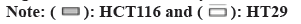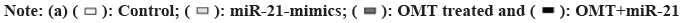- *Corresponding Author:
- Y. Zhao
Department of Gastroenterology, College of Medicine, Hebei North University, Zhangjiakou, Hebei 075000, China
E-mail: yfzhao1107@163.com
| This article was originally published in a special issue, “Advanced Targeted Therapies in Biomedical and Pharmaceutical Sciences” |
| Indian J Pharm Sci 2023:85(1) Spl Issue “119-128” |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
Colon cancer constitutes a major challenge all over the world, which has the fourth highest incidence and the third mortality rate of all cancers. Although the therapeutic modalities have been improved in recent years, the 5 y survival rate remains low. Oxymatrine is one of the main active compounds extracted from roots of Sophora flavescens and was found with therapeutic effect for colon cancers. Recently, microRNAs have been recognized as a type of diagnostic and prognostic biomarker and are implicated in a variety of biological activities underlying the tumourogenesis and cancer development. This study firstly evaluated the anti-tumor efficacy of oxymatrine in colon cancer via cell viability assay, colony formation assay and quantification of E-cadherin/vimentin as a metric of epithelial-mesenchymal transition. MicroRNA profile was conducted to identify effective microRNAs involved in the pharmacological mechanism of oxymatrine. MicroRNA-21 and microRNA-141 were the most differentially expressed microRNAs, and overexpression of both resulted in mitigation of anti-tumor effect of oxymatrine. Phosphatase and tensin homolog was predicted to be microRNA-21 and microRNA-141, which was validated via luciferase assay. Silencing of phosphatase and tensin homolog increased the level of phosphorylated-protein kinase B and phosphorylated-nuclear factorkappa B and transfection of microRNA-21 and microRNA-141 antagomir yielded increase of them, suggesting that microRNA-21/phosphatase and tensin homolog/protein kinase B and microRNA-141/phosphatase and tensin homolog/protein kinase B were involved in the pharmacological effect of oxymatrine.
Keywords
Colon cancer, microRNA-21 and microRNA-141, epithelial-mesenchymal transition, messenger RNA
Colon cancer is the most common human malignant cancer, which has the fourth highest incidence and the third mortality rate of all cancers. According to recent global cancer statistics, there were 10 96 601 new colon cancer cases, accounting for 6.1 % of all cancer patients and 5 51 269 deaths of colon cancer patients accounted for 5.8 % of all cancer- related deaths in 2018[1]. The colon cancer prognosis is not particularly ideal. In European countries, the 5 y survival rate in colon cancer is below 60 %[2]. Previous studies have shown that chemotherapy drug tolerance and cancer metastasis are the main reasons for poor prognosis in advanced colon cancer. Data show that the rate of metastatic cancer in advanced colon cancer is as high as 50 %[3].
In very early studies, it was found that Oxymatrine (OMT) activates the inflammatory responses such as p53 pathway in the human colon cancer cell line SW1116, exhibits a dose-dependent killing effect and arrest cancer cells in Growth (G) 1/G0 period[4]. In recent study, Liang et al.[5] found that OMT can inhibit the Nuclear Factor Kappa B (NF-κB) pathway activation by regulating the Epithelial- Mesenchymal Transition (EMT) process of cancer cells, thereby impeding the proliferation and migration of colon cancer cells and reducing their invasion. In addition, the latest studies have shown that OMT can up-regulate Nuclear Factor Erythroid 2-related factor 2 (Nrf2)-mediated antioxidant stress genes Heme Oxygenase 1 (HO-1) and NAD (P) H: Quinone Oxidoreductase 1 (NQO1) in mice at messenger Ribonucleic Acid (mRNA) and protein levels, which play a critical role in colon cancer[6].
Due to the heterogeneity of colon cancer and the limitations of diagnostic techniques, traditional methods of examination cannot be able to meet the requirements of clinical practice. Biomarkers are widely used for screening, diagnosis, prognosis and treatment evaluation of various cancers because of their convenience (from blood, body fluids or excretions) and accuracy. At the same time, it also has an important guiding role for tumor localization, staging and treatment options[7]. Common markers for colon cancer currently include site specific hemoglobin, Carcinoembryonic Antigen (CEA) and Carbohydrate Antigen (CA) 19-9[8], in addition to Adenomatous Polyposis Coli (APC)[9], MutL Homolog 1 (MLH1), Mut Shomologue 2 (MSH2), MSH6[10] and so on. However, the existing markers are not enough to meet the needs of clinical work; therefore, we need to conduct more in-depth research.
MicroRNA (miRNA), a kind of small non-coding RNA which has about 22 nucleotides, plays a vital role in RNA silencing and post-transcriptional regulation[11]. As a negative regulator of biological activity, miRNA can regulate a variety of biological processes including cell cycle, proliferation, apoptosis, tumor growth, metastasis and invasion[12].
The first known disease associated with miRNA dysfunction in human history is Chronic Lymphocytic Leukemia (CLL)[13]. In addition, many cancers have been shown to be associated with miRNAs. For example, studies by Lei et al.[14] indicated that in gastric cancer cells, the decrease of miR-143 and miR-145 can up-regulate its target gene Myosin VI (MYO6), affecting the EMT of gastric cells and leading to the malignant invasive manifestations of gastric cancer. The study by Li et al.[15] pointed out that miR-221/222 can enhance breast cancer development by increasing the Phosphatase and Tensin Homolog (PTEN), increase the ability of breast cancer balloon growth and increase the invasive ability of cancer cells. In addition, studies have shown that miR-21 as well as miR-155 can increase the occurrence and progression of Non- Small Cell Lung Cancer (NSCLC) by partially down- regulating Suppressor of Cytokine Signaling (SOCS) 1, SOCS6 as well as PTEN[16]. Similarly, there are many related studies on the role of miRNAs in colon cancer. For example, Zeng et al.[17] found that miR- 378 can block the metastasis and invasion of cancer cells by inhibiting SDAD1, while miR-378 can also reduce the malignancy of colon cancer cells, which may be through inhibition of Wnt/Beta (β)-catenin pathway is achieved. In addition, studies by Zhang et al.[18] showed that miR-185 also inhibits the β-catenin pathway to suppress the proliferation and invasion ability of colon cancer cells, at the same time, this inhibition is also achieved by the target gene Wnt1 of miR-185.
Materials and Methods
Cell culture:
The human Colorectal cancer (CRC) cell lines, SW480 and HCT116 were purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China). The cells were cultured in T25 cell culture flasks, containing Roswell Park Memorial Institute (RPMI)-1640 culture medium supplemented with 10 % (Fetal Bovine Serum (FBS); Gibco Laboratories, Grand Island, New York), penicillin (100 IU/ml) and streptomycin (100 μg/ml), and then incubated at 37°, 95 % humidified atmosphere with 5 % Carbon dioxide (CO2).
RNA extraction and Reverse Transcription Polymerase Chain Reaction (RT-PCR):
Total miRNA from tissues or cells was extracted using the mirVana™ miRNA isolation Kit (Ambion). mRNA and miRNA were reverse transcribed from total mRNA using the RevertAid First Strand complementary DNA (cDNA) synthesis Kit (Thermo Fisher, United States of America (USA)) according to the manufacturer’s protocol. cDNA was amplified and quantified on the CFX96 system (BIO-RAD, USA) using iQ SYBR® Green (BIO-RAD, USA). RT- PCR was performed according to a standard method, as described previously. Primers for U6, miR-21 and miR-141 were synthesized and purified by Ribo Bio (Guangzhou, China). U6 or Glyceraldehyde-3- Phosphate Dehydrogenase (GAPDH) was used as the endogenous controls. Relative fold expressions were calculated with the comparative threshold cycle (2-ΔΔCt) method.
Plasmid, small interfering RNA transfection:
Scrambled control or miR-21 (or miR-141) mimic (Thermo Fisher, Ambion) were introduced into HCT116 and SW480 cells using Lipofectamine™ RNAiMAX (ThermoFisher) for 6 h. Cells were harvested for assays 2 d after transfection. For PTEN small interfering RNA (siRNA) transfection, lentivirus expressing PTEN siRNA was constructed and amplified as described by Wang et al.[19].
Scrambled siRNA (Scr-siRNA) were prepared as negative control, respectively.
Cell viability assay:
Cell viability was assessed by 3-(4,5-Dimethylthiazol- 2-yl)-2,5-Diphenyltetrazolium Bromide (MTT) assay. The HCT116 and SW480 cells were seeded into each well of 12-well culture plates and then cultured in RPMI-1640 supplemented with 10 % heat-inactivated FBS containing 20 µg/ml penicillin and streptomycin for 72 h. MTT solution (100 µl; 5 mg/ml in Phosphate-Buffered Saline (PBS)) was then added to each well and the plates were incubated at 37° for 4 h. After removing the supernatant and shaking with 200 µl of dimethyl sulfoxide (Jersey Lab Supply, Livingston, New Jersey, USA) for 30 min and absorbance were measured at 570 nm. All experiments were repeated at least 3 times.
Luciferase assay:
The 3′-Untranslated Region (3′-UTR) of human PTEN transcript was cloned downstream of the luciferase gene between the Xho I and Sal I sites of the pmirGLO dual luciferase vector (Promega, Wisconsin, USA). A pmirGLO dual luciferase vector containing one mutated seed sequence of miR-21 (or miR-141) was constructed. The sequencing of constructed plasmids was verified by Shanghai Sangon Biotechnology Co. (Shanghai, China). 1.5×105 786- O cells in 12-well plate were co-transfected with 300 ng DNA (pmirGLO-3′ UTR constructs or derived mutants) and 30 nm of either miR-21 (or miR-141) mimics using transfection reagent Lipofectamine 2000 (Invitrogen). Luciferase activity was measured 48 h later using the Dual Luciferase Reporter Assay System (Promega)[31] with a Tecan Infinite M200 Pro microplate reader (Tecan, Mannedorf, Switzerland).
Western blot:
Total protein was extracted from HCT116 and SW480 cells, following the same protocol. Briefly, the cells were washed with 1X PBS once and each well in the 12-well cell culture plate was lysed with 100 μl of ice cold Radioimmunoprecipitation assay buffer (25 mm Tris-Cl, pH 7.4, 150 mM Sodium chloride (NaCl), 50 mM Potassium chloride (KCl), 1 % Sodium Dodecyl Sulfate (SDS), 2 mM Ethylenediaminetetraacetic Acid (EDTA), 0.5 % glycerol, 50 mM Sodium fluoride (NaF)) with 1:100 (v/v) dilution of the proteinase inhibitor and phosphatase I and II inhibitor mixture (Sigma- Aldrich; Merck KGaA). This was then briefly vortexes and placed on ice for 15 min, spun down at 8000 ×g for 15 min to pellet the debris and the supernatant was collected for the Western blotting. The protein concentration was determined using a bicinchoninic acid protein assay kit (Sigma-Aldrich; Merck KGaA), according to the manufacturer’s instructions. Then, 30 μg protein lysate was boiled with 500 mM Dithiothreitol (DTT) in 2× sample buffer for 5 min, separated by 10 % Sodium Dodecyl Sulphate-Polyacrylamide Gel Electrophoresis (SDS-PAGE) and transferred onto a nitrocellulose membrane. The membrane was blocked with Tris-buffered saline containing 5 % bovine serum albumin (Sigma-Aldrich; Merck KGaA) at room temperature for 30 min, followed by incubation with the appropriate primary antibody (1:1000) overnight at 4°. β-actin served as an internal control. Following this, membranes were incubated with a secondary antibody (1:5000) at room temperature for 1 h. Proteins were visualized by Enhanced Chemiluminescence (EMD Millipore, Billerica, Massachusetts, USA).
Scratch wound healing assay:
HCT116 and SW480 cells at the 3rd to 5th passage were digested by 0.25 % trypsin, then suspended cells in Dulbecco's Modified Eagle's Medium (DMEM) containing 10 % FBS and counted. Cells were inoculated in a 6-well plate at the total cell density of 1×105 cells/well. When the cultured cells proliferated to form a full layer, using serum-free medium for 24 h made cells synchronize. A scratch was made in the cell monolayer by drawing a sterile p-200 pipette tip across the surface of the culture dish. PBS was used to wash the plate three times. Cell migration into the scratched area was imaged after 0 h, 24 h, 48 h and 72 h. The areas of cells were measured using Image Pro plus 6.0. The migration rate of the HCT116 or SW480 cells was calculated as the ratio of the migration area to the original scratch area.
Statistical analysis:
Results are expressed as mean±standard deviation and a Student’s t-test was used for testing statistical significance. p<0.05 is considered as significant, whereas p<0.001 represents extreme significance.
Results and Discussion
We used human colon cancer cell lines SW480 and HCT116 to examine the antitumor effect of OMT.
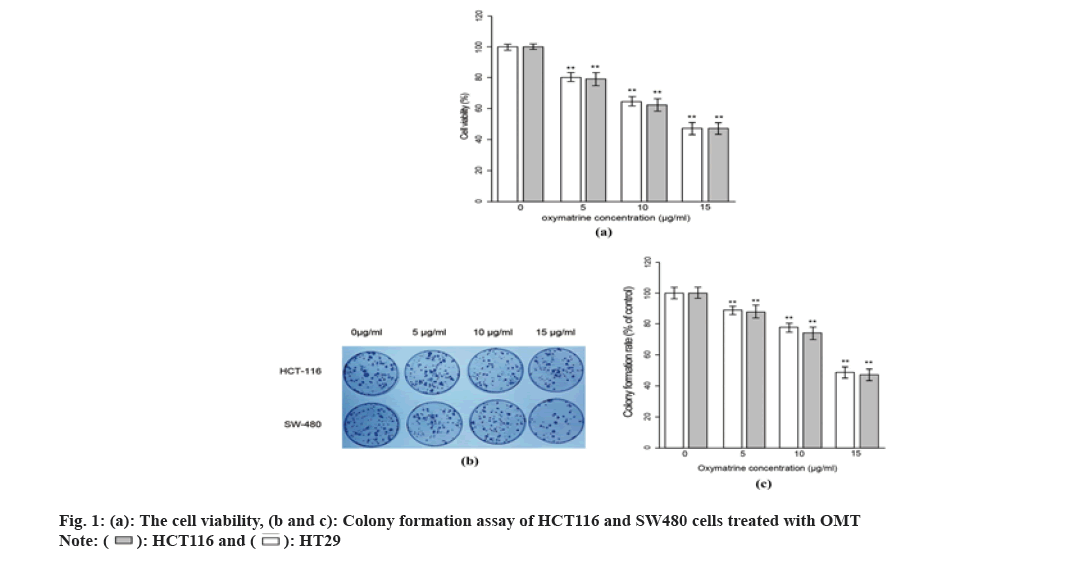
MTT assay was performed to assess the cell viability across the observation period after OMT treatment, which represents the cell proliferation rate of cells. As shown in fig. 1A, both SW480 and HCT116 cells exhibited a reduction in cell viability after being treated with different concentration of OMT. The HCT116 and SW480 cells were treated with 5, 10 and 15 μg/ml OMT respectively and were harvested for MTT assay after 48 h. The cell viability presents a dose-dependent reduction, suggesting a strong inhibitory effect on proliferation. To evaluate the changes in malignancy after OMT treatment, colony formation test was performed. As shown in fig. 1B and fig. 1C, when the OMT increased from 5 to 15 μg/ml, the colony formation rate was reduced. Finally, 10 μg/ml was chosen as an optimal dose in the following experiments since it could produce significant effect and phenotypes while avoiding evanescence of them.
EMT is a hallmark that occurs during tumourogenesis and tumor progression, which is characterized by loss of E-cadherin and gain of vimentin in the cells. The ratio of E-cadherin/vimentin serves as a metric of degree of EMT. To this end, we quantified the mRNA and protein level of both E-cadherin and vimentin and calculated the ratio of E-cadherin/vimentin. 2 d after 10 μg/ml OMT treatment, the transcription level of E-cadherin of both HCT116 and SW480 was substantially decreased so as the protein level (fig. 2A and fig. 2B). Whereas the transcription and protein expression of vimentin exhibited opposite trend.
The EMT also confers tumor cells the ability to migrate, which can be measured by wound healing assay. Thus, wound healing assay was performed to evaluate the EMT progress. As shown in fig. 2C and fig. 2D, 10 μg/ml OMT treatment resulted in larger distance between cells located on two sides of the open wound field, indicating reduced ability to migrate.
Fig. 2: Treatment with OMT reduces EMT phenotype in HCT116 and SW480 cells, (a and b): The protein level was quantified by western blot, which showed that E-cadherin expression in HCT116 and SW480 cells was mitigated and vimentin increased, and (c and d): The scratch wound healing assay was performed to quantified the ability to migrate 
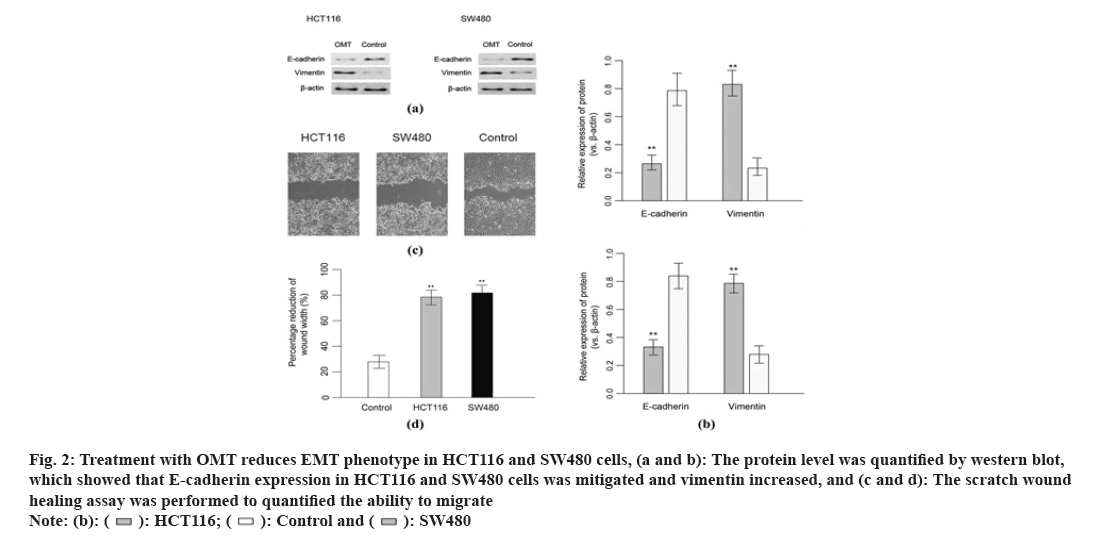
MiRNA profile was performed to sort out regulation of miRNAs in response to OMT treatment. MiRNA expression level of totally HCT116 and SW480 cells treated with or without OMT were obtained by quantifying the optical density of each probe-miRNA hybrids. A total of 53 miRNAs were significantly differentially expressed in HCT116 and SW480 cells in response to OMT treatment, in which 29 miRNAs were up regulated and 24 were down regulated. Among the 63 significant DE miRNAs, miR-21, miR-141, miR-556 and miR-639 have the largest fold change of expression, which were then subject to qRT-PCR validation (fig. 3A). Experimental validation showed that miR-21 and miR-141 have more than 3-fold decreased expression in OMT treated cells, while the other miRNAs did not show significant differential expressions (fig. 3B), suggesting that miR-21 and miR-141 have potential as prognostic biomarker and therapeutic target for colon cancer and evaluating OMT treatment.
Fig. 3: miRNA profile of CRC cells derived from patients and the validation of differentially expressed miRNAs, (a): Heat map of miRNA expression level, the samples and cluster of miRNAs were clustered with hierarchical clustering method and (b): Validation of differentially expressed miRNAs derived from profile 
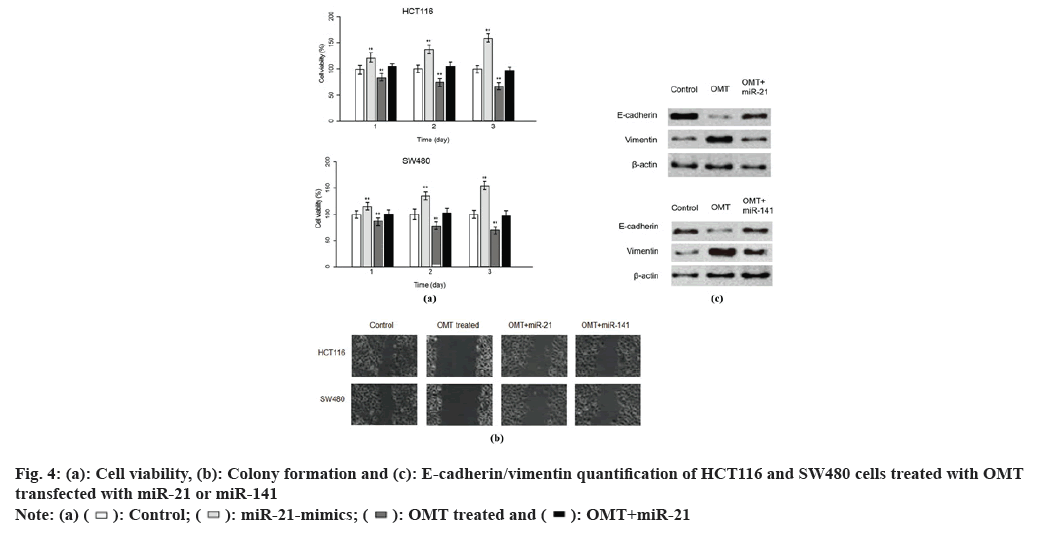
To examine the role of miR-21 and miR-141 in the antitumor mechanism of OMT, we sought to up regulate miR-21 and miR141 in HCT116 and SW480 cells treated with OMT. Lipofectamine 2000 was used to deliver antagomir of miR-21 and/or antagomir of miR-141 into HCT116 and SW480 cells, treated with or without OMT. Cell viability assay was performed after 3 d of culture. Every consecutive of 3 d, the cells were harvested and dyed with MTT and the absorbance of them were measured at 570 nm. It was obvious that transfection of miR-21 mimics or miR-141 mimics resulted in increased cell viability of HCT116 and SW480 cells treated with OMT and the co-transfection of miR-21 and miR-141 mimics led to elevated cell viability even more remarkable (fig. 4A and fig. 4B), suggesting that miR-21 and miR-141 synergistically promote cell proliferation. To further investigate the effect of miR-21 and miR- 141 on EMT phenotypes of colon cancer, wound healing assay and measurement of E-cadherin and vimentin were conducted. In wound healing assay, HCT116 and SW480 cells transfected with miR- 21 mimics or miR-141 mimics have smaller width of open wound field and co-transfection of miR-21 and miR-141 resulted in even smaller width (fig. 4C). Similarly, E-cadherin was less expressed in OMT-treated HCT116 and SW480 cells transfected with miR-21 or miR-141 mimics than HCT116 and SW480 cells treated with OMT alone (fig. 4D).
Since miRNAs regulate biological activities via targeting activator or suppressor of genes involved in related pathways, identifying the target of miR- 21 and miR141 holds the key to decipher the role of miR-21 and miR-141 in antitumor mechanism of OMT. Target Scan 7.1 calculates the target ability between miRNAs and protein-coding genes based on the complementation of 3’-UTR of genes with the sequence of miRNAs, taking into account of experimental findings. Target Scan server predicts PTEN as a target of miR-141 and we noticed that it was also a target of miR-21 reported in various cancer researches (fig. 5A). To validate our presumptions, luciferase assay was performed. The vectors carrying 3’-UTR sequence of wild type PETN have decreased luminescence compared with mutant 3’-UTR sequence, suggesting that the luciferase expression was suppressed by both miR-21 and miR-141 (fig. 5B).
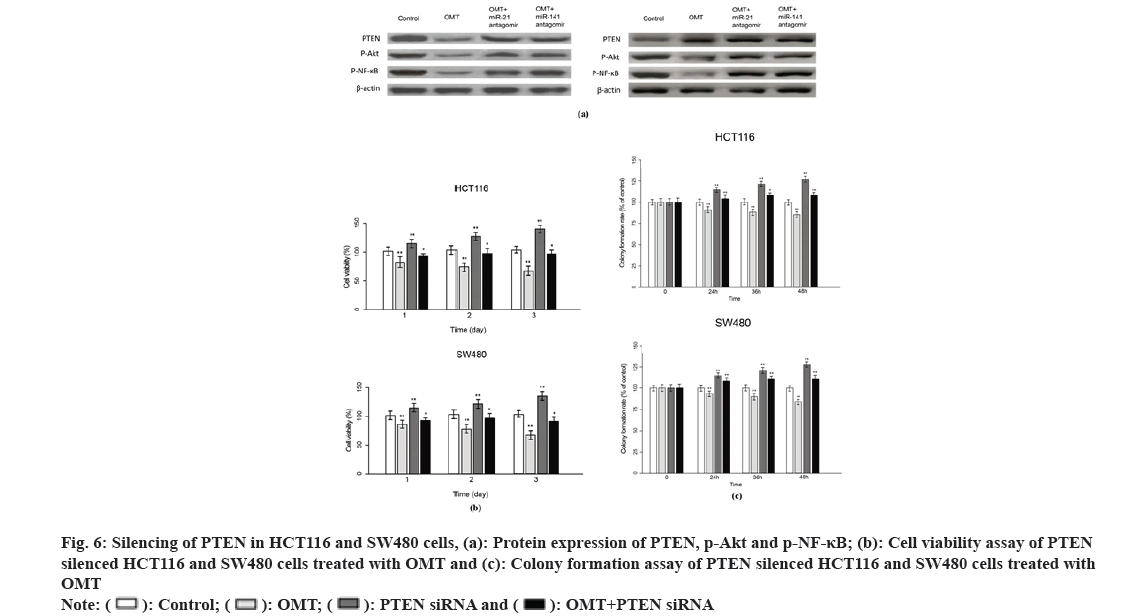
Previous studies suggest that PTEN upstream suppressor of the oncogenic NF-κB pathway, thus we presumed that NF-κB is an effector underlying the antitumor mechanism of miR-21 and miR-141 and OMT as well. To examine this, we measured the expression of proteins implicated in PTEN/NF- κB pathway. The mRNA and PTEN, Protein Kinase B (Akt) and NFKB1 were quantified by qRT-PCR and protein level of PTEN, Akt, p-Akt, NF-κB and p-NF-κB were measured by Western blot. As shown in fig. 6A, OMT treated HCT116 and SW480 cells express higher level of PTEN, and lower level of p-Akt and p-NF-κB, compared with control group. Overexpression of miR-21 and miR-141 reduced PTEN, and increased the transcription and protein level of p-Akt and p-NF-κB (fig. 6A). Silencing of PTEN in OMT treated HCT116 and SW480 cells resulted in increased expression of p-Akt and p-NF- κB, as well as the cell proliferation and colony formation rate (fig. 6B and fig. 6C).
MiRNAs have been recognized as a good marker for cancer screening, diagnosis, prognostic evaluation and guided therapy in recent studies. The most direct method of traditional cancer diagnosis is pathological biopsy, but this method is not only invasive, but also the implementation is limited by the location of the material, which can’t meet the needs of clinical work. In contrast, miRNAs can be extracted from the patient’s peripheral blood, secretions and even excretions, greatly improving the efficiency of detection. In addition, miRNAs are stable in different sample types[19], which is the basis for the accurate analysis of diseases using miRNAs. With the advancement of modern analytical techniques, miRNAs have become more widely used in cancer.
MiR-21 has been considered as a potential biomarker for a variety of cancers in previous studies. Furthermore, it is vital in biological processes including cell proliferation, migration as well as apoptosis[20]. For example, Zhang et al.[21] pointed out that miR-21 may be dual in the progression of breast cancer. Compared with normal cells, miR- 21 is highly expressed in breast cancer cells and is associated with peripheral blood sex hormone levels in patients. At the same time, high expression of miR-21 causes an increase in the proliferation and metastasis activity of cancer cells and vice versa. In addition, the authors also found that transfection of breast cancer cells with miR-21 analogs reduce the expression level of protein Signal Transducer and Activator of Transcription 3 (STAT3), a transcriptional activator that plays a key role in cell growth and apoptosis. These may be the cause of high invasiveness and high lymphatic metastasis in breast cancer. Similarly, Zhang et al.[22] also found that miR-21 is up-regulated in Osteosarcoma (OS), which suggests that X-Inactive Specific Transcripts (XIST) and Programmed Cell Death 4 (PDCD4) associated with miR-21 may be potential OS markers. The possible mechanism is that miR-21 interacts with the tumor suppressor XIST, leading to uncontrolled growth of OS cells, increasing the degree of malignancy of cancer. Similarly, there are many studies on miR-21 in colon cancer. For example, Gong et al.[23] found that RASA1 is a target gene of miR-21 and the rise of pre-miR-21 levels is often associated with high proliferation and low colon cancer cells apoptosis. The expression levels of related components of the RAS-Raf-Mitogen- Activated Protein Kinase (MAPK) pathway are also up-regulated, in which cancer cells express high pre- miR-21, resulting in colon cancer cells showing a higher degree of malignancy. At the same time, Wu et al.[24] also pointed out that miR-21 can also promote the colon cancer cells proliferation and metastasis by repressing PTEN protein expression.
There are also many studies about the role of miR- 141 in cancer. Debeb BG’s studies in mouse models have shown that high expression of miR-141 can promote the metastasis of breast cancer to brain tissue, therefore, miR-141 can become a potential therapeutic target for tumor brain metastasis[25]. Ding et al.[26] found that miR-141 can down-regulate the expression of the tumor inhibitor gene MAP2K4, thereby aggravating the proliferation of colon cancer, especially colonic adenocarcinoma cells. Looking back at more studies, we found that miR-141 appears to be involved in a variety of tumor processes by activating the NF-κB pathway. For example, a study by Quan et al.[27] found that miR-141 can inhibit the p38 MAPK and NF-κB pathways and increase the endotoxin-induced inflammatory response in WI-38 fibroblasts by increasing NADPH Oxidase 2 (NOX2) expression, but p38 MAPK suppression of the NF- κB pathway is often responsible for the malignant progression of many cancers. Similarly, Huang et al.[28] also found that miR-141 can also inhibit NF-κB pathway activation by TRAF5/6, thereby inducing bone metastasis of prostate cancer.
Interestingly, miR-21 also has an effect on the NF- κB pathway. For example, Zhou et al.[29] found that the target cell of miR-21 in NSCLC is p53 Apoptosis Stimulating Protein (ASPP2), and inhibition of high expression of miR-21 can simultaneously induce the inhibition of Phosphoinositide 3-Kinase (PI3K)/ Akt/NF-κB signaling pathway, thereby inducing apoptosis. In addition, some current researches have shown that the combination of miR-21 with miR-141 is important for the diagnosis and identification of some genitourinary cancers[30,31].
EMT is a procedure in which epithelial cells lose polarity and intercellular adhesion as well as transform into mesenchymal stem cells. Current research suggests that EMT can be broadly classified into three types, development (type I), fibrosis and wound healing (type II), as well as cancer (type III) [32,33]. Studies have shown that the premise of cancer metastasis is the invasion of cancer cells, which is achieved through the EMT process[34]. The primary tumor cells lose intercellular adhesion through EMT, thereby breaking through the basement membrane to gain increased invasiveness and entering the blood circulation and when the cancer cells in the circulating blood reach an environment suitable for proliferation, they are grown by MET cloning[35]. After reaching a new planting site, cancer cells can create better proliferation conditions through special signaling pathways. For example, a study by Song et al.[36] showed that NF-κB pathway activation induces, at the same time, maintains one of the most fundamental factors of EMT. In general, activation of the NF-κB pathway facilitates mesenchymal stem cell-type cancer cells to converse to normal immune surveillance mechanisms, reducing apoptosis, thereby facilitating better proliferation of cancer cells at the implant site.
At present, the relevant research on OMT treatment of colon cancer is not very thorough. In the early studies, it was pointed out that OMT has a killing effect on human colon cancer SW1116 cell line[4] and animal experiments by Zhang et al.[37] showed that the level of serum Tumor Necrosis Factor alpha (TNFα) in mice can be reduced. In fact, studies have shown that TNF-α is an effective activator of the NF- κB pathway[38], which indirectly indicates that the anti-malignant effect of OMT is produced by NF-κB.
In recent years, the mechanism of action of OMT in the treatment of colon cancer has been more clearly defined. For example, the combination of OMT and oxaliplatin can arrest colon cancer cells in the G0/ G1 phase by increasing the expression of P21, P27 and decreasing the level of cyclin D. In addition, the combination of the two can also down-regulate PI3K/AKT/mammalian Target of Rapamycin (mTOR) pathway to induce apoptosis in cancer cells, thereby playing a role in anti-tumor[39]. It is worth noting that recent studies have shown that OMT can affect the nuclear translocation process of NF- κB, inhibit the transcriptional activation of NF-κB and improve the intestinal epithelial barrier function through the NF-κB pathway[40], which means that NF-κB pathway may be an important way to improve colon cancer. Indeed, Liang et al. showed that OMT can inhibit the invasion of RKO cells by NF-κB signaling, and also regulate EMT markers of colon cancer cells through this pathway, such as increasing the expression of E-cadherin in cancer cells. At the same time, the level of N-cadherin and Snail was adversely reduced. Therefore, OMT plays a role in anti-colon cancer and reducing the malignancy of colon cancer.
In this study, we have explored the regulatory role of miRNA in colon cancer cell lines. In the future, Circulating Tumor Cells (CTCs) can be used to capture real colon cancer cells in the peripheral blood of colon cancer patients to verify whether miRNA has the same regulatory effect in real colon cancer cells. A crucial step in CTCs test is the enrichment of CTCs. However, our team has invented a kind of magnetic nano materials, which has the good capture efficiency for CTCs in peripheral blood of prostate cancer patients. Based on it, we can use the material to capture CTCs in peripheral blood of colon cancer patients in next experiments.
Funding:
This research was supported by the Provincial Medical Research Fund of Hebei (20210951), organized by the Provincial Health Committee of Hebei.
Conflict of interests:
The authors declared no conflict of interests.
References
- Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2018;68(6):394-424.
[Crossref] [Google Scholar] [PubMed]
- Cunningham, D. Colorectal cancer. Lancet 2010;375(9719):1030-47.
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin 2015;65(1):5-29.
- Zou J, Ran ZH, Xu Q, Xiao SD. Experimental study of the killing effects of oxymatrine on human colon cancer cell line SW1116. Chin J Dig Dis 2005;6(1):15-20.
[Crossref] [Google Scholar] [PubMed]
- Liang L, Huang J. Oxymatrine inhibits epithelial-mesenchymal transition through regulation of NF-κB signaling in colorectal cancer cells. Oncol Rep 2016;36(3):1333-8.
[Crossref] [Google Scholar] [PubMed]
- Fang R, Wu R, Zuo Q, Yin R, Zhang C, Wang C, et al. Sophora flavescens containing-QYJD formula activates Nrf2 anti-oxidant response, blocks cellular transformation and protects against DSS-induced colitis in mouse model. Am J Chin Med 2018;46(7):1609-23.
[Crossref] [Google Scholar] [PubMed]
- Sidransky D. Emerging molecular markers of cancer. Nat Rev Cancer 2002;2(3):210-9.
[Crossref] [Google Scholar] [PubMed]
- Das V, Kalita J, Pal M. Predictive and prognostic biomarkers in colorectal cancer: A systematic review of recent advances and challenges. Biomed Pharmacother 2017;87:8-19.
[Crossref] [Google Scholar] [PubMed]
- Ting WC, Chen LM, Pao JB, Yang YP, You BJ, Chang TY, et al. Common genetic variants in Wnt signaling pathway genes as potential prognostic biomarkers for colorectal cancer. PloS One 2013;8(2):e56196.
[Crossref] [Google Scholar] [PubMed]
- Scarpa M, Scarpa M, Castagliuolo I, Erroi F, Kotsafti A, Basato S, et al. Aberrant gene methylation in non-neoplastic mucosa as a predictive marker of ulcerative colitis-associated CRC. Oncotarget 2016;7(9):10322-31.
[Crossref] [Google Scholar] [PubMed]
- Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. Microarray analysis shows that some microRNAs down regulate large numbers of target mRNAs. Nature 2005;433(7027):769-73.
[Crossref] [Google Scholar] [PubMed]
- Chi Y, Zhou D. MicroRNAs in colorectal carcinoma-from pathogenesis to therapy. J Exp Clin Cancer Res 2016;35(1):43.
[Crossref] [Google Scholar] [PubMed]
- Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E, et al. Frequent deletions and down-regulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci 2002;99(24):15524-9.
[Crossref] [Google Scholar] [PubMed]
- Lei C, Du F, Sun L, Li T, Li T, Min Y, et al. miR-143 and miR-145 inhibit gastric cancer cell migration and metastasis by suppressing MYO6. Cell Death Dis 2017;8(10):e3101.
[Crossref] [Google Scholar] [PubMed]
- Li B, Lu Y, Yu L, Han X, Wang H, Mao J, et al. miR-221/222 promote cancer stem-like cell properties and tumor growth of breast cancer via targeting PTEN and sustained Akt/NF-κB/COX-2 activation. Chem Biol Int 2017;277:33-42.
[Crossref] [Google Scholar] [PubMed]
- Xue X, Liu Y, Wang Y, Meng M, Wang K, Zang X, et al. MiR-21 and MiR-155 promote non-small cell lung cancer progression by down regulating SOCS1, SOCS6 and PTEN. Oncotarget 2016;7(51):84508.
[Crossref] [Google Scholar] [PubMed]
- Zeng M, Zhu L, Li L, Kang C. miR-378 suppresses the proliferation, migration and invasion of colon cancer cells by inhibiting SDAD1. Cell Mol Biol Lett 2017;22:1-3.
[Crossref] [Google Scholar] [PubMed]
- Zhang W, Sun Z, Su L, Wang F, Jiang Y, Yu D, et al. miRNA-185 serves as a prognostic factor and suppresses migration and invasion through Wnt1 in colon cancer. Eur J Pharmacol 2018;825:75-84.
[Crossref] [Google Scholar] [PubMed]
- Weber JA, Baxter DH, Zhang S, Huang DY, How Huang K, Jen Lee M, et al. The microRNA spectrum in 12 body fluids. Clin Chem 2010;56(11):1733-41.
[Crossref] [Google Scholar] [PubMed]
- Pfeffer SR, Yang CH, Pfeffer LM. The role of miR-21 in cancer. Drug Dev Res 2015;76(6):270-7.
- Zhang C, Liu K, Li T, Fang J, Ding Y, Sun L, et al. miR-21: A gene of dual regulation in breast cancer. Int J Oncol 2016;48(1):161-72.
[Crossref] [Google Scholar] [PubMed]
- Zhang R, Xia T. Long non-coding RNA XIST regulates PDCD4 expression by interacting with miR-21-5p and inhibits osteosarcoma cell growth and metastasis. Int J Oncol 2017;51(5):1460-70.
[Crossref] [Google Scholar] [PubMed]
- Gong B, Liu WW, Nie WJ, Li DF, Xie ZJ, Liu C, et al. MiR-21/RASA1 axis affects malignancy of colon cancer cells via RAS pathways. World J Gastroenterol 2015;21(5):1488.
[Crossref] [Google Scholar] [PubMed]
- Wu Y, Song Y, Xiong Y, Wang X, Xu K, Han B, et. MicroRNA-21 (Mir-21) promotes cell growth and invasion by repressing tumor suppressor PTEN in colorectal cancer. Cell Physiol Biochem 2017;43(3):945-58.
[Crossref] [Google Scholar] [PubMed]
- Debeb BG, Lacerda L, Anfossi S, Diagaradjane P, Chu K, Bambhroliya A, et al. miR-141-mediated regulation of brain metastasis from breast cancer. J the Natl Cancer Inst 2016;108(8).
[Crossref] [Google Scholar] [PubMed]
- Ding L, Yu LL, Han N, Zhang BT. miR-141 promotes colon cancer cell proliferation by inhibiting MAP2K4. Oncol Lett 2017;13(3):1665-71.
[Crossref] [Google Scholar] [PubMed]
- Quan B, Zhang H, Xue R. miR-141 alleviates LPS-induced inflammation injury in WI-38 fibroblasts by up-regulation of NOX2. Life Sci 2019;216:271-8.
[Crossref] [Google Scholar] [PubMed]
- Huang S, Wa Q, Pan J, Peng X, Ren D, Huang Y, et al. Down regulation of miR-141-3p promotes bone metastasis via activating NF-κB signaling in prostate cancer. J Exp Clin Cancer Res 2017;36(1):173.
[Crossref] [Google Scholar] [PubMed]
- Zhou B, Wang D, Sun G, Mei F, Cui Y, Xu H. Effect of miR-21 on apoptosis in lung cancer cell through inhibiting the PI3K/Akt/NF-κB signaling pathway in vitro and in vivo. Cell Physiol Biochem 2018;46(3):999-1008.
[Crossref] [Google Scholar] [PubMed]
- Ghorbanmehr N, Gharbi S, Korsching E, Tavallaei M, Einollahi B, Mowla SJ. miR-21-5p, miR-141-3p and miR-205-5p levels in urine-promising biomarkers for the identification of prostate and bladder cancer. Prostate 2019;79(1):88-95.
[Crossref] [Google Scholar] [PubMed]
- Porzycki P, Ciszkowicz E, Semik M, Tyrka M. Combination of three miRNA (miR-141, miR-21, and miR-375) as potential diagnostic tool for prostate cancer recognition. Int Urol Nephrol 2018;50:1619-26.
[Crossref] [Google Scholar] [PubMed]
- Li L, Li W. Epithelial–mesenchymal transition in human cancer: Comprehensive reprogramming of metabolism, epigenetics and differentiation. Pharmacol Ther 2015;150:33-46.
[Crossref] [Google Scholar] [PubMed]
- Sciacovelli M, Frezza C. Metabolic reprogramming and epithelial-to-mesenchymal transition in cancer. FEBS J 2017;284(19):3132-44.
[Crossref] [Google Scholar] [PubMed]
- Hanahan D, Weinberg RA. Hallmarks of cancer: The next generation. Cell 2011;144(5):646-74.
[Crossref] [Google Scholar] [PubMed]
- Chaffer CL, Weinberg RA. A perspective on cancer cell metastasis. Science 2011;331(6024):1559-64.
[Crossref] [Google Scholar] [PubMed]
- Song R, Song H, Liang Y, Yin D, Zhang H, Zheng T, et al. Reciprocal activation between ATPase inhibitory factor 1 and NF-κB drives hepatocellular carcinoma angiogenesis and metastasis. Hepatology 2014;60(5):1659-73.
[Crossref] [Google Scholar] [PubMed]
- Zhang Y, Wang S, Li Y, Xiao Z, Hu Z, Zhang J. Sophocarpine and matrine inhibit the production of TNF-α and IL-6 in murine macrophages and prevent cachexia-related symptoms induced by colon26 adenocarcinoma in mice. Int Immunopharmacol 2008;8(13-14):1767-72.
[Crossref] [Google Scholar] [PubMed]
- Chandel NS, Trzyna WC, McClintock DS, Schumacker PT. Role of oxidants in NF-κB activation and TNF-α gene transcription induced by hypoxia and endotoxin. J Immunol 2000;165(2):1013-21.
[Crossref] [Google Scholar] [PubMed]
- Liu Y, Bi T, Wang Z, Wu G, Qian L, Gao Q, et al. Oxymatrine synergistically enhances antitumor activity of oxaliplatin in colon carcinoma through PI3K/AKT/mTOR pathway. Apoptosis 2016;21(12):1398-407.
[Crossref] [Google Scholar] [PubMed]
- Chen H, Zhang J, Luo J, Lai F, Wang Z, Tong H, et al. Antiangiogenic effects of oxymatrine on pancreatic cancer by inhibition of the NF-κB-mediated VEGF signaling pathway. Oncol Rep 2013;30(2):589-95.
[Crossref] [Google Scholar] [PubMed]