- *Corresponding Author:
- Jiangang Liu
Department of Radiation Oncology, China
E-mail: 13685104715@163.com
| Date of Received | 30 March 2022 |
| Date of Revision | 12 February 2023 |
| Date of Acceptance | 28 September 2023 |
| Indian J Pharm Sci 2023;85(5):1398-1407 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
The main aim of this study was to investigate the biological characteristics of myristoylated alanine-rich C-kinase substrate analog-1 expression in esophageal squamous cell carcinoma tissues. Tumor tissues and associated paracancerous tissues were obtained from 74 individuals with esophageal squamous cell carcinoma hospitalized in Nanjing Lishui District People's Hospital between January 2016 and July 2019. According to immunohistochemically findings, the expression level of myristoylated alanine-rich C-kinase substrate analog-1 was comparable to that of patients, and its H-score in esophageal squamous cell carcinoma tissues was substantially greater than that in the equivalent paracancerous tissues (120.10±1.935 vs. 35.65±2.012, t=30.25, p<0.01). The tumor, node and metastasis staging was closely related (χ2=7.835, p<0.01). The expression of myristoylated alanine-rich C-kinase substrate analog-1 emerged as a significant independent risk indicator for overall survival in esophageal squamous cell carcinoma patients, as determined by Cox regression univariate and multivariate analysis (p=0.007; p=0.017). Cytological tests revealed that myristoylated alanine-rich C-kinase substrate analog-1 knockdown may substantially decrease esophageal squamous cell carcinoma cell proliferation, invasion, and migration. According to Western blotting experiments, the epithelial-mesenchymal transition process in esophageal squamous cell carcinoma cells might be inhibited by suppressing myristoylated alanine-rich C-kinase substrate analog-1. The outcomes of the bioinformatics study revealed that post-transcriptional regulation was connected to the dysregulated expression of myristoylated alanine-rich C-kinase substrate analog-1 in the tissues of esophageal cancer. Upstream mechanism experiments confirmed that microRNA-34a, the "suppressor" of esophageal cancer, can target myristoylated alanine-rich C-kinase substrate analog-1 and inhibit its expression at the post-transcriptional level. The current study demonstrated that myristoylated alanine-rich C-kinase substrate analog-1 may function as an "oncogene" in esophageal squamous cell carcinoma. The outcome of this investigation suggests that myristoylated alanine-rich C-kinase substrate analog-1 could become a potentially new biomarker in diagnosing esophageal squamous cell carcinoma and a new target for tumor therapy.
Keywords
Myristoylated alanine-rich C-kinase substrate analog-1, esophageal squamous cell carcinoma, invasion, migration, microRNA-34a
Esophageal Cancer (EC) stands out as a highly prevalent malignancy within the digestive tract. In 2020, there were approximately 604 000 new approximately of EC, ranking it as the 7th most prevalent among the 36 types of malignancies. Remarkably, it stood 3rd in terms of mortality rates[1]. In China, Esophageal Squamous Cell Carcinoma (ESCC) constitutes over 50 % of fresh EC diagnoses, marked by an unfavorable prognosis and an overall 5 y survival rate below 20 %[2]. Despite recent major advancements in radiation technology, the Overall Survival (OS) rate has not substantially increased[3]. In order to discover relevant tumor indicators and assist those with EC, it is therefore important to investigate the onset and progression of ESCC at the molecular level.
One of the members of the family of myristoylated alanine-rich C-kinase substrate proteins and a significant protein kinase C byproduct is Myristoylated Alanine-Rich C-Kinase Substrate Analog-1 (MARCKSL1)[4,5]. MARCKSL1 can combine with actin and calmodulin through its effector structure, affect its stability, enhance the movement ability of filopodia, regulate the formation of the cytoskeleton, and are crucial in the process of cell-cell interaction[6]. MARCKSL1, an essential regulator of cell motility, is a "cancer- promoting factor" in developing aggressive tumors[7-9]. However, further research is warranted to delve into the precise function of MARCKSL1 in ESCC. Therefore, this study aimed to scrutinize the distinctive expression pattern of MARCKSL1 in ESCC tissues, analyze its biological function from the cellular level, and further search for its differences. The possible mechanism of expression to find new molecular markers of EC.
Materials and Methods
Clinical sample data:
In this study, data were gathered from 74 ESCC patients who underwent surgery in various institutions and were treated at Nanjing Lishui District People's Hospital between January 2016 and July 2019. Among them were 64 males and 10 females (aged 46-72 y). The obtained EC tissues were all confirmed by postoperative pathology as ESCC; the paracancerous tissues were esophageal tissues more than 5 cm away from the tumor and pathologically confirmed as normal mucosal epithelial tissues. The staging of EC followed the 8th edition of the American Joint Committee on Cancer criteria, encompassing 40 cases categorized as stage I-II and 34 cases as stage III-IV. None of the patients had undergone prior radiotherapy, chemotherapy or immunotherapy. The study received ethical committee approval, and all participants provided informed consent.
Follow-up:
All patients with EC were followed up by outpatient visits, telephone calls, and WeChat contacts. Starting from the postoperative time, the deadline for follow-up is July 1st 2021, and the follow-up time ranges from 3 mo to 65 mo, with a median follow-up time of 33 mo. The term OS describes the interval from the moment of surgery until the last follow-up or death from any cause. The duration from the time after surgery until a confirmed disease progression or mortality from any reason is Progression-Free Survival (PFS).
Main reagents:
Eca-109 and KYSE30 EC cell lines were obtained from the American Type Culture Collection (ATCC). Rabbit anti-human MARCKSL1 antibody (ab184546), Horseradish Peroxidase (HRP)-labeled goat anti-rabbit antibody (ab205718), and β-actin (ab8227) were supplied by AbcamTM (Cambridge, United Kingdom (UK)). Diaminobenzidine (DAB) chromogen was procured through Shanghai Biyuntian Biotechnology Co., Ltd. Both Si-MARCKSL1 and si-NC were purchased from Ribo Company in Guangzhou, China. The Lipofectamine 2000TM transfection kit, OPTIMEM reagent, and Transwell chamber were all acquired from Thermo Fisher Scientific (United States of America (USA)). The Cell Counting Kit- 8 (CCK-8) reagent kit was purchased from MCE Company, USA.
Immunohistochemistry and interpretation of results:
Tissue sections from ESCC patients were dewaxed with xylene and dehydrated with an ethanol gradient. The antigen was then extracted using citrate buffer and blocked with 3 % Hydrogen Peroxide (H2O2). After washing the slides, the primary antibody to MARCKSL1 (1:100, ab184546, Abcam) was added dropwise, subsequently placed into incubation (4°/overnight). Post-washing slides again, drop HRP-conjugated secondary antibody (1:2000, ab205718, Abcam) and incubate for 20 min at 37°. The slides were cleaned with Phosphate Buffer Saline (PBS) before being counterstained with hematoxylin, processed with DAB, dehydrated with ethanol gradient, and coated with neutral gum.
The slice data was scanned into a file using the Panoramic MIDI slice scanner, and the slice results were then immediately recognized by the Quant Center program. Set strong positives as dark brown, moderate positives as brownish yellow, weak positives as light yellow, and negatives as blue nucleus. At the same time, the software can identify the corresponding stained area on the section and perform a Histochemistry score (H-score) on the positive percentage[10].
Reverse Transcription-Polymerase Chain Reaction (RT-PCR) detection of MARCKSL1 expression level:
The total Ribonucleic Acid (RNA) of ESCC cell lines Eca-109 and KYSE30 was extracted according to the operating instructions and analyzed for concentration and purity. On a PCR machine, the collected RNA was then reverse transcribed into complementary Deoxyribonucleic Acid (cDNA). ABI7500 FAST system detected the CT value of MARCKSL1 and Beta (β)-actin by q-PCR on cDNA.
MARCKSL1 primer; Forward: 5'-CAAGGGTGAAGGGGAGTCG-3 and Reverse: 5'-GACAGGCCGCTCAATTTGAAA-3. β-actin prime; Forward: 5'-GGACTTCGAGCAAGAGATGG-3 and Reverse: 5'-AGCACTGTGTTGGCGTACAG-3. And relative expression for MARCKSL1 was identified through 2-ΔΔCt technique using β-actin as an internal reference.
Cell transfection:
The Eca-109 and KYSE30 cells in good condition were resuspended, seeded in a 6-well plate, and cultured in the incubator. When the cells covered 90 % of the bottom of the dish, si-MARCKSL1 and si-NC were transfected into ESCC cells using Lipofectamine 2000 reagent. RNA was extracted to detect the transfection efficiency after continuing to culture in the incubator for 1 d.
Detection of cell proliferation activity by CCK-8:
After MARCKSL1 suppression, the proliferative activity for ESCC cultures was assessed using the CCK-8 assay. After being digested and centrifuged, the si-MARCKSL1 or si-NC-transfected cells were seeded in a 96-well plate at a density of 3000 cells per well. At 0 h, 24 h, 48 h, and 72 h, respectively, add 10 μl of CCK-8 reagent to each well, continue culturing in the incubator for 1 h and take it out. A multifunctional microplate reader detected the absorbance of the cells at 450 nm, and the proliferative activity of the cells was calculated.
Transwell assay to detect cell invasion and migration:
For the invasion experiment, 100 μl of serum-free medium containing Matrigel should be introduced into Transwell chamber-bottom in advance with subsequent placing into incubation for 6 h. The assays for Transwell migration have no requirement for Matrigel. Centrifuge the transfected cells, resuspended them in a serum-free medium and plant them in the upper chamber at a density of 2×104/ well. Place the upper chamber into the lower chamber gently, add a medium containing 20 % serum, and continue culturing for another 24 h in the incubator. The Transwell chamber was excluded, with cultures from the base within top chamber being observed under a microscope after being treated with formaldehyde and stained with crystal violet.
Western blot test to detect protein expression:
Radioimmunoprecipitation Assay (RIPA) lysate was used to extract the total protein from ESCC cells, and the Bicinchoninic Acid (BCA) technique was used to determine the protein concentration. After the protein was added to the precast gel, exposed to 60 min Sodium Dodecyl Sulphate-Polyacrylamide Gel Electrophoresis (SDS-PAGE) gel electrophoresis, membrane was subsequently placed for 2 h at a constant current of 300 mA in the transfer fluid, and the protein on the gel was transferred to the Polyvinylidene Difluoride (PVDF) membrane. After blocking the PVDF membrane, add MARCKSL1 (1:1000, ab184546, Abcam), Snail (1:1000, 3879, Cell Signaling Technology), Vimentin (1:1000, 5741, Cell Signaling Technology), N-cadherin (1:1000, 13116, Cell Signaling Technology), and E-cadherin (1:1000, 3195, Cell Signaling Technology), other primary antibody dilutions, and incubated overnight in a 4° refrigerator. Remove the PVDF membrane, wash it with PBS, add the secondary antibody dilution (1:5000, ab205718, Abcam), and incubate on a shaker for 1 h. Finally, the protein on the PVDF membrane was imaged by the Electroluminescence (ECL) chemical exposure method.
Dual Luciferase Reporter (DLR) assay:
A luciferase reporter plasmid containing the wildtype region and the mutant that microRNA (miR)-34a may bind to MARCKSL1 has been developed following the potential binding sites listed on the Star base website. When the cell density is appropriate, add 500 μl of serum-free medium to each well, and then add the corresponding plasmid (PGL3-MARCKSL1-3′ Untranslated Regions (3′ UTRs) or mutant-MARCKSL1-UTR, miRNA negative control or miR-34a mimic, empty plasmid and Lipofectamine 2000. Continue culturing for 1 d after transfection, remove the old culture medium, wash with PBS twice, and then add 100 μl PLB to each well. Lyse for at room temperature 15 min. Then add the corresponding amount of lysate and substrate luciferin. Use the instrument to record the value of the detected firefly luciferase. Add 100 μl of stop solution to the dish again. Use the device to collect the value of Renilla luciferase. Compare the assembled two groups of luciferase values calculated.
Gene Expression Omnibus (GEO) database analysis of MARCKSL1 expression:
The ESCC datasets GSE23400 and GSE45670 were acquired from the human tumor-associated gene expression repository, GEO (https://www.ncbi.nlm.nih.gov/gds/). The GSE23400 dataset encompassed 53 ESCC tissues alongside their corresponding normal esophageal mucosa tissues. MARCKSL1 expression was determined through GPL96 [HG-U133A] Affymetrix Human Genome U133A Array; GSE45670 included 28 ESCC tissues, and 10 normal esophageal mucosa tissues were analyzed for the expression of MARCKSL1 using the GPL570 [HG-U133_Plus_2] Affymetrix Human Genome U133 plus 2.0 Array.
Statistical methods:
The Statistical Package for the Social Sciences (SPSS) software (v.20.0) was used for data analysis. The expression of MARCKSL1 while clinic- pathological ESCC case profiles were compared through Chi-square (χ2) test. In order to produce the OS curve, Kaplan-Meier and Log-rank tests were applied. Prognostic variables were examined using Cox regression analysis. Measurements were determined to be significant at p<0.05.
Results and Discussion
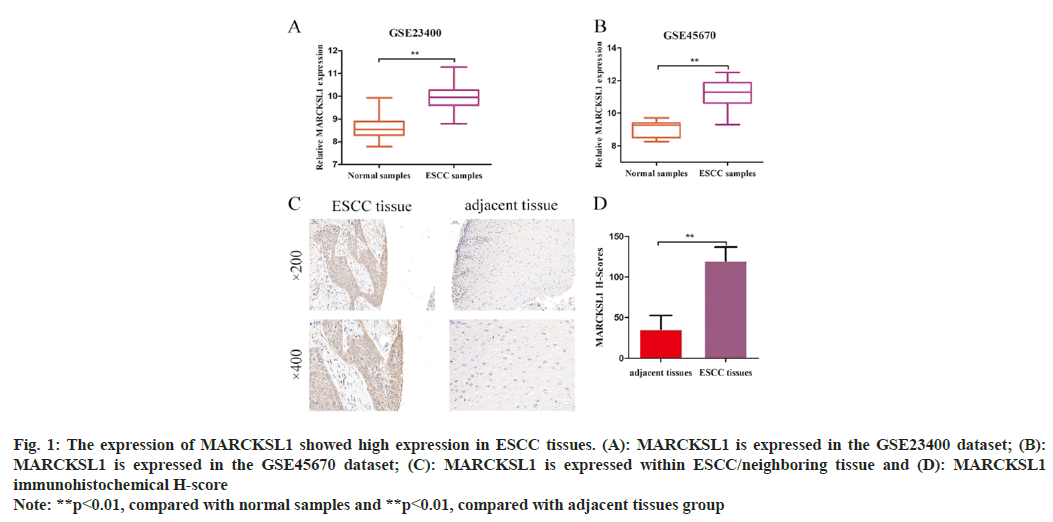
Online analysis was conducted through the GEO database (https://www.ncbi.nlm.nih.gov/gds/); we found that the expression of MARCKSL1 in the GSE23400 and GSE45670 datasets was 9.972±0.074 and 11.20±0.161, which were higher than the corresponding controls. The difference between the group expressions of 8.644±0.067 and 9.056±0.157 was statistically significant (t=13.25, p<0.01; t=7.472, p<0.01) as shown in fig. 1A and fig. 1B. Immunohistochemistry consequently identified MARCKSL1 expression level within ESCC tissue and its equivalent paracancerous tissues (fig. 1C). According to the results of immunohistochemistry H-score evaluation, MARCKSL1 exhibited a considerably higher H-score in malignant tissues than in paracancerous tissues (120.10±1.935 vs. 35.65±2.012, t=30.25, p<0.01, fig. 1D).
Fig. 1: The expression of MARCKSL1 showed high expression in ESCC tissues. (A): MARCKSL1 is expressed in the GSE23400 dataset; (B):
MARCKSL1 is expressed in the GSE45670 dataset; (C): MARCKSL1 is expressed within ESCC/neighboring tissue and (D): MARCKSL1
immunohistochemical H-score
Note: **p<0.01, compared with normal samples and **p<0.01, compared with adjacent tissues group
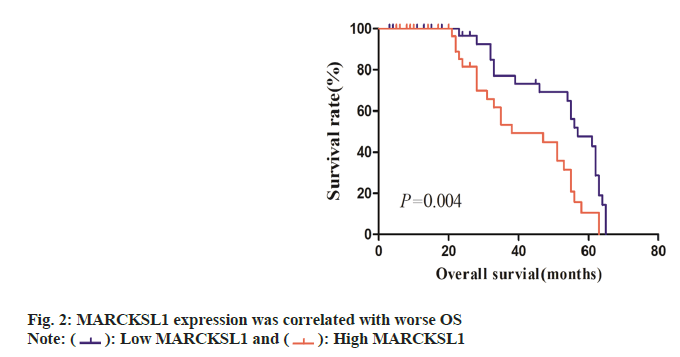
The ESCC tissues were segregated within MARCKSL1 high-expression group and MARCKSL1 low-expression group, depending upon immunohistochemistry scorings for MARCKSL1. Links across MARCKSL1 expression-profile characteristics and clinic-pathological profiles within ESCC cases were examined through χ2 test (Table 1). Such assessments revealed a strong correlation between the patient’s clinical Tumor, Nodes and Metastases (TNM) stage and the expression of MARCKSL1 (χ2=7.835, p<0.01). However, the expression of MARCKSL1 has no obvious relationship with the patient's age, gender, tumor size (T stage), and lymphatic metastasis (N stage). Kaplan-Meier analysis indicated that individuals displaying elevated MARCKSL1 expression experienced inferior OS compared to those with lower MARCKSL1 expression (p<0.05, fig. 2).
| Clinic pathological characteristics | n | MARCKSL1 low expression | MARCKSL1 high expression | χ2 | p |
|---|---|---|---|---|---|
| Age (years) | |||||
| ≤62 | 43 | 23 | 20 | 0.5 | 0.48 |
| >62 | 31 | 14 | 17 | ||
| Gender | |||||
| Female | 10 | 4 | 6 | 0.463 | 0.496 |
| Male | 64 | 33 | 31 | ||
| TNM stage | |||||
| I-II | 40 | 26 | 14 | 7.835 | 0.005 |
| III-IV | 34 | 11 | 23 | ||
| T stage | |||||
| T1-T2 | 11 | 4 | 7 | 0.961 | 0.327 |
| T3-T4 | 63 | 33 | 30 | ||
| N stage | |||||
| N0-N1 | 61 | 33 | 28 | 2.333 | 0.127 |
| N2-N3 | 13 | 4 | 9 | ||
Table 1: Correlation between MARCKSL1 Expression and Clinical Features of ESCC Patients
The Cox regression model subjected the Prognostic factors to univariate and multivariate analyses. Univariate analysis revealed a relationship between MARCKSL1 expression and OS (Hazard Ratios (HR)=2.355, 95 % Confidence Intervals (CI): 1.261-4.397, p=0.007; Table 2). Only the expression of MARCKSL1 was identified as a separate risk parameter concerning OS by multivariate analysis (HR=2.314, 95 % CI: 1.162- 4.610 and p=0.017).
| Variables | Univariate assessment | Multivariate assessment | ||||
|---|---|---|---|---|---|---|
| HR | 95 % CI | p | HR | 95 % CI | p | |
| Ages (≤62 vs. >62) | 1.145 | 0.628-2.090 | 0.659 | 1.196 | 0.245-2.940 | 0.603 |
| Gender (female vs. male) | 0.703 | 0.326-1.515 | 0.368 | 1.495 | 0.275-1.630 | 0.376 |
| TMN Stage(I-II vs. III-IV) | 1.276 | 0.671-2.425 | 0.457 | 0.986 | 0.432-2.249 | 0.974 |
| T Stage (T1-T2 vs. T3-T4 ) | 0.719 | 0.342-1.511 | 0.384 | 0.778 | 0.353-1.713 | 0.533 |
| N Stage (N0-N1 vs. N2-N3) | 1.45 | 0.643-3.273 | 0.371 | 0.977 | 0.325-2.939 | 0.967 |
| MARCKSL1 expression (low vs. high) | 2.355 | 1.261-4.397 | 0.007 | 2.314 | 1.162-4.610 | 0.017 |
Table 2: Shows Uni-/Multi-Variate Assessments for OS Predictors
The results of RT-PCR and protein studies demonstrated that Small interfering RNA (siRNA) could successfully decrease the expression of MARCKSL1 in cells by silencing MARCKSL1 expression within ESCC cellular cultures Eca- 109/KYSE30 (Eca-109: si-NC vs. si-MARCKSL1: 1.02±0.06 vs. 0.29±0.05, t=12.28, p<0.01; KYSE: si-NC vs. si-MARCKSL1: 1.00±0.021 vs. 0.303±0.08, t=12.59, p<0.01, fig. 3A and fig. 3B). Subsequently, the cellular proliferation ability of ESCC was identified through CCK-8 assessment. Such comparative assessment findings demonstrated knocking down MARCKSL1 reduced the ability of EC cells to proliferate as shown in fig. 3C.
Because MARCKSL1 is associated with the function of cell motility, our research team has continued to investigate the ability of ESCC cells to invade and migrate following various therapies. According to the Transwell invasion assay findings, the amount of Matrigel that invaded and transferred to the small chamber's outer membrane in the control group was Eca109: 114±6.85; KYSE30: 83.67±4.50, whereas the amount in the MARCKSL1 silencing group was Eca109: 44.67±5.43; and KYSE30: 30.33±4.50. This variation was statistically significant (p<0.01, fig. 4A and fig. 4B). Transwell migration assessment results revealed total amount concerning migrating cellular population within control group to be Eca109: 130.67±8.22; KYSE30: 86.67±4.92, while number of migrating cellular population within MARCKSL1 silencing cohort to be Eca109: 55±3.74; KYSE30: 44.67±4.62, with a statistically significant difference (p<0.01, fig. 4B-fig. 4D).
After suppressing MARCKSL1, the expression of key molecular markers of the Epithelial- Mesenchymal Transition (EMT) process was examined using Western blotting. The expression increased, while mesenchymal markers N-cadherin, vimentin and snail decreased significantly as shown in fig. 5.
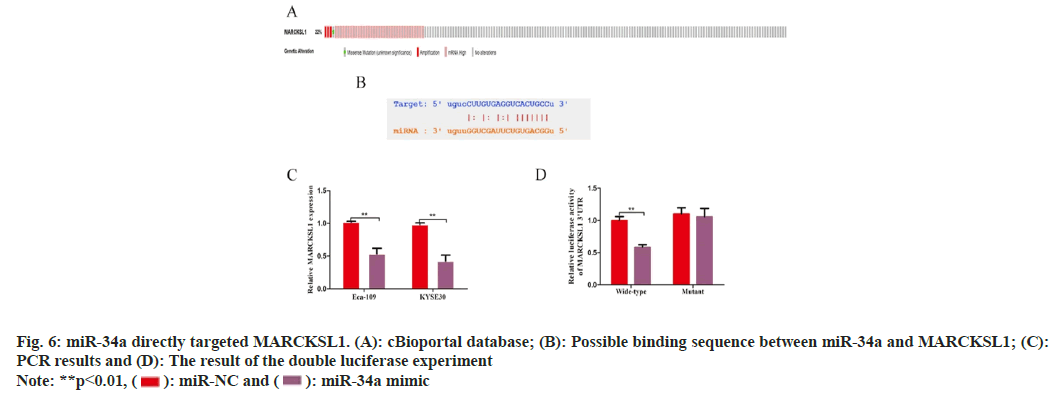
The cBioportal database (www.cbioportal.org) was utilized to investigate the causes of MARCKSL1 overexpression in EC tissues. The findings showed that the gene mutation or DNA copy number increase of MARCKSL1 in EC tissues was not obvious. Still, the mRNA level was significantly amplified, suggesting that MARCKSL1 Expression regulation mainly occurs at the post-transcriptional level (fig. 6A). Because of their distinctive architectures that can target and control genomic expression within post-transcription- level, non-coding RNAs, particularly micro non- coding RNAs (miRNAs), are now hot molecules in post-transcriptional modification research.
Using the Star base database to search for possible target miRNAs of MARCKSL1, miR-34a caught our attention. PCR studies were performed after overexpressing miR-34a using miR-34a analogs, and the outcomes indicated overexpressing miR-34a can strongly downregulate MARCKSL1 (KYSE: 1.017±0.012 vs. 0.533±0.069, t=9.688, p<0.05; Eca-109: 0.980±0.022 vs. 0.423±0.074, t=10.20, p<0.05, fig. 6B and fig. 6C). The dual luciferase assay was then used to investigate a potential targeted regulatory interaction between miR-34a and MARCKSL1. Overexpression of miR-34a substantially reduced the luciferase activity of MARCKSL1 wild type (0.997±0.051) in comparison to control cohort. Luciferase activity influence was not obvious (1.100±0.078 vs. 1.054±0.108, t=0.489, p>0.05, fig. 6C and fig. 6D). According to these findings, miR-34a could target and control the expression of MARCKSL1.
EC holds a significant position among the most common malignancies within the digestive tract in China. The pathological types include carcinosarcoma, small-cell undifferentiated carcinoma, adenosquamous cell carcinoma, adenocarcinoma and squamous cell carcinoma[11]. In China, ESCC accounts for more than 90 % of the pathological type of EC, and the incidence clearly varies by area. The northern Jiangsu area is a high- incidence area for EC in China[12]. Although the current treatment technology for EC continues to improve, many patients with ESCC are discovered late, thus missing the best time for surgery. Even though some patients have undergone surgery, the postoperative tumor is still vulnerable to metastasis and recurrence, and the death rate for those suffering from ESCC is still high. Therefore, looking for molecular markers related to ESCC, exploring its possible molecular mechanism of action, and looking for potential molecular targeted drugs are currently hot issues in ESCC research[13]. Using proteomic technologies, Karagoz et al.[14] examined the tissues from 99 cases of ESCC and the matched normal esophageal mucosa tissues in 2016. They discovered 51 differential genes, of which 21 had high expression, and 30 had low expression. MARCKSL1 was highly expressed in ESCC for the first time.
In this study, we also used the data analysis from the online GEO database to find that MARCKSL1 was highly expressed in the two EC datasets. Then our research group used an immunohistochemically technique to detect the expression of MARCKSL1 in 74 cases of ESCC tissues collected and found that MARCKSL1 corroborated to online-database-derived dataset outcomes and MARCKSL1 was significantly highly expressed in cancer tissues. Our investigation did not end there; we also investigated the relationship between MARCKSL1 expression and the clinic pathological characteristics of patients. We found that the high expression of MARCKSL1 was related to the later clinical stage, while other factors (age, gender, T stage, N stage) are irrelevant. Among them, MARCKSL1 expression has no obvious correlation with tumor size (T stage) and lymphatic metastasis (N stage), which aroused our thinking.
In a recent study focused on ESCC, Zhao et al.[15] discovered that individuals with elevated MARCKSL1 expression exhibited a notably higher rate of lymph node metastasis compared to those with lower expression levels. Moreover, the study revealed an association between high MARCKSL1 expression and poorer tumor differentiation. The entire sample size is too small to investigate the causes of the discrepancies between the experiment's findings and the study’s conclusions, and the data cannot show a connection between patient’s clinical stages and MARCKSL1 expression. As early as 2012, some researchers found that higher the expression of MARCKSL1 in patients with lymph node-negative breast cancer, the greater the risk of distant metastasis and significantly shortened survival time[16]. Therefore, in follow-up experiments, this project needs to increase the number of samples and collect tissues from patients with distant metastasis of ESCC to explain the association between the expression of MARCKSL1 and distant metastasis of patients with ESCC.
In both lung adenocarcinoma and breast cancer, it has been reported that high expression of MARCKSL1 indicates poor prognosis[17]. The Kaplan-Meier method was utilized in this study to examine the association between MARCKSL1 expression and OS in ESCC patients. The findings demonstrated that shorter OS was predicted by greater MARCKSL1 expression. Only the expression of MARCKSL1 was identified as separate predictive biomarker for OS within ESCC cases through Cox regression model, uni-/ multi-variate assessments of pertinent covariates. MARCKSL1 has a high clinical guiding value, possibly serving as a separate prognostic biomarker for ESCC cases, even if the sample size of this study is rather limited compared to the current literature.
The protein MARCKSL1, belonging to the myristoylated alanine-rich C-kinase substrate family, regulates actin movement and the development of the cytoskeleton[18]. According to certain studies, MARCKSL1 can promote lung adenocarcinoma cell’s EMT, promoting tumor cell proliferative, invasive and migrative properties[16]. A separate investigation revealed dephosphorylation of MARCKSL1 in tumor tissue increased the mobility of actin and induced tumor cell migration[19]. As a result, we hypothesize that MARCKSL1 may also control the invasion and migration of cells with ESCC. First, we discovered that MARCKSL1 silencing may substantially decrease the growth of EC cells using CCK-8 detection. The results of subsequent Transwell invasion and migration assays demonstrated that lowering the expression of MARCKSL1 might significantly lessen the ability of tumor cells to invade and migrate.
Further downstream mechanism exploration experiments confirmed that silencing MARCKSL1 could reverse the EMT process of EC cells. This study is the first to prove at the cell level in vitro that MARCKSL1 can play an important role as a "tumor-promoting factor" in ESCC and accelerate the malignant process of ESCC. However, there are limitations in this study at the cellular level. It does not yet have a thorough understanding of MARCKSL1's mechanism of action; it merely explains the biological function of MARCKSL1. The molecular mechanism of MARCKSL1 in advancing EC will be clarified in the follow- up experiments using protein spectrum, electron microscopy, and other procedures. This work will also serve as a crucial theoretical foundation for the development of MARCKSL1-targeting drugs in the future.
In this experiment, we analyzed the influence of MARCKSL1 on downstream related genes by protein experiment and the possible reasons for the high expression of MARCKSL1 in EC using the bioinformatics website. The findings demonstrated that MARCKSL1 DNA levels did not vary noticeably. However, the mRNA level increased abnormally, indicating that improper post-transcriptional regulation may cause the dysregulation of MARCKSL1 expression in EC. Currently, a large number of literature data have confirmed that there is a class of endogenous non-coding small molecule single-stranded RNA, which is composed of 18-25 nucleotides, through complete or incomplete complementary combination with the 3'-UTRs of the target gene, in the post-transcriptional level causes the degradation or translational repression of target gene mRNA[20,21]. In the study of EC, miRNA can be employed as a molecular target that serves a crucial regulatory function in the tumor growth of EC, in addition to being used as a molecular marker to detect cancers early and assess the prognosis of patients[22,23]. Therefore, we speculate that it is likely that the dysfunction of key tumor suppressor miRNAs in EC leads to the uncontrolled expression of MARCKSL1, which leads to the malignant transformation of tumor cells. As a well-known tumor suppressor function miRNA, miR-34a is lowly expressed in EC tissue, which could thwart proliferative, invasive, and migrative properties for EC cells[24,25].
This investigation employed the Star base website to identify a potential binding link between miR-34a and MARCKSL1, and we then conducted further related tests to support this finding. The low expression of the "tumor suppressor" miR-34a in EC tissue results in the weakening of the post-transcriptional inhibition of MARCKSL1 and the increase of MARCKSL1 expression level, which promotes the EMT of EC cells and aggravates the tumorigenesis of EC cells-vicious process. The existing experimental data strongly confirm this. Many known post-transcriptional regulations exist, such as splicing of mRNA precursors, m6A methylation, etc. This study selected miRNA, a hotspot molecule currently studied, to find the mechanism of post-transcriptional regulation of MARCKSL1 from the perspective of non- coding RNA. It is still limited, and the relevant experimental data still needs to be continuously improved in the later stage to clarify the upstream mechanism of MARCKSL1.
In conclusion, our work is the first to identify differentially elevated MARCKSL1 expression in ESCC, and its expression level is directly correlated with the advanced stage of the disease and the patients' bad prognosis. Additional COX univariate and multivariate analysis supported that MARCKSL1 can function as an individual risk factor for ESCC patients. The findings at the in vitro cell level suggested that MARCKSL1 contributes significantly as a "tumor-promoting factor" to the malignant development of ESCC. In addition, this work is the first to explain the causes of MARCKSL1's differential expression in tumor research. It is found that the "tumor suppressor" miR-34a can inhibit the expression of MARCKSL1 at the post-transcriptional level to develop a molecular target for MARCKSL1 from a molecular perspective and provide important mechanistic clues to drugs.
Funding:
This work was supported by Nanjing Medical and Health Science Research Project (YKK20214).
Author’s contributions:
Zhang Lei proposed and designed the research top-ic and wrote the first draft of the thesis; Zhang Renmin worked on research topic data analysis; Hongping Zhou on data acquisition; Du Mingyu has revised the paper; and Liu Jiangang has done the review and guidance.
Acknowledgements:
Zhang Lei and Hou Ying have contributed equally to this work.
Conflict of interests:
The authors declared no conflict of interests.
References
- Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021;71(3):209-49.
[Crossref] [Google Scholar] [PubMed]
- Roy Chattopadhyay N, Chakrabarti S, Chatterjee K, Deb Roy S, Kumar Sahu S, Reddy RR, et al. Histocompatibility locus antigens regions contribute to the ethnicity bias of Epstein-Barr virus-associated nasopharyngeal carcinoma in higher-incidence populations. Scandinavian J Immunol 2019;90(4):e12796.
[Crossref] [Google Scholar] [PubMed]
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68(6):394-424.
[Crossref] [Google Scholar] [PubMed]
- van Itallie CM, Tietgens AJ, Aponte A, Gucek M, Cartagena-Rivera AX, Chadwick RS, et al. MARCKS-related protein regulates cytoskeletal organization at cell–cell and cell–substrate contacts in epithelial cells. J Cell Sci 2018;131(3):jcs210237.
[Crossref] [Google Scholar] [PubMed]
- Aderem A. The MARCKS family of protein kinase-C substrates. Biochem Soc Trans 1995;23(3):587-91.
[Crossref] [Google Scholar] [PubMed]
- El Amri M, Fitzgerald U, Schlosser G. MARCKS and MARCKS-like proteins in development and regeneration. J Biomed Sci 2018;25(1):43.
[Crossref] [Google Scholar] [PubMed]
- Li F, Zhao J, Wang L, Chi Y, Huang X, Liu W. METTL14-mediated miR-30c-1-3p maturation represses the progression of lung cancer via regulation of MARCKSL1 expression. Mol Biotechnol 2022;64(2):199-212.
[Crossref] [Google Scholar] [PubMed]
- Jiang M, Qi F, Zhang K, Zhang X, Ma J, Xia S, et al. MARCKSL1–2 reverses docetaxel-resistance of lung adenocarcinoma cells by recruiting SUZ12 to suppress HDAC1 and elevate miR-200b. Mol Cancer 2022;21(1):150.
[Crossref] [Google Scholar] [PubMed]
- Liu Y, Liu H, Bian Q. Identification of potential biomarkers associated with basal cell carcinoma. Biomed Res Int 2020;2020:2073690.
[Crossref] [Google Scholar] [PubMed]
- Du M, Peng Y, Li Y, Sun W, Zhu H, Wu J, et al. MYC-activated RNA N6-methyladenosine reader IGF2BP3 promotes cell proliferation and metastasis in nasopharyngeal carcinoma. Cell Death Discov 2022;8(1):53.
[Crossref] [Google Scholar] [PubMed]
- Zeng H, Zheng R, Zhang S, Zuo T, Xia C, Zou X, et al. Esophageal cancer statistics in China, 2011: Estimates based on 177 cancer registries. Thoracic Cancer 2016;7(2):232-7.
[Crossref] [Google Scholar] [PubMed]
- Pan R, Zhu M, Yu C, Lv J, Guo Y, Bian Z, et al. Cancer incidence and mortality: A cohort study in China, 2008–2013. Int J Cancer 2017;141(7):1315-23.
[Crossref] [Google Scholar] [PubMed]
- Zhan J, Wang W, Tang Y, Zhou N, Jiang D. Development of a prognostic signature for esophageal cancer based on a novel 7-DNA damage repair genes signature. Biocell 2022;46(12):2601-13.
- Karagoz K, L Lehman H, B Stairs D, Sinha R, Y Arga K. Proteomic and metabolic signatures of esophageal squamous cell carcinoma. Curr Cancer Drug Targets 2016;16(8):721-36.
[Google Scholar] [PubMed]
- Zhao Y, Xie X, Tian L, Liu F, Sun Y, Lu H, et al. MARCKSL1 interacted with F-actin to promote esophageal squamous cell carcinoma mobility by modulating the formation of invadopodia. Cancer Med 2023;12(3):3299-312.
[Crossref] [Google Scholar] [PubMed]
- Egeland NG, Austdal M, van Diermen-Hidle B, Rewcastle E, Gudlaugsson EG, Baak JP, et al. Validation study of MARCKSL1 as a prognostic factor in lymph node-negative breast cancer patients. Plos One 2019;14(3):e0212527.
[Crossref] [Google Scholar] [PubMed]
- Liang W, Gao R, Yang M, Wang X, Cheng K, Shi X, et al. MARCKSL1 promotes the proliferation, migration and invasion of lung adenocarcinoma cells. Oncol Lett 2020;19(3):2272-80.
[Crossref] [Google Scholar] [PubMed]
- Kondrychyn I, Kelly DJ, Carretero NT, Nomori A, Kato K, Chong J, et al. Marcksl1 modulates endothelial cell mechanoresponse to haemodynamic forces to control blood vessel shape and size. Nat Commun 2020;11(1):5476.
[Crossref] [Google Scholar] [PubMed]
- Bjorkblom B, Padzik A, Mohammad H, Westerlund N, Komulainen E, Hollos P, et al. c-Jun N-terminal kinase phosphorylation of MARCKSL1 determines actin stability and migration in neurons and in cancer cells. Mol Cell Biol 2012;32(17):3513-26.
[Crossref] [Google Scholar] [PubMed]
- Fabian MR, Sonenberg N. The mechanics of miRNA-mediated gene silencing: A look under the hood of miRISC. Nat Struct Mol Biol 2012;19(6):586-93.
[Crossref] [Google Scholar] [PubMed]
- Hill M, Tran N. miRNA interplay: Mechanisms and consequences in cancer. Dis Models Mech 2021;14(4):dmm047662.
[Crossref] [Google Scholar] [PubMed]
- Xue ST, Zheng B, Cao SQ, Ding JC, Hu GS, Liu W, et al. Long non-coding RNA LINC00680 functions as a ceRNA to promote esophageal squamous cell carcinoma progression through the miR-423-5p/PAK6 axis. Mol Cancer 2022;21(1):69.
[Crossref] [Google Scholar] [PubMed]
- Miyoshi J, Zhu Z, Luo A, Toden S, Zhou X, Izumi D, et al. A microRNA-based liquid biopsy signature for the early detection of esophageal squamous cell carcinoma: A retrospective, prospective and multicenter study. Mol Cancer 2022;21(1):1-3.
[Crossref] [Google Scholar] [PubMed]
- Ye Z, Xie T, Yan F, Wang L, Fang J, Wang Z, et al. miR-34a reverses radiation resistance on ECA-109 cells by inhibiting PI3K/AKT/mTOR signal pathway through downregulating the expression of SIRT1. Int J Radiation Biol 2021;97(4):452-63.
[Crossref] [Google Scholar] [PubMed]
- Wang X, Zhao Y, Lu Q, Fei X, Lu C, Li C, et al. miR-34a-5p inhibits proliferation, migration, invasion and epithelial-mesenchymal transition in esophageal squamous cell carcinoma by targeting LEF1 and inactivation of the hippo-YAP1/TAZ signaling pathway. J Cancer 2020;11(10):3072.
[Crossref] [Google Scholar] [PubMed]
 High MARCKSL1
High MARCKSL1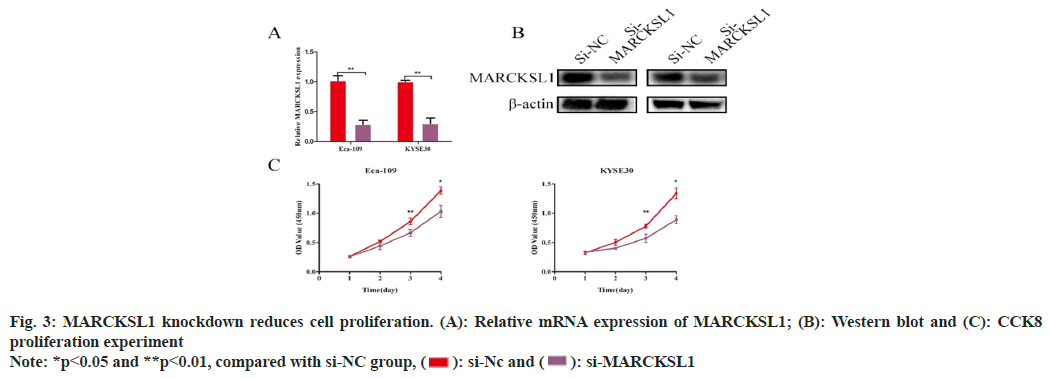
 si-MARCKSL1
si-MARCKSL1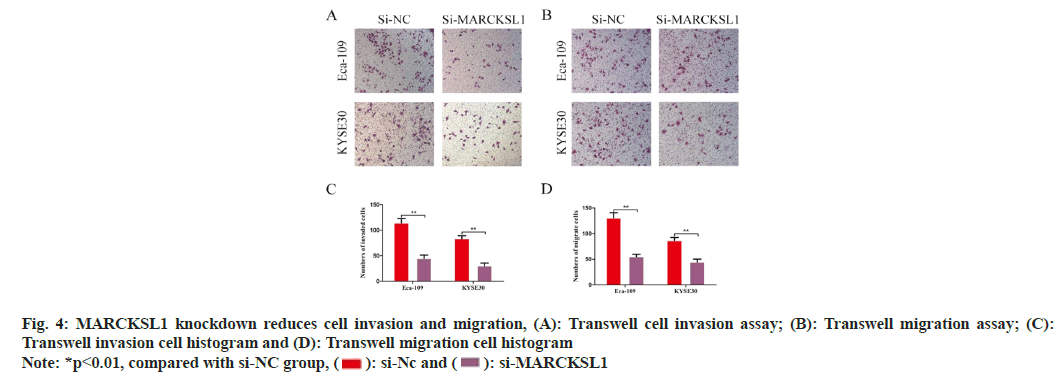
 si-MARCKSL1
si-MARCKSL1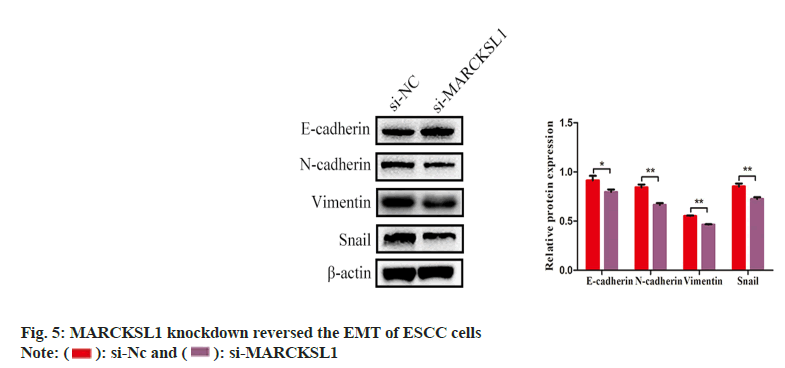
 miR-34a mimic
miR-34a mimic



