- *Corresponding Author:
- Y. Dong
Department of Traditional Chinese Medicine, College of Pharmacy and Biotechnology, Tianjin Medical College, Tianjin 300222, China
E-mail: zxsd411@126.com
| Date of Received | 10 July 2021 |
| Date of Revision | 19 August 2022 |
| Date of Acceptance | 03 February 2023 |
| Indian J Pharm Sci 2023;85(1):233-240 |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
Qing’E pills are a classical formula in traditional Chinese medicine that has the effect of tonifying the kidneys and strengthening the bones, first appearing in the tonic formulae in prescriptions of peaceful benevolent dispensary (10th century CE). To preliminarily elucidate the potential mechanisms of Qing’E pills in the treatment of postmenopausal osteoporosis based on network pharmacology. The principal components and corresponding protein targets of Qing’E pills were searched on traditional Chinese medicine systems pharmacology, bioinformatics analysis tool for molecular mechanism-traditional Chinese medicine, SymMap and encyclopedia of traditional Chinese medicine and the compound-target network was constructed by Cytoscape 3.9.1. The targets of postmenopausal osteoporosis were searched on online mendelian inheritance in man, GeneCards, histidine triad nucleotide binding protein and DisGeNET databases. The intersection of compound target and disease target was obtained and the coincidence target was imported into the search tool for retrieval of interacting genes database to construct a protein-protein interaction network. We further performed gene ontology and Kyoto encyclopedia of genes and genomes enrichment analyses on the targets. The 33 active compounds in Qing’E pills corresponded to 433 targets. 1647 targets were associated with postmenopausal osteoporosis disease. The active compounds with high degree values were quercetin, stearic acid and marine. The proteins with higher values in the protein-protein interaction network were mitogenactivated protein kinase 1, protein kinase B, phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha, phosphoinositide-3-kinase regulatory 1, amyloid beta precursor protein, Src proto-oncogene, nonreceptor tyrosine kinase, mitogen-activated protein kinase 8, heat shock protein 90 alpha family class A member 1, epidermal growth factor receptor, and Janus kinase 2. Gene ontology and Kyoto encyclopedia of genes and genomes analysis showed that Qing’E pills treatment of postmenopausal osteoporosis involves signaling pathways that mainly include the tumor necrosis factor signaling pathway, osteoclast differentiation, Wnt signaling pathway and estrogen signaling pathway. We revealed the active ingredients and potential molecular mechanism of Qing’E pills in the treatment of postmenopausal osteoporosis and provided a reference for subsequent basic research
Keywords
Qing’E pills, network pharmacology, traditional Chinese medicine, postmenopausal osteoporosis
Postmenopausal Osteoporosis (PMO) is a metabolic disorder caused by estrogen deficiency and characterized by reduced Bone Mineral Density (BMD) and deterioration of bone microarchitecture[1,2]. Previous studies have shown a high incidence of PMO[3]. Patients with PMO have an increased susceptibility to hypo traumatic fractures, which are often accompanied by severe painful manifestations after fracture, which severely reduces the quality of life of patients. According to epidemiological studies, PMO affects half the women in the 6th and 7th decade of life. More than 57 % of women at risk in 27 European Union countries (EU27) do not receive bone-specific treatment[4,5]. As the population ages, aging comorbidities such as PMO pose a huge economic and health burden on society[6], and have become a growing public health problem[7]. The International Osteoporosis Foundation (IOF) and the European Federation of Pharmaceutical Industries Association (EFPIA) report that the cost of PMO treatment will increase by 25 % by 2025. The search for rational and effective treatments has become an important issue for clinicians to address.
In the late 1980s, estrogen replacement, perhaps calcitonin, along with calcium and vitamin D supplementation were used as the main treatment strategy for PMO[7]. In 2017, selective estrogen receptor modulators[8] (raloxifene), bisphosphonates[9] (alendronate, risedronate, ibandronate and zoledronic acid), human anti-Receptor Activator of Nuclear Factor Kappa-Β Ligand (RANKL) antibody[10] (ANKL; denosumab) and parathyroid hormone analog[11] (teriparatide) are included in the range of therapeutic agents. Despite significant advances in the development of drugs for the prevention and treatment of fractures, there are still major challenges in implementing treatment such as adverse effects, for example, osteonecrosis of the jaw and atypical femoral fractures have been reported following teriparatide treatment[3].
Qing’E Pills (QEP) is a mixture of Traditional Chinese Medicine (TCM) consisting of Eucommiae Cortex (Eucommia ulmoides (E. ulmoides), Psoraleae Fructus (Psoralea corylifolia), Juglandis Semen (Juglans regia (J. regia)) and Garlic Rhizoma[12-14]. However, the mechanism by which QEP improves the balance of bone metabolism in PMO patients is not known. As an emerging discipline, network pharmacology can accelerate the clinical translation of precise and effective therapeutic interventions[15]. The targets of the main active components of QEP and the targets of sepsis were collected by using the network pharmacology technology. The QEP component target network was built, as well as the Protein-Protein Interaction (PPI) network of chemical and disease interaction targets. The signaling pathways of biological function and efficacy were then investigated using Gene Ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis.
Materials and Methods
Data sources:
Procurement and screening of active compounds in QEP: The compounds and the corresponding targets of Eucommiae Cortex (E. ulmoides), Psoralea Fructus (Psoralea corylifolia), Juglandis Semen (J. regia) in QEP were obtained from the following databases, namely Traditional Chinese Medicine Systems Pharmacology (TCMSP) (tcmsp)[16], Bioinformatics Analysis Tool for Molecular Mechanism (BATMAN)-TCM (batman-tcm) [17], SymMap (http://www.symmap.org/)[18] and Encyclopedia of Traditional Chinese Medicine (ETCM) (http://www.tcmip.cn/ETCM/index.php/ Home/Index/)[19]. Oral Bioavailability (OB) refers to the amount of medicine that enters the circulatory system. Drug similarity (DL) is the similarity between a chemical and a known drug[16]. In the TCMSP system, OB ≥30 % and DL ≥0.18 were set as the filters for retrieving pharmacokinetic data.
Target prediction for the active compounds of QEP: The 2-dimensional structure files of the active ingredients were obtained by accessing the organic small molecule bioactivity database (https://pubchem.ncbi.nlm.nih.gov/, https:/pubchem.ncbi.nlm.nih.gov/)[20]. The structures of the active ingredients were cross-validated for ligand similarity and analytically aligned via the SwissTargetPrediction platform (http://www.swisstargetprediction.ch/)[21] to complement the TCMSP system with compound components not included and their corresponding targets of action. The active ingredient targets of the QEP were screened using the criterion of "probability value >0". The top 10 related targets (related target proteins) of each compound were selected and uploaded to the PharmMapper database (http://lilab.ecust.edu.cn/)[22]. The "Homo sapiens" was selected for the identification of potential target proteins (hereafter referred to as "targets").
Target prediction: Searching through GeneCards database (https://www.genecards.org/)[23], Online Mendelian Inheritance in Man (OMIM) database (https://omim.org/)[24], DisGeNET database (https://www.disgenet.org/)[25] for PMO-related target retrieval. The potential targets for PMO treatment with QEP were mapped through the ImageGP platform (http://www.ehbio.com/ImageGP/).
PPI network construction:
To further study the interaction targets, the protein identifier or protein sequence provided by Search Tool for Retrieval of Interacting Genes (STRING) (https://string-db.org/)[26] was used for analysis. Select “Homo sapiens” in the organism column and the minimum required interaction score is “highest confidence (0.9)”. Then input the correlation information between the targets into Cytoscape 3.7.2 for visualization and construct a PPI network.
Functional enrichment analysis of QEP'S therapeutic targets for PMO:
The Meta scape system (http://metascape.org/, version 3.0) was used to study the biological function of potential targets in PMO[27]. We collected terms with p<0.01, a minimum count of 3 and an enrichment factor >1.5. The Cytoscape 3.8.0 software package (Boston, MA, United States of America (USA)) was used to explain the complicated relationships among compounds, genes, pathways and diseases.
Results and Discussion
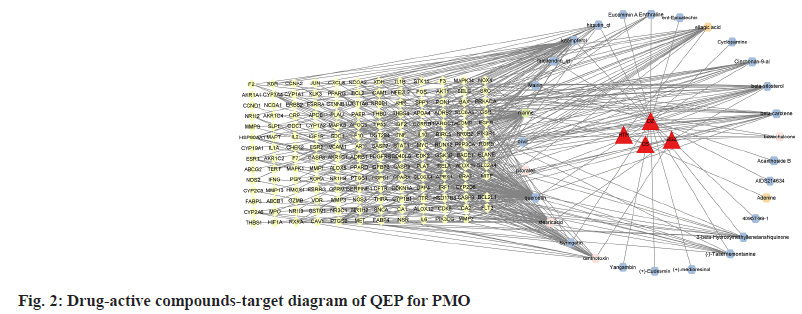
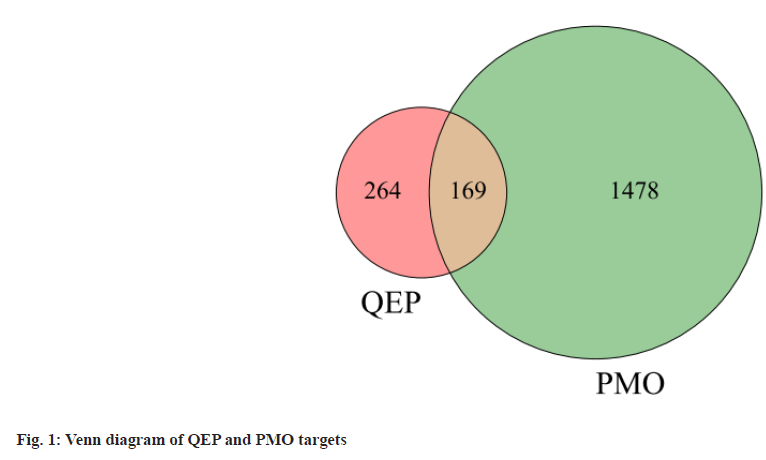
After removing duplications of the initial ingredients, a total of 33 initial chemical components of QEP were obtained through the TCMSP system and the SwissTarget system retrieval and manual retrieval. E. ulmoides Oliv. contains 25 active compounds, Psoralea corylifolia L contains 5 active compounds, J. regia L. and Allium sativum L include 2, 1 active compounds respectively. The 433 targets of 33 active ingredients were selected as common targets with PMO (1647 targets) (fig. 1) to construct a drug-active ingredient-target network for QEP for PMO (fig. 2). The active compounds with high QEP median values are shown in Table 1.
| Type | Name | Degree | Stress | Radiality |
|---|---|---|---|---|
| Eucommia ulmoides Oliv. | Quercetin | 127 | 1394918 | 0.99 |
| Psoralea corylifolia L | Stearic acid | 115 | 638198 | 0.99 |
| Allium sativum L | Marine | 101 | 889700 | 0.99 |
| Psoralea corylifolia L | Xanthotoxin | 81 | 1101216 | 0.99 |
| Juglans regia L. | Ellagic acid | 58 | 781208 | 0.989 |
| Eucommia ulmoides Oliv. | Kaempferol | 54 | 536792 | 0.99 |
| Eucommia ulmoides Oliv. | Beta-sitosterol | 36 | 251990 | 0.99 |
| Eucommia ulmoides Oliv. | Cinchonan-9-al | 30 | 204364 | 0.98 |
| Eucommia ulmoides Oliv. | Tabernemontanine | 28 | 133726 | 0.99 |
| Eucommia ulmoides Oliv. | Beta-carotene | 18 | 67074 | 0.98 |
Table 1: Active Compounds with High Moderate QEP Values
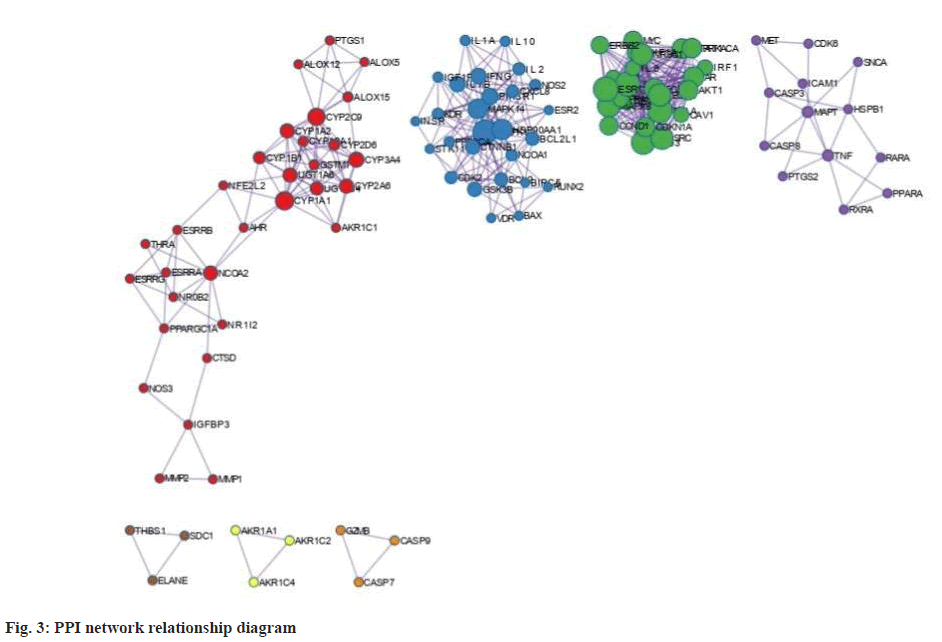
We performed PPI network construction for the QEP-PMO common target based on Cytoscape software results (fig. 3). If the network contains between 3 and 500 proteins, the molecular complex detection algorithm was applied to identify densely connected network components. The nodes in fig. 3 represent target proteins and the edges represent two proteins connected, the larger the node, the larger the position in the network. The top 10 nodes are Mitogen-Activated Protein Kinase (MAPK) 1, Protein Kinase B (AKT1), Phosphatidylinositol-4, 5-Bisphosphate 3-Kinase Catalytic Subunit Alpha (PIK3CA), Phosphoinositide-3-Kinase Regulatory 1 (PIK3R1), Amyloid Beta Precursor Protein (APP), SRC Proto-Oncogene, Non-Receptor Tyrosine Kinase (SRC), MAPK8, Heat Shock Protein 90 Alpha Family Class A Member 1 (HSP90AA1), Epidermal Growth Factor Receptor (EGFR) and Janus Kinase 2 (JAK2), which are presumed to be important in the QEP treatment of PMO.
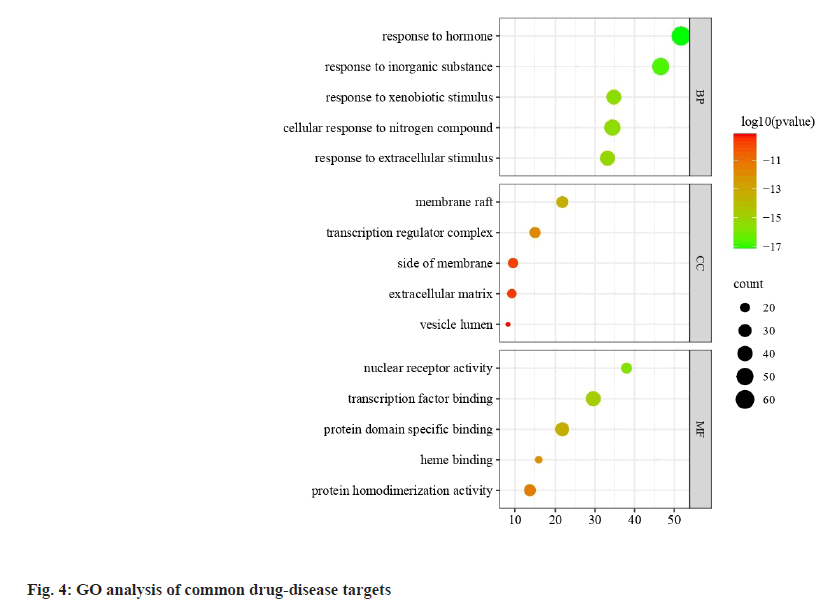
During the GO enrichment analysis of the targets, 150 biological processes, 28 cellular components and 38 molecular functions were identified, indicating that the biological functions of QEP interfering with PMO were mainly in the areas of estrogen regulation, regulation of bone mineralization and regulation of factor binding. We screened 15 biological functions, cellular components and molecular functional pathways that had a significant impact in GO analysis for visualization (fig. 4), which related to positive regulation of transcription from nuclear receptor activity, transcription factor binding, protein domain specific binding, heme binding, protein homodimerization activity, response to hormone, response to inorganic substance, response to xenobiotic stimulus, cellular response to nitrogen compound, response to extracellular stimulus, membrane raft, transcription regulator complex, side of membrane and extracellular matrix.
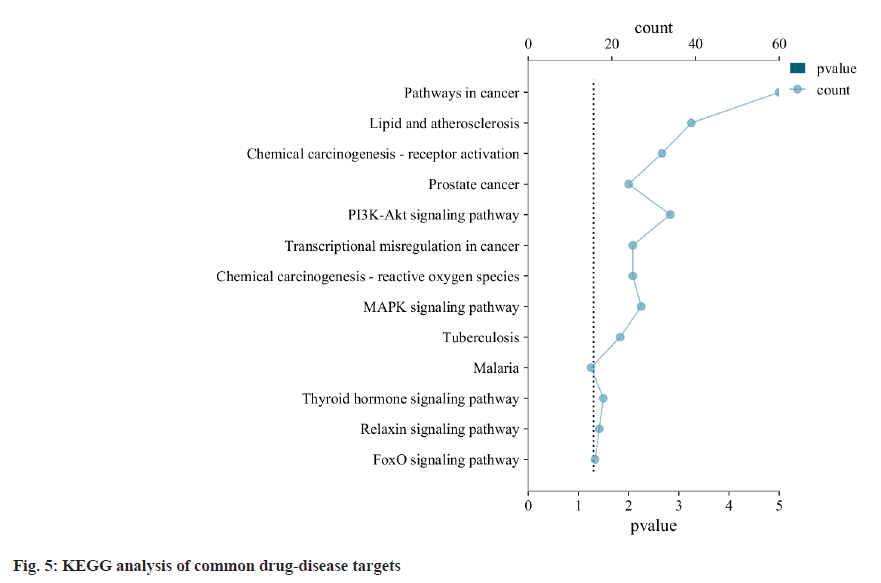
In order to explore the signal pathway mechanism of QEP in the treatment of PMO, we performed KEGG enrichment analysis (fig. 5). We found that the targets were mainly related to 36 key pathways, including the calcium signaling pathway, the estrogen signaling pathway and the cyclic Guanosine Monophosphate-activates Protein Kinase G (cGMP-PKG) signaling pathway. At the same time the pathways in cancer, lipid and atherosclerosis, chemical carcinogenesis-receptor activation, prostate cancer, Phosphoinositide-3-Kinase (PI3K)-Akt signaling pathway, transcriptional misregulation in cancer, chemical carcinogenesis-reactive oxygen species, MAPK signaling pathway and tuberculosis were of great importance in the QEP efficacy research.
As a recommended remedy in the PMO Chinese Medicine Guideline, QEP can promote bone calcium absorption and thus regulate the balance between calcium and phosphorus. Pharmacological studies have shown that QEP can inhibit osteoclast activity, maintain stable bone metabolism and regulate estrogen metabolism, thereby preventing and treating PMO. However, there is a lack of research on the mechanism of QEP efficacy. We investigate the potential mechanism of QEP in the treatment of PMO through a predictive study, with a view to providing directional options for the clinical management of PMO.
Thirty-three chemical components were collected from the TCMSP database, which can act in combination with other active components at multiple targets in the network. Six of the ten active ingredients with high degree values were from Eucommia tulipifera (E. tulipifera) and the associated components of E. tulipifera, such as quercetin and kaempferol, were connected to drug-disease common targets with high density, suggesting that E. tulipifera may play a good role in the treatment of PMO. The lignans and phenylpropanoids in Eucommia are immune-enhancing, and the sterol in bone marrow is used as a raw material for the production of vitamin D3, so it is assumed that QEP may prevent PMO exacerbation to some extent[28-34]. The topological analysis of the main compounds and the major intersecting targets revealed the highest binding of the target CYP3A4 and hirsutine. CYP3A4 (cytochrome P450 3A4) is a hemoglobin thioredoxin that completes a variety of oxidative reactions that affect bone loss[35]. Evellyn et al. showed that stimulation of the estrogen signaling pathway could regulate the induction of osteogenesis. Evellyn et al. also found that stimulation of the estrogen signaling pathway could regulate the body's secretion of estrogen. It suggested that QEP may improve bone loss due to estrogen reduction by modulating the calcium signaling pathway, estrogen signaling pathway and cGMP-PKG signaling pathway, which are the key effector pathways[36].
Among the high value targets, MAPK1 and MAKE8 are members of the MAPK signaling pathway, which intervenes in osteoblast apoptosis via the mitochondrial pathway[37-42]. AKT1 was a member of the AKT gene family[43]. AKT1 was found to have limited skeletal development in rats treated with AKT1 knockout, suggesting that AKTI may play a role in promoting osteogenesis[44]. AKT1, PIK3CA and PIK3R1 are members of the PI3K-Akt signaling pathway, through which AKTI plays an anti-inflammatory role in the metabolic process of the body[45-49]. AK2, a member of the JAK/signal transducer and activator of transcription signaling pathway, is involved in the development of bone destruction through inflammatory responses[50]. The key targets, including AKT1, PIK3CA and PIK3R1 are presumed by GO enrichment analysis to function mainly at the plasma membrane, membrane rafts and nucleus and to perform molecular functions such as kinase activity, protein binding and protein kinase binding through positive regulation of nitric oxide biosynthesis, signal transduction, protein phosphorylation and other biological processes. Although the cancer signaling pathway was ranked first in the enrichment analysis of this study, there are few studies on this pathway in relation to PMO and the mechanisms mediating it need to be investigated in subsequent studies.
There are still some limitations in this study. We only used modern bioinformatics methods to predict the role of QEP in PMO through network pharmacology. However, the current network information technology needs to be further improved and the accuracy and timeliness of the database data needs to be scientifically validated. Also, unidentified and undocumented compounds or targets may not be included in our analysis. Secondly, the bioactive components in this study are not fully representative of QEP. We believe that the pharmacodynamics and molecular biology experiments should be considered to further validate our results.
We conducted a preliminary exploration of the main chemical components and potential mechanisms of QEP for the treatment of PMO based on network pharmacology techniques. The study suggested that dulcimer may be the main drug for the treatment of PMO. Quercetin, stearic acid and marine may be the main active component of QEP. MAPK1, AKT1, PIK3CA, PIK3R1, APP, SRC, MAPK8, HSP90AA1, EGFR and JAK2 may be the potential therapeutic targets of QEP for PMO. Several signaling pathways such as tumor necrosis factor signaling pathway, osteoclast differentiation, Wnt signaling pathway and estrogen signaling pathway may be involved in the mechanism of QEP efficacy.
Conflict of interests:
The authors declared no conflict of interests.
References
- Blake J, Cosman FA, Lewiecki EM, McClung MR, Pinkerton J, Shapiro M. Management of osteoporosis in postmenopausal women: The 2021 position statement of The North American Menopause Society. Menopause 2021;28(9):973-97.
[Crossref] [Google Scholar] [PubMed]
- Li J, Chen X, Lu L, Yu X. The relationship between bone marrow adipose tissue and bone metabolism in postmenopausal osteoporosis. Cytokine Growth Factor Rev 2020;52:88-98.
[Crossref] [Google Scholar] [PubMed]
- Black DM, Rosen CJ. Clinical practice. Postmenopausal osteoporosis. N Engl J Med 2016;374(3):254-62.
[Crossref] [Google Scholar] [PubMed]
- Hernlund E, Svedbom A, Ivergård M, Compston J, Cooper C, Stenmark J, et al. Osteoporosis in the European Union: Medical management, epidemiology and economic burden: A report prepared in collaboration with the International Osteoporosis Foundation (IOF) and the European Federation of Pharmaceutical Industry Associations (EFPIA). Arch Osteoporos 2013;8(1):136.
[Crossref] [Google Scholar] [PubMed]
- Wu D, Cline-Smith A, Shashkova E, Perla A, Katyal A, Aurora R. T-cell mediated inflammation in postmenopausal osteoporosis. Front Immunol 2021;12:687551.
[Crossref] [Google Scholar] [PubMed]
- Baccaro LF, Conde DM, Costa-Paiva L, Pinto-Neto AM. The epidemiology and management of postmenopausal osteoporosis: A viewpoint from Brazil. Clin Interv Aging 2015:583-91.
[Crossref] [Google Scholar] [PubMed]
- Khosla S, Hofbauer LC. Osteoporosis treatment: Recent developments and ongoing challenges. Lancet Diabetes Endocrinol 2017;5(11):898-907.
[Crossref] [Google Scholar] [PubMed]
- Riggs BL, Hartmann LC. Selective estrogen-receptor modulators-mechanisms of action and application to clinical practice. N Engl J Med 2003;348(7):618-29.
[Crossref] [Google Scholar] [PubMed]
- Khosla S, Bilezikian JP, Dempster DW, Lewiecki EM, Miller PD, Neer RM, et al. Benefits and risks of bisphosphonate therapy for osteoporosis. J Clin Endocrinol Metab 2012;97(7):2272-82.
[Crossref] [Google Scholar] [PubMed]
- Kearns AE, Khosla S, Kostenuik PJ. Receptor activator of nuclear factor κB ligand and osteoprotegerin regulation of bone remodeling in health and disease. Endocr Rev 2008;29(2):155-92.
[Crossref] [Google Scholar] [PubMed]
- Arnaud Jr CD, Tenenhouse AM, Rasmussen HO. Parathyroid hormone. Annual Rev Physiol 1967;29(1):349-72.
- Hou J, Lin S, Lu J, Wu Y, Wu L, Chen Z, et al. Establishment of a UPLC-MS/MS method for studying the effect of salt-processing on tissue distribution of twelve major bioactive components of Qing'e pills in rats. J Anal Methods Chem 2020;2020:8832736.
[Crossref] [Google Scholar] [PubMed]
- Li CG, Shen L, Yang YP, Xu XJ, Shuai B, Ma C. Effects of modified Qing’e pill on expression of adiponectin, bone morphogenetic protein 2 and coagulation-related factors in patients with nontraumatic osteonecrosis of femoral head. Chin J Integr Med 2017;23(3):183-9.
[Crossref] [Google Scholar] [PubMed]
- Xie H, Hua Z, Guo M, Lin S, Zhou Y, Weng Z, et al. Gut microbiota and metabonomics used to explore the mechanism of Qing’e pills in alleviating osteoporosis. Pharm Biol 2022;60(1):785-800.
[Crossref] [Google Scholar] [PubMed]
- Nogales C, Mamdouh ZM, List M, Kiel C, Casas AI, Schmidt HH. Network pharmacology: Curing causal mechanisms instead of treating symptoms. Trends Pharmacol Sci 2022;43(2):136-50.
[Crossref] [Google Scholar] [PubMed]
- Ru J, Li P, Wang J, Zhou W, Li B, Huang C, et al. TCMSP: A database of systems pharmacology for drug discovery from herbal medicines. J Cheminform 2014;6:13.
[Crossref] [Google Scholar] [PubMed]
- Zhang P, Chen H, Shen G, Zhang Z, Yu X, Shang Q, et al. Network pharmacology integrated with experimental validation reveals the regulatory mechanism of plastrum testudinis in treating senile osteoporosis. J Ethnopharmacol 2021;276:114198.
[Crossref] [Google Scholar] [PubMed]
- Wu Y, Zhang F, Yang K, Fang S, Bu D, Li H, et al. SymMap: An integrative database of traditional Chinese medicine enhanced by symptom mapping. Nucleic Acids Res 2019;47(D1):D1110-7.
[Crossref] [Google Scholar] [PubMed]
- Xu HY, Zhang YQ, Liu ZM, Chen T, Lv CY, Tang SH, et al. ETCM: An encyclopaedia of traditional Chinese medicine. Nucleic acids Res 2019;47(D1):D976-82.
[Crossref] [Google Scholar] [PubMed]
- Kim S, Chen J, Cheng T, Gindulyte A, He J, He S, et al. PubChem in 2021: New data content and improved web interfaces. Nucleic Acids Res 2021;49(D1):D1388-95.
[Crossref] [Google Scholar] [PubMed]
- Gfeller D, Grosdidier A, Wirth M, Daina A, Michielin O, Zoete V. SwissTargetPrediction: A web server for target prediction of bioactive small molecules. Nucleic Acids Res 2014;42(W1):W32-8.
[Crossref] [Google Scholar] [PubMed]
- Wang X, Shen Y, Wang S, Li S, Zhang W, Liu X, et al. PharmMapper 2017 update: A web server for potential drug target identification with a comprehensive target pharmacophore database. Nucleic Acids Res 2017;45(W1):W356-60.
[Crossref] [Google Scholar] [PubMed]
- Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S, et al. The GeneCards suite: From gene data mining to disease genome sequence analyses. Curr Protoc Bioinform 2016;54(1):1-30.
[Crossref] [Google Scholar] [PubMed]
- Amberger JS, Bocchini CA, Scott AF, Hamosh A. OMIM. org: Leveraging knowledge across phenotype–gene relationships. Nucl Acids Res 2019;47(D1):D1038-43.
[Crossref] [Google Scholar] [PubMed]
- Piñero J, Ramírez-Anguita JM, Saüch-Pitarch J, Ronzano F, Centeno E, Sanz F, et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res 2020;48(D1):D845-55.
[Crossref] [Google Scholar] [PubMed]
- Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, et al. The STRING database in 2021: Customizable protein-protein networks and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res 2021;49(D1):D605-12.
[Crossref] [Google Scholar] [PubMed]
- Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun 2019;10(1):1523.
[Crossref] [Google Scholar] [PubMed]
- Sun Y, Huang K, Mo L, Ahmad A, Wang D, Rong Z, et al. Eucommia ulmoides polysaccharides attenuate rabbit osteoarthritis by regulating the function of macrophages. Front Pharmacol 2021;12:730557.
[Crossref] [Google Scholar] [PubMed]
- Wang CY, Tang L, He JW, Li J, Wang YZ. Ethnobotany, phytochemistry and pharmacological properties of Eucommia ulmoides: A review. Am J Chin Med 2019;47(2):259-300.
[Crossref] [Google Scholar] [PubMed]
- Alam F, Khan GN, Asad MH. Psoralea corylifolia L: Ethnobotanical, biological and chemical aspects: A review. Phytother Res 2018;32(4):597-615.
[Crossref] [Google Scholar] [PubMed]
- Pai FT, Lu CY, Lin CH, Wang J, Huang MC, Liu CT, et al. Psoralea corylifolia L. ameliorates collagen-induced arthritis by reducing proinflammatory cytokines and up-regulating myeloid-derived suppressor cells. Life 2021;11(6):587.
[Crossref] [Google Scholar] [PubMed]
- Tian D, Hu Z. CYP3A4-mediated pharmacokinetic interactions in cancer therapy. Curr Drug Metab 2014;15(8):808-17.
[Crossref] [Google Scholar] [PubMed]
- Werk AN, Cascorbi I. Functional gene variants of CYP3A4. Clin Pharmacol Ther 2014;96(3):340-8.
[Crossref] [Google Scholar] [PubMed]
- Wang J, Wang M, Chen F, Wei Y, Chen X, Zhou Y, et al. Nano-hydroxyapatite coating promotes porous calcium phosphate ceramic-induced osteogenesis via BMP/Smad signaling pathway. Int J Nanomed 2019:7987-8000.
[Crossref] [Google Scholar] [PubMed]
- Agidigbi TS, Kim C. Reactive oxygen species in osteoclast differentiation and possible pharmaceutical targets of ROS-mediated osteoclast diseases. Int J Mol Sci 2019;20(14):3576.
[Crossref] [Google Scholar] [PubMed]
- Tian W, Teng F, Gao J, Gao C, Liu G, Zhang Y, et al. Estrogen and insulin synergistically promote endometrial cancer progression via crosstalk between their receptor signaling pathways. Cancer Biol Med 2019;16(1):55.
[Crossref] [Google Scholar] [PubMed]
- Fang JY, Richardson BC. The MAPK signalling pathways and colorectal cancer. Lancet Oncol 2005;6(5):322-7.
[Crossref] [Google Scholar] [PubMed]
- Yong HY, Koh MS, Moon A. The p38 MAPK inhibitors for the treatment of inflammatory diseases and cancer. Expert Opin Investig Drugs 2009;18(12):1893-905.
[Crossref] [Google Scholar] [PubMed]
- Guo YJ, Pan WW, Liu SB, Shen ZF, Xu Y, Hu LL. ERK/MAPK signalling pathway and tumorigenesis. Exp Ther Med 2020;19(3):1997-2007.
[Crossref] [Google Scholar] [PubMed]
- Deng R, Zhang HL, Huang JH, Cai RZ, Wang Y, Chen YH, et al. MAPK1/3 kinase-dependent ULK1 degradation attenuates mitophagy and promotes breast cancer bone metastasis. Autophagy 2021;17(10):3011-29.
[Crossref] [Google Scholar] [PubMed]
- Hirota Y, Yamashita SI, Kurihara Y, Jin X, Aihara M, Saigusa T, et al. Mitophagy is primarily due to alternative autophagy and requires the MAPK1 and MAPK14 signaling pathways. Autophagy 2015;11(2):332-43.
[Crossref] [Google Scholar] [PubMed]
- Li Q, Wu M, Fang G, Li K, Cui W, Li L, et al. MicroRNA-186-5p downregulation inhibits osteoarthritis development by targeting MAPK1. Mol Med Rep 2021;23(4):253.
[Crossref] [Google Scholar] [PubMed]
- McKenna M, Balasuriya N, Zhong S, Li SS, O'Donoghue P. Phospho-form specific substrates of protein kinase B (AKT1). Front Bioeng Biotechnol 2021;8:619252.
[Crossref] [Google Scholar] [PubMed]
- Peng XD, Xu PZ, Chen ML, Hahn-Windgassen A, Skeen J, Jacobs J, et al. Dwarfism, impaired skin development, skeletal muscle atrophy, delayed bone development and impeded adipogenesis in mice lacking Akt1 and Akt2. Genes Dev 2003;17(11):1352-65.
[Crossref] [Google Scholar] [PubMed]
- Xu F, Na L, Li Y, Chen L. Retracted article: Roles of the PI3K/AKT/mTOR signalling pathways in neurodegenerative diseases and tumours. Cell Biosci 2020;10(1):1-2.
[Crossref] [Google Scholar] [PubMed]
- Xue JF, Shi ZM, Zou J, Li XL. Inhibition of PI3K/AKT/mTOR signaling pathway promotes autophagy of articular chondrocytes and attenuates inflammatory response in rats with osteoarthritis. Biomed Pharmacother 2017;89:1252-61.
[Crossref] [Google Scholar] [PubMed]
- Yang Q, Jiang W, Hou P. Emerging role of PI3K/AKT in tumor-related epigenetic regulation. Semin Cancer Biol 2019;59:112-24.
[Crossref] [Google Scholar] [PubMed]
- Liang D, Yang M, Guo B, Cao J, Yang L, Guo X, et al. Zinc inhibits H2O2-induced MC3T3-E1 cells apoptosis via MAPK and PI3K/AKT pathways. Biol Trace Element Res 2012;148(3):420-9.
[Crossref] [Google Scholar] [PubMed]
- Gu YX, Du J, Si MS, Mo JJ, Qiao SC, Lai HC. The roles of PI3K/Akt signaling pathway in regulating MC3T3-E1 preosteoblast proliferation and differentiation on SLA and SLActive titanium surfaces. J Biomed Mater Res A 2013;101(3):748-54.
[Crossref] [Google Scholar] [PubMed]
- Perner F, Perner C, Ernst T, Heidel FH. Roles of JAK2 in aging, inflammation, hematopoiesis and malignant transformation. Cells 2019;8(8):854.
[Crossref] [Google Scholar] [PubMed]