2NV6 with good binding energy. Presence of number of common residues between EGCG and isoniazid already present in the receptor further indicates similar binding sites. Presence of five hydrogen bonded residues in the docked structure further substantiates our in silico findings. He et al. have earlier reported synthetic carboxamides to inhibit the enoyl ACP protein [21]. However, to the best of our knowledge, there are no reports of inhibition of InhA of Mtb by EGCG alone by docking studies.
The above findings were corroborated by the results of our in vitro findings wherein we observed that EGCG was able to inhibit growth of M. smegmatis, a Mycobacterium genetically similar to MI, both in the microplate dilution inhibition assay as well as the sensitive alamarBlue assay. However, the derivatives permethyl EGCG and EGCG perpropionate showed only slight inhibition of M. smegmatis in the microplate dilution assay and no inhibition in the alamarBlue assay. Utenova et al. [16] have earlier tested EGCG and few derivatives, such as permethyl EGCG and EGCG perpropionate for their TB inhibition properties against MT (H37RV) in Bactec 12B medium using the alamarBlue assay [14]. They have observed that while permethyl EGCG and EGCG perpropionate showed some inhibition, EGCG itself did not show any TB inhibitory activity. On the other hand, present findings regarding the MICs of EGCG (MIC between 8-16 μg/ml) are broadly in agreement with those of Sivakumar et al. who have studied the antimycobacterial properties of few medicinal plants, such as Allium species by alamarBlue assay [22]. They have reported the inhibitory properties of these compounds towards Mycobacteria at concentrations of 50-100 μg/ml. Ugwu et al. have also shown that the inhibitory concentrations of carboxamides against MT is in the range of 25-50 μg/ml [23]. Cho et al. showed that the inhibitory concentrations of EGCG against the antibiotic resistant Imipenem Klebsiella pneumonia strains to be in the range of 300-650 μg/ml [12].
The Ames test is a well-established, robust, in vitro mutagenicity assay used by regulatory agencies, such as OECD to screen lead compounds for their mutagenic potential [24]. EGCG was found to be non-mutagenic by this assay and actively suggests towards being a promising candidate therapeutic molecule for further investigations. In conclusion, present in silico and in vitro studies strongly suggests that EGCG is able to inhibit M. smegmatis. This compound is also non-mutagenic as observed in the Ames test. Further investigations towards the development of EGCG as an adjunct therapy for TB are recommended based on the present promising findings. Additional studies, such as molecular dynamics simulation of EGCG, 2NV6 and their complex are being performed by the authors.
Acknowledgements
The authors are grateful to Lakshmi Mundkur, Vrushali Dandavate for their assistance with the in vitro assays; to Sangeetha, Gireesh Kamath for the Ames assay. Financial assistance from Syngene Higher Education Fund and permission from Jain University, Bengaluru, India are gratefully acknowledged.
Financial assistance
None.
Conflict of interests
None declared.
References
- http://www.who.int/tb/publications/global_report/en/.
- Cifuentes DP, Ocampo M, Curtidor H, Vanegas M, Forero M, Patarroyo ME, et al. Mycobacterium tuberculosisRv0679c protein sequences involved in host-cell infection: Potential TB vaccine candidate antigen. BMC Microbiol 2010;10:109.
- Kimerling ME, Kluge H, Vezhnina N, Iacovazzi T, Demeulenaere T, Portaels F, et al. Inadequacy of the current WHO re-treatment regimen in a central Siberian prison: Treatment failure and MDR-TB. Int J Tuberc Lung Dis 1999;3:451-53.
- Griffith DE, Brown-Elliott BA, Shepherd S, McLarty J, Griffith L, Wallace RJ. Ethambutol ocular toxicity in treatment regimens for Mycobacterium avium complex lung disease. Am J Resp Crit Care Med 2005;172;250-53.
- https://extranet.who.int/prequal/sites/default/files/documents/Comparator-TB2017-15September.pdf.
- Dutt AK, Moers D, Stead WW. Undesirable side effects of isoniazid and rifampin in largely twice-weekly short-course chemotherapy for tuberculosis. Am Rev Respir Dis 1983;128:419-24.
- Takayama K, Wang C, Besra GS. Pathway to synthesis and processing of mycolic acids in Mycobacterium tuberculosis. Clin Microbiol Rev 2005;18:81-101.
- Das S, Tanwar J, Hameed S, Fatima Z, Manesar G. Antimicrobial potential of epigallocatechin-3-gallate (EGCG): a green tea polyphenol. J Biochem Pharmacol Res 2014;2:167-74.
- Sharma SK, Kumar G, Kapoor M, Surolia A. Combined effect of epigallocatechin gallate and triclosan on Enoyl-ACP reductase of Mycobacteriumtuberculosis. Biochem Biophys Res Commun 2008;368:12-7.
- Kohda C, Yanagawa Y, Shimamura T. Epigallocatechin gallate inhibits intracellular survival of Listeria monocytogenes in macrophages. Biochem Biophys Res Commun 2008;365:310-5.
- Hu ZQ, Zhao WH, HaraY, Shimamura T. Epigallocatechin gallate synergy with Ampicillin/sulbactam against 28 clinical isolates of methicillin-resistant Staphylococcus aureus. J Antimicrob Chemother 2001;48:361-64.
- Cho YS, Oh JJ, Oh KH. Synergistic anti-bacterial and proteomic effects of epigallocatechin gallate on clinical isolates of imipenem-resistant Klebsiella pneumonia. Phytomedicine 2001;18:941-6.
- Novy P, Rondevaldova J, Kourimska L, Koroska L. Synergistic interactions of epigallocatechin gallate and oxytetracycline against various drug resistant Staphylococcus aureusstrains in vitro. Phytomedicine 2013;20:432-35.
- Rampersad SN. Multiple applications of Alamar Blue as an indicator of metabolic function and cellular health in cell viability bioassays. Sensors (Basel) 2012;12:12347-60.
- Maron DM, Ames B. Revised methods for the Salmonellamutagenicity test. Mutat Res 1983;113:173-215.
- Utenova BT, Malterud KE, Rise F. Antioxidant activity of O-protected derivatives of (-)-epigallocatechin-3-gallate: inhibition of soybean and rabbit 15-lipoxygenases. Arkiv Org Chem 2007;9:6-16.
- Pople JA, Head-Gordon M, Fox DJ, Raghavachari K, Curtiss LA. Gaussian-1 theory: A general procedure for prediction of molecular energies. J Chem Phys 1989;90:5622-29.
- Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J Comput Chem 2009;30:2785-91.
- Guex N, Peitsch MC, Schwede T. Automated comparative protein structure modelling with SWISS-MODEL and Swiss-Pdb Viewer: a historical perspective. Electrophoresis 2009;30:S162-73.
- Ramesh KV, Chandy S, Pai D, Deshmukh S. In silicodocking of herbal based epigallocatechin onto homology modeled ketoacyl-ACP reductase domain of FAS protein from Mycobacteriumtuberculosis H37Rv. Indian J Biotechnol 2012;11:257-66.
- He X, Alian A, Stroud R, Ortiz de Montellano PR. Pyrrolidinecarboxamides as a novel class of inhibitors of enoyl acyl carrier protein reductase from Mycobacterium tuberculosis. J Med Chem 2006;49:6308-23.
- Sivakumar A, Emerson IA, Jayaraman G. Docking studies on transcription factor sp1: The transcriptional down-regulation of TACO Gene by EGCG and the Importance of TACO in M. tuberculosissurvival. Int J Drug Design Discov 2010;1:265-72.
- Ugwu DI, Ezema BE, Eze FU, Ugwuja DI. Synthesis and structural activity relationship study of antitubercular carboxamides. Intl J Med Chem 2014; 614808.
- https://www.oecd.org/chemicalsafety/risk-assessment/1948418.pdf.