- *Corresponding Author:
- Tian E Zhang
Department of Basic Medical Sciences, Chengdu University of Traditional Chinese Medicine, Chengdu, Wenjiang 611137, China
E-mail: zhte2003@cdutcm.edu.cn
| This article was originally published in a special issue, “Recent Developments in Biomedical Research and Pharmaceutical Sciences” |
| Indian J Pharm Sci 2022:84(4) Spl Issue “7-24” |
This is an open access article distributed under the terms of the Creative Commons Attribution-NonCommercial-ShareAlike 3.0 License, which allows others to remix, tweak, and build upon the work non-commercially, as long as the author is credited and the new creations are licensed under the identical terms
Abstract
Purpose of this study was to screen the active compounds of Zhenwu decoction and their gene targets in the treatment of non-alcoholic fatty liver disease by network pharmacology and cytological validation test. The candidate compounds and related targets of Zhenwu decoction were obtained from traditional Chinese medicine system pharmacology database and PharmMapper. The non-alcoholic fatty liver disease-related genes were obtained from online Mendelian inheritance in man, GeneCards and DisGeNET databases. Molecular docking was carried out to simulate the binding affinities between potential core compounds and key target genes, and the chemical constituents of Zhenwu decoction were analyzed and identified by Ultra-Performance Liquid Chromatography-Quadrupole Time-of-Flight Mass Spectrometry technology. Therapeutic effect and mechanism of Zhenwu decoction on non-alcoholic fatty liver disease were validated using in vitro analysis. The protein-protein interaction network analysis identified the albumin, protein kinase B1, epidermal growth factor receptor, caspase 3 and peroxisome proliferator activated receptor gamma as the key target genes. Gene ontology annotation analysis showed that the Zhenwu decoctionnon- alcoholic fatty liver disease target genes were mainly involved in the response to steroid hormone and lipid catabolic process. The results of the Kyoto encyclopedia of genes and genomes pathway enrichment analysis were mainly related to the Forkhead box O signaling pathway and phosphatidylinositol 3-kinaseprotein kinase B signaling pathway. Paeoniflorgenone had the highest binding affinities for albumin, protein kinase B1, epidermal growth factor receptor, caspase 3 and peroxisome proliferator activated receptor gamma. The in vitro experiment showed that 19.5 mg/ml (p≤0.05) and 39 mg/ml (p≤0.01) of Zhenwu decoction could reduce lipid accumulation. Real-time quantitative polymerase chain reaction analyses revealed that Zhenwu decoction induced down-regulation of epidermal growth factor receptor messenger ribonucleic acid levels and up-regulation of heat shock protein 90 alpha family class a member 1, mitogen-activated protein kinase 1 and phosphatidylinositol 3-kinase messenger ribonucleic acid levels.
Keywords
Non-alcoholic fatty liver disease, Zhenwu decoction, network pharmacology, molecular docking, cytological validation
Non-Alcoholic Fatty Liver Disease (NAFLD) is defined as the presence of hepatic steatosis determined by imaging or histology, excluding the secondary causes of fat accumulation in the liver[1]. NAFLD can be divided into the Non-Alcoholic Fatty Liver (NAFL) and Non-Alcoholic Steatohepatitis (NASH)[1]. NAFLD is usually associated with metabolic complications such as obesity, diabetes and dyslipidemia. It is the major cause of liver diseases worldwide with a significantly increasing burden. At present, the incidence of NAFLD is about 200/10 000 persons per year. It is likely to be the leading cause of End-Stage Liver Disease (ESLD), which affects both adults and children[2]. Over the past decade, chronic liver diseases, cardiovascular diseases and Type 2 Diabetes Mellitus (T2DM) have been focused on among the NAFLD-related chronic diseases. A recent meta-analysis showed that the mortality rate of NAFLD, which is mainly caused by liver-related and cardiovascular diseases, had increased by 57 %, while the risk of T2DM and NAFLD-related chronic kidney disease had nearly been tripled[3]. It is crucial to implement novel therapeutic interventions that can prevent or reverse the damaging effects of NAFLD[4].
The famous Chinese medicine formula, Zhenwu Decoction (ZWD), was first described in the “Treatise on Febrile Diseases” by Zhang Zhongjing. ZWD is composed of Aconiti Lateralis Radix Praeparata (Fuzi in Chinese, lateral radix of Aconitum carmichaelii Debx.), Zingiber officinale Roscoe (Shengjiang in Chinese, the rhizome of Zingiber officinale Rosc.), Poria cocos (Schw.) Wolf. (Fuling in Chinese, the sclerotium of Poria cocos (Schw.) Wolf.), Paeoniae Radix Alba (Bai Shao in Chinese, a radix of Paeonia lactiflora Pall.) and Atractylodis macrocephalae Rhizoma (Bazhu in Chinese, a radix of Atractylodes macrocephalae Koidz.) in the ratio of 9:9:9:9:6, respectively. ZWD has a clear hypolipidemic effect, lowering serum triglycerides, total cholesterol and raising high density lipoprotein levels[5].
Clinical studies have proved the better therapeutic potential of ZWD in combination with Sini powder on T2DM overlapping NAFLD (T2DM-NAFLD) as compared to the control group but their mechanism of action is still not clear[6]. ZWD could regulate energy metabolism, reduce oxidative stress by lowering serum lysophospholipid levels in rats[7].
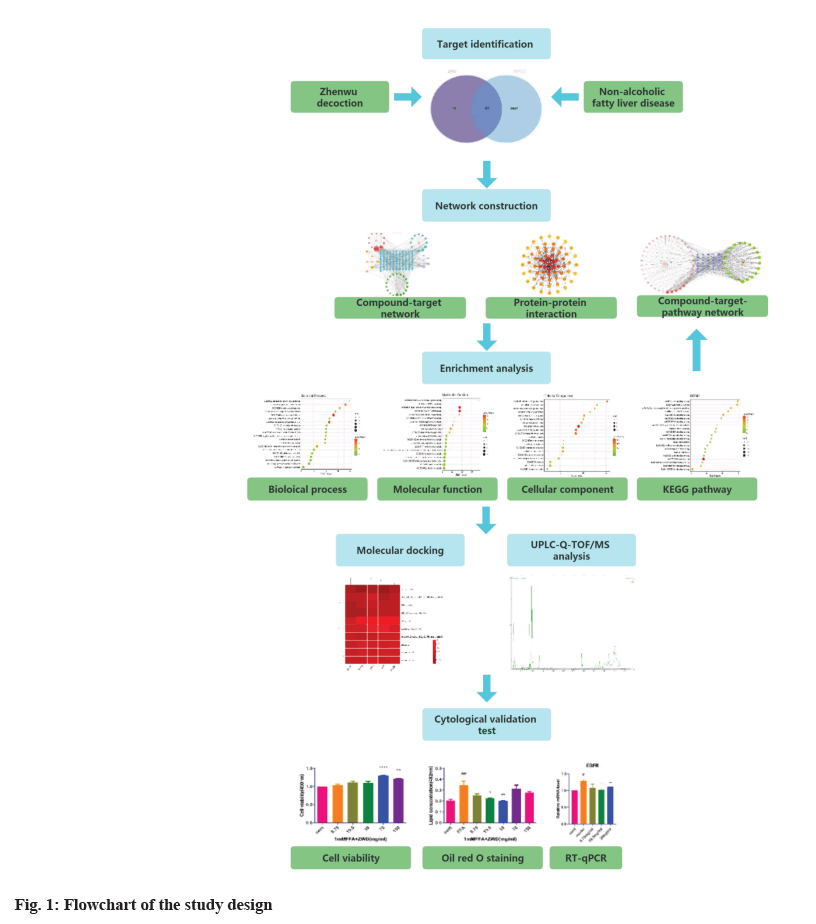
Network pharmacology adopts the “multi-compound, multi-target and multi-pathway” method and involves system biology, bioinformatics and pharmacology. It is usually used to study the systematic effects of traditional Chinese medicine and is considered to be a novel strategy for drug discovery[8] for significantly improving the success rate of drug development, reducing the cost and predicting the side effects of drugs[9]. This study was the first to elucidate the mechanisms of ZWD in NAFLD treatment with a network pharmacology approach. The flowchart of the study is presented in fig. 1.
Materials and Methods
Extraction and screening of the active compounds of ZWD:
Chinese names of the five traditional Chinese herbs present in ZWD were used to search the active compounds of each medicine in the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) (http://tcmspw.com/tcmsp.php). TCMSP contains Chinese medicinal herbs, chemicals, targets and drug-target networks. The active compounds in ZWD were obtained using Oral Bioavailability (OB)≥30 % and Drug-Likeness, (DL)≥0.18 as screening conditions.
Screening of the targets and core compounds of ZWD:
The screened active compounds of ZWD were submitted into the PharmMapper database (http://lilab-ecust.cn/ pharmmapper/index.html) to obtain the corresponding targets using “Norm Fit≥0.90” as screening criteria. The corresponding gene names of the target proteins were obtained from through UniProt database (https://www.uniprot.org/). Duplicate gene names were then removed. The compound-target network of ZWD was constructed and visualized using Cytoscape 3.8.2, and the core compounds of ZWD were determined.
Collection of known therapeutic targets acting on the NAFLD and candidate genes:
The therapeutic target genes of NAFLD were obtained from three databases, including GeneCards (https://www.genecards.org/), Online Mendelian Inheritance in Man (OMIM) (http://www.omim.org/) and DisGeNET (https://www.disgenet.org/). The search term “nonalcoholic fatty liver” was used in all three databases and the species were limited to “Homo sapiens”. The therapeutic targets were collected and duplicate targets were removed. NAFLD-related protein targets were imported into the UniProt database. Then, the obtained genes were submitted to Venny 2.1 (https://bioinfogp.cnb.csic.es/tools/venny/) to draw a Venn diagram. The intersected genes in the two datasets were selected as candidate genes.
Construction of the Protein-Protein Interaction (PPI) network:
Search Tool for the Retrieval of Interacting Genes/ Proteins (STRING) database (https://string-db.org/) was used to study the association of the retrieved active ingredient targets of ZWD and NAFLD target genes. The confidence score of relevance was set to ≥0.4 as the critical value to obtain the PPI network. The PPI network was visualized using Cytoscape 3.8.2 and the key genes were identified by degree, betweenness and closeness.
Enrichment analyses of Gene Ontology (GO) annotation and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway:
The gene targets of ZWD-NAFLD were subjected to GO annotation and KEGG pathway enrichment analysis using Metascape (https://metascape.org/). The target species was set to Homo sapiens. The results were presented using an R software package.
Construction of the compound-target-pathway network:
The active compound of ZWD, selected gene targets and related pathways were submitted to Cytoscape 3.8.2 in order to build a compound-target-pathway network. The graphic network represented the connections and interactions among the target genes, proteins or molecules and pathways. The value of each node represented the number of connections between the node and other nodes and the greater the degree, the more important the node.
Molecular docking between the main active components of ZWD and core proteins:
In order to further verify the reliability of this study, the top five targets in the PPI network and the top 10 core compounds of ZWD were selected for molecular docking. First, the Spatial Data File (SDF) files of the core compounds of ZWD were downloaded from the PubChem database (https://www.uniprot.org/) and then converted to mol2 format files using Open Babel 2.4.1 software. The best three Dimensional (3D) protein structures of the main protein targets were obtained from Protein Data Bank (PDB) and saved in PDB format. Then, the downloaded protein structures and active compounds of ZWD were opened in PyMOL to remove the water molecules and ligands. The active compounds and protein targets were then imported to AutoDock software for docking. The smaller the value of docking binding force, the more stable the compound binds to the protein and the more likely it is to interact. Finally, the PDB file was imported to LigPlot software to find the interaction between small molecules and large proteins.
Ultra-Performance Liquid Chromatography- Quadrupole Time-of-Flight Mass Spectrometry (UPLC-QTOF/MS) analysis of ZWD:
Standard preparation: 0.3 ml, 0.6 ml, 1.2 ml of ZWD were diluted into 1.2 ml, 0.9 ml, 0.3 ml of 50 % methanol solution, ultrasonic treatment for 30 min and then filtered with 0.22 μm microporous membrane. UPLC analysis conditions were as follows. The results were performed on a liquid chromatography-mass spectrometer (SNAPT XS) equipped with ACQUITY UPLC CSH C18 (100 mm×2.1 mm, 1.7 μm) at 40°. The mobile phase consisted of 0.1 % formic acid aqueous solution (A) and acetonitrile (B). The optimized gradient program was 0-5 min, 95 %-90 % A; 5-8 min, 85 % A; 8-15 min, 85 %-83 % A; 15-18 min, 83 %-80 % A; 18-22 min, 80 %-70 % A; 22-28 min, 70 %-5 % A; 28-31 min, 5 %-5 % A; 31-35 min, 5 %-95 % A. The flow rate was 0.2 ml/min and the injection volume was 2 μl.
MS analysis conditions were as follows. MS analysis was performed on a liquid chromatographymass spectrometer (SNAPT XS) equipped with an Electrospray Ionization (ESI) source. The capillary voltage is set to 2000 V, the source temperature is 200°, the desolvation temperature is 450°, the cone gas flow is 50 l/h and the desolvation gas flow is 600 l/h. The sample infusion flow rate was 5 μ/min. The full scan quality range of the scan is m/z 50-1200. Data is obtained in continuous mode.
Preparation of ZWD:
The dried raw herbs of ZWD were purchased from the outpatient department of Chengdu University of traditional Chinese medicine (Chengdu, China). Zingiber officinale Roscoe was homemade (fresh Chengdu-origin ginger was washed and sliced). Aconiti Lateralis Radix Praeparata, Paeoniae radix Alba, Poria cocos (Schw.) Wolf., Zingiber officinale Roscoe and Atractylodes macrocephala Koidz. (Production batch number: 21090104, 21081906, 21101061, C038210701, respectively) were mixed at the proportion of 9:9:9:9:6, that is 42 g soaked in water (500 ml) for 30 min and then boiled for 30 min. The residues were filtered and concentrated to 0.78 g/ml and after centrifugation, the supernatant was collected and five concentration gradients (9.75 mg/ml, 19.5 mg/ml, 39 mg/ml, 78 mg/ ml and 156 mg/ml) were prepared, which were then stored at 4°.
Cell cultures and treatments:
Liver hepatocellular carcinoma cell line (HepG2) cells were cultured in 90 % high glucose sugar Dulbecco’s Modified Eagle Medium (DMEM, Gibco), containing 10 % Fetal Bovine Serum (FBS, Gibco), 1 % penicillin and streptomycin. The cells were maintained at 37° under 5 % Carbon dioxide (CO2). Free Fatty Acids (FFA) are a mixture of oleic acid and palmitic acid at 2:1. The cells were treated with different concentrations of FFAs and cell viability was observed.
The HepG2 cell viability was determined using Cell Counting Kit-8 (CCK-8). For this purpose, HepG2 cells in the logarithmic phase (2×103 cells per well) were inoculated into 96-well plates (100 μl/well) and treated with the different concentrations of ZWD (9.75 mg/ml, 19.5 mg/ml, 39 mg/ml, 78 mg/ml, 156 mg/ml) for 24 h. Then, the bottom solution was removed and 10 μl of CCK-8 solution was added to 100 μl complete medium. The CCK-8 solution was added to each well and the plate was incubated at 37° for 2 h. The absorbance was measured at 450 nm using a microplate reader.
Oil Red O (ORO) staining:
The cells were washed with Phosphate Buffered Saline (PBS) solution and fixed with 4 % paraformaldehyde for 30 min. The ORO storage solution (0.5 %) was prepared with 0.25 g oil red dry powder and 50 ml isopropanol and filtered through a 0.22 μm filter. The ORO working solution was prepared by mixing the ORO storage solution and deionized water at a ratio of 3:2. The fixed HepG2 cells were stained with an ORO working solution for 30 min at 37° and then immediately washed with PBS for 3 times. The qualitative analysis of lipids and quantification of lipid accumulation under the microscope were based on the Optical Density (OD) value of the target ORO.
Ribonucleic Acid (RNA) isolation and Real-Time quantitative Polymerase Chain Reaction (RTqPCR):
RT-qPCR was performed to measure the hepatic cell expression of the predicted NAFLD-related genes. Total RNA was collected with the RNA isolation kit following the instructions, complementary Deoxyribonucleic Acid (cDNA) synthesis kit with genomic DNA (gDNA) eraser was used to carry out the reverse transcription reactions. RT-qPCR was performed to measure the relative expression of messenger RNA (mRNA), using SYBR Green RT-PCR master mix. The reaction conditions were 95°, 3 min; 95°, 10 s; 55°, 20 s and 72°, 30 s, for 40 cycles. Beta-actin was used as a control and the 2-ΔΔCt method was conducted for the data analysis. The primer sequence of target genes was synthesized by Invitrogen Biotech Co., Ltd (Table 1). The targets Epidermal Growth Factor Receptor (EGFR), Peroxisome Proliferator Activated Receptor Gamma (PPARG), Heat Shock Protein 90 Alpha Family Class A Member 1 (HSP90AA1) and Mitogen-Activated Protein Kinase 1 (MAPK1) formed the PPI network analysis and Phosphatidylinositol 3-Kinase (PI3K) formed the PI3K-Protein Kinase B (AKT) signaling pathway.
| Gene name | Amplicon length | Sequences |
|---|---|---|
| EGFR-F | 97 | 5’-ACCCATATGTACCATCGATGTC-3’ |
| EGFR-R | 97 | 5’-GAATTCGATGATCAACTCACGG-3’ |
| PPARG-F | 120 | 5’-AGATCATTTACACAATGCTGGC-3’ |
| PPARG-R | 120 | 5’-TAAAGTCACCAAAAGGCTTTCG-3’ |
| HSP90AA1-F | 191 | 5’-CCAGTTCGGTGTTGGTTTTTAT-3’ |
| HSP90AA1-R | 191 | 5’-CAGTTTGGTCTTCTTTCAGGTG-3’ |
| MAPK1-F | 188 | 5’-ATGGTGTGCTCTGCTTATGATA-3’ |
| MAPK1-R | 188 | 5’-TCTTTCATTTGCTCGATGGTTG-3’ |
| PI3K-F | 180 | 5’-GAGATTGCAAGCAGTGATAGTG-3’ |
| PI3K-R | 180 | 5’-TAATTTTGGCAGTGATTGTGGG-3’ |
| Beta-actin-F | 285 | 5’-AGCGAGCATCCCCCAAAGTT-3’ |
| Beta-actin-R | 285 | 5’-GGGCACGAAGGCTCATCATT-3’ |
Note: F: Forward primer and R: Reverse primer
Table 1: The Nucleotide Sequences of The Primer Pairs Used for Quantitative Gene Expression
Statistical analysis:
All the data were expressed as mean±Standard Error (SE). One-way Analysis of Variance (ANOVA) in GraphPad Prism v8.3.0 was used to test and analyze the data. The difference with p<0.05 was considered statistically significant.
Results and Discussion
According to the screening criteria of OB≥30 % and DL≥0.18, a total of 61 active compounds were obtained for the five medicinal herbs of ZWD. After removing the duplicate data, a total of 59 compounds were obtained of which 20 were from Fuzi, 12 from Bai Shao, 7 from Baizhu, 15 from Fuling and 5 from Sheng Jiang. After removing the duplicate data and screening the target genes using PharmMapper and through the standardized transformation of Uniprot, a total of 133 target genes were obtained.
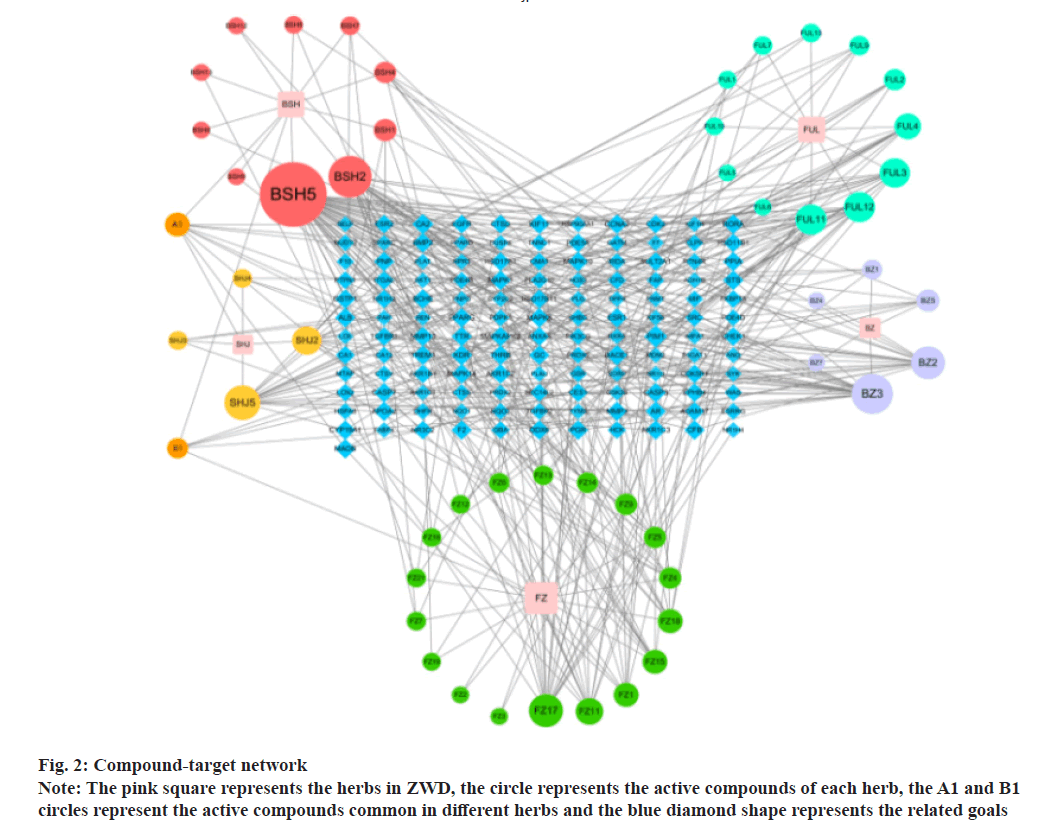
The compound-target network was constructed to show the complex compounds and multiple target genes and the correlations between them. There were 303 nodes and 824 edges in the compound-target network of ZWD. It was suggested that the compounds of ZWD might play a synergistic effect on these target genes and might have pharmacological effects on NAFLD and other diseases. The highest connectivity was that of BSH5 with 55 target genes with an OB value of 53.87 %; this might be considered as the potential key active compound as shown in fig. 2.
As shown by the compound-target network, some compounds play a crucial role in the network, thereby proving to be a great potential in the treatment of NAFLD, such as paeoniflorin, dihydrocapsaicin and poricoic acid B. Zhang found that paeoniflorin could prevent NAFLD by restoring the serum Alanine Aminotransferase (ALT), Aspartate Aminotransferase (AST), Serum Total Cholesterol (TC), Triglyceride (TG), High-Density Lipoprotein (HDL) and Low- Density Lipoprotein (LDL) levels. Paeoniflorin could alleviate the infiltration of liver fats induced by a high-fat diet through reducing steatosis, inflammation, balloon degeneration and necrosis. Suggested mechanisms may relate to the protection on cardiovascular system as body weight and hyperlipidemia reduced, inflammation blocked and lipid deposition inhibited[8]. In addition, paeoniflorin could also significantly reduce serum insulin and glucagon levels, enhance insulin sensitivity, restore serum lipid spectrum and alleviate liver steatosis[9]. The dietary dihydrocapsaicin could also reduce blood lipid levels and improve cholesterol metabolism in hyperlipidemia rats[10]. It is worth mentioning that the three compounds in Poria cocos Wolf, including poricoic acid, pachymic acid and ergosterol, could significantly reduce the TG level in HepG2 cells treated with FFAs[11].
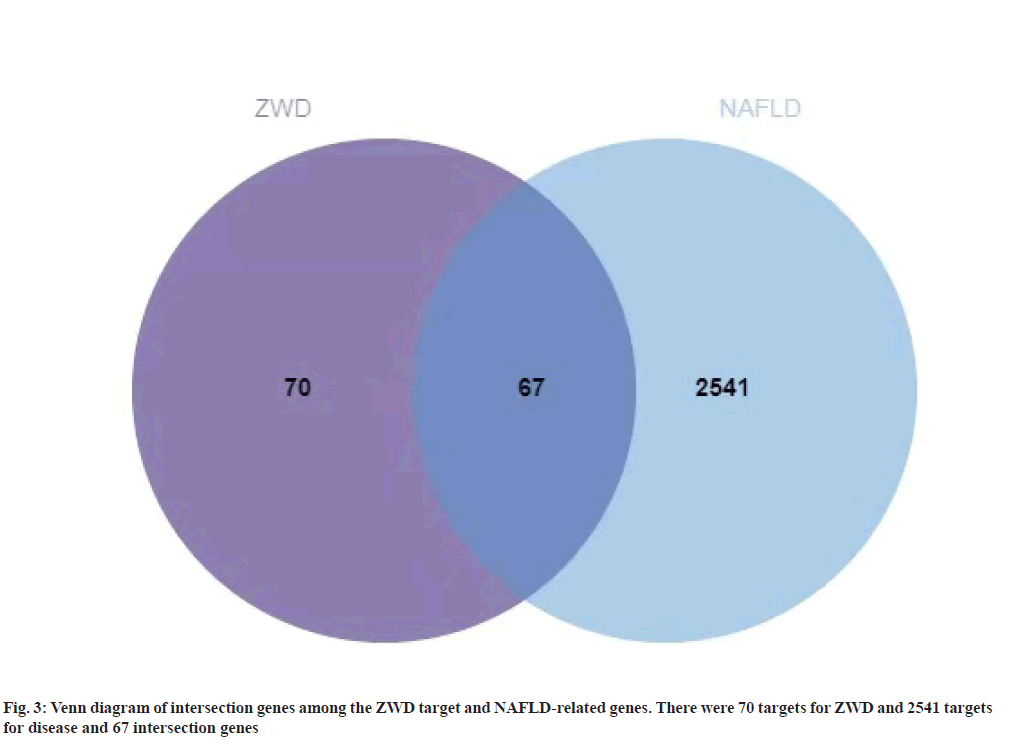
A total of 3181 NAFLD-related target genes were obtained from OMIM, GeneCards and DisGeNET databases. After removing the duplicate data, a total of 2608 target genes were obtained as shown in Table 2. The intersection genes among the NAFLD-related and ZWD target genes were extracted, which were 67 target genes and were considered as a candidate gene. The intersection genes are shown in fig. 3.
| Source | Compound | Degree | Binding affinity/KJ/mol | ||||
|---|---|---|---|---|---|---|---|
| ALB | AKT1 | EGFR | CASP3 | PPARG | |||
| BSH5 | Paeoniflorin | 56 | -1.74 | -1.63 | -1.24 | -2.36 | -1.35 |
| BSH2 | Paeoniflorgenone | 30 | -4.42 | -3.8 | -4.55 | -3.83 | -3.67 |
| BZ3 | 14-acetyl-12-senecioyl-2E,8Z,10E-atractylentriol | 28 | -1.74 | -2.31 | -2.82 | -2.05 | -1.55 |
| SHJ5 | Dihydrocapsaicin | 22 | -2.42 | -2.26 | -2.45 | -3.22 | -2.43 |
| FZ17 | Jesaconitine | 20 | 0.88 | 0.26 | -0.02 | -1.64 | 0.74 |
| BZ2 | 14-acetyl-12-senecioyl-2E,8E,10E-atractylentriol | 20 | -3.34 | -1.83 | -2.41 | -2.59 | -2.56 |
| FUL12 | Poricoic acid B | 17 | -1.56 | -1.93 | -1.84 | -2.08 | -1.77 |
| FUL11 | Poricoic acid A | 17 | -1.93 | -1.35 | -1.98 | -2.14 | -1.75 |
| FUL3 | 7,9(11)-dehydropachymic acid | 16 | -2.5 | -2.42 | -3.03 | -3.45 | -2.46 |
| SHJ2 | 6-methylgingediacetate 2 | 15 | -0.36 | -0.51 | -1.82 | -1.78 | -1.04 |
Table 2: The Binding Energy Values Of Core Compounds Of Zwd And Core Targets
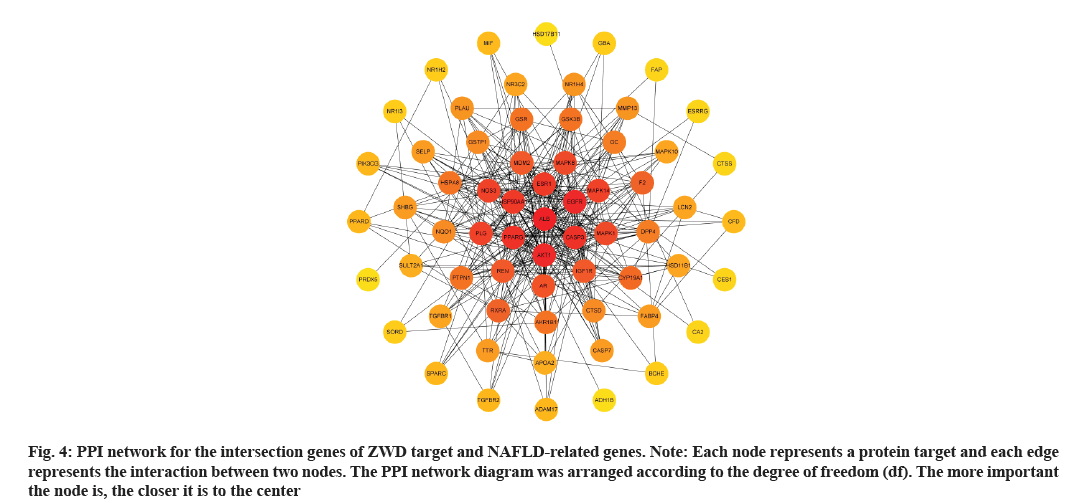
A ZWD-NAFLD target PPI network was obtained, which contained 64 nodes and 718 edges (fig. 4). The nodes represented proteins and edges represented protein interactions. The average node degree value was 10.7 and the average local clustering coefficient was 0.627. In the PPI network, some nodes had a high degree, such as Albumin (ALB), Protein Kinase B1 (AKT1), EGFR, Caspase 3 (CASP3) and PPARG.
Fig. 4: PPI network for the intersection genes of ZWD target and NAFLD-related genes.
Note: Each node represents a protein target and each edge represents the interaction between two nodes. The PPI network diagram was arranged according to the degree of freedom (df). The more important the node is, the closer it is to the center
These target nodes were identified as key genes and might be potential therapeutic targets for the treatment of NAFLD by ZWD. Alkaline phosphatase, ALT, AST, ALB and total serum protein were significantly correlated with the severity of NAFLD[12]. Besides, there were significant differences in the ALB-binding function between healthy subjects and different severity groups of NAFLD[13]. Differential regulation of AKT1 and AKT2 is consistent with upregulation of Forkhead Box Protein O1 (FOXO1), may justify the paradoxical state of insulin resistance relative to the glucoregulatory pathway and augmented insulin sensitivity of the liporegulatory pathway, typical of steatosis and the metabolic syndrome in obese patients[14]. Liang et al. demonstrated that the EGFR was phosphorylated in the liver tissues of the highfat diet murine model of NAFLD. The inhibition of EGFR could prevent diet-induced lipid accumulation, oxidative stress, hepatic stellate cell activation and matrix deposition[15]. Hepatocyte death is an important feature of NASH. The CASP3 activation in hepatocytes played an important role in the NASH-related apoptosis and fibrosis[16]. Finally, a study reported PPARG was positively correlated with liver TG levels in male mice[17]. Therefore, ALB, AKT1, EGFR, CASP3 and PPARG might be the potential therapeutic targets for the treatment of NAFLD.
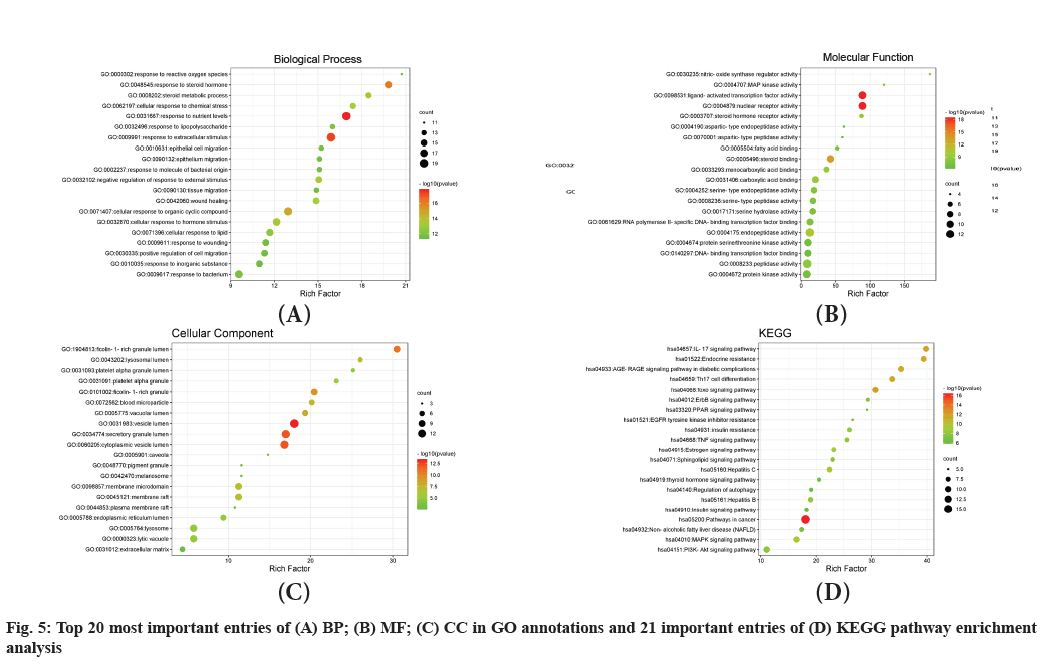
The GO annotations of the candidate genes were carried out in three categories, including Biological Process (BP), Molecular Function (MF) and Cell Composition (CC). The top 20 significantly-enriched terms in BP, MF and CC categories (adjusted p<0.01) are presented in fig. 5A-fig. 5C, respectively. A total of 793 entries were displayed in the BP enrichment analysis, among which, many BPs such as response to nutrient levels, response to a steroid hormone, cellular response to lipids and lipid catabolic process, were closely associated with the pathogenesis of NAFLD. The CC enrichment results showed 45 terms, including vesicle lumen, secretory granule lumen, ficolin-1-rich granule, lytic vacuole and pigment granule. A total of 67 enrichment processes were identified related to MF, including nuclear receptor activity, steroid binding, steroid hormone receptor activity, DNA-binding, transcription factor binding and beta-catenin binding. In order to further investigate the biological process of these targets, the KEGG pathways enrichment analysis was conducted, which resulted in a total of 202 entries (p<0.01), including FOXO signal and PI3K-AKT signaling pathway, endocrine resistance and estrogen signaling pathway.
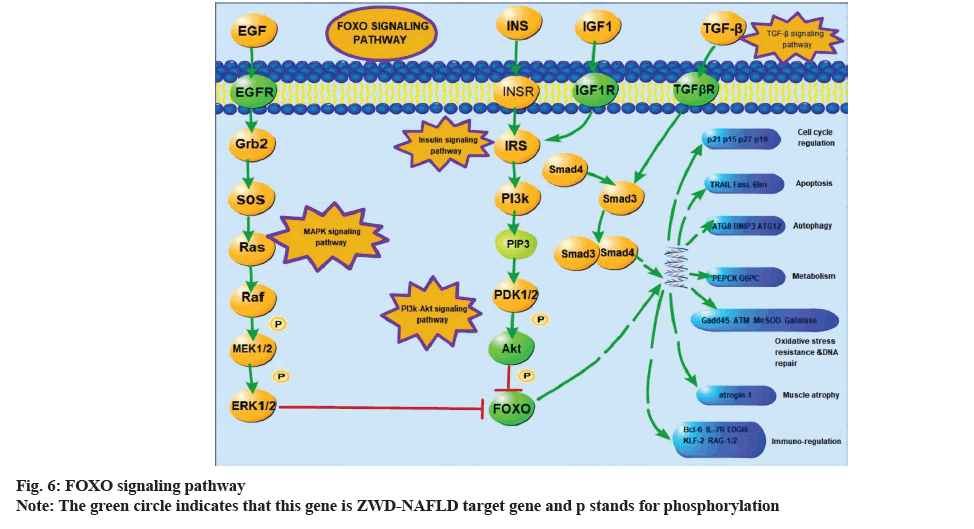
The pathogenesis of NAFLD is closely related to insulin resistance[18]. A core regulatory mechanism in the FOXO signaling pathway is the phosphorylation of AKT which is the downstream of the PI3K for insulin response (fig. 6). A study showed that the FOXO signal pathway could lead to lipid accumulation and the denaturation of hepatocytes and promote the occurrence and development of NAFLD[19]. PI3K-AKT signal transduction pathway in hepatocytes is a common molecular mechanism, which is involved in metabolic dysfunction, including obesity, metabolic syndrome and NAFLD[20]. Cai et al. found that the insulin-PI3K/ AKT-p70 S6 Kinase (p70S6K) pathway played an important role in the early activation of hepatic stellate cells[21]. Interestingly, a study reported that paeoniflorin could have positive influence on improving NAFLD and obesity by regulating the PI3K/AKT pathway[22]. The results of our compound-target-pathway suggested that paeoniflorin could interact with the PI3K-AKT signaling pathway through AKT1, EGFR, Glycogen Synthase Kinase-3 Beta (GSK3B), HSP90AA1, Mouse Double Minute 2 homolog (MDM2) and Insulin-like Growth Factor 1 Receptor (IGF1R), while paeoniflorgenone could interact with this pathway through MAPK1 and Retinoid X Receptor Alpha (RXRA). In addition, there are also numerous endocrine-related pathways, such as endocrine resistance, estrogen signaling pathway and thyroid hormone signaling pathway. Zhu et al. found that the estrogen signaling pathway could inhibit fatty acid oxidation and TGs in the liver and increase peripheral fat mobilization[23].
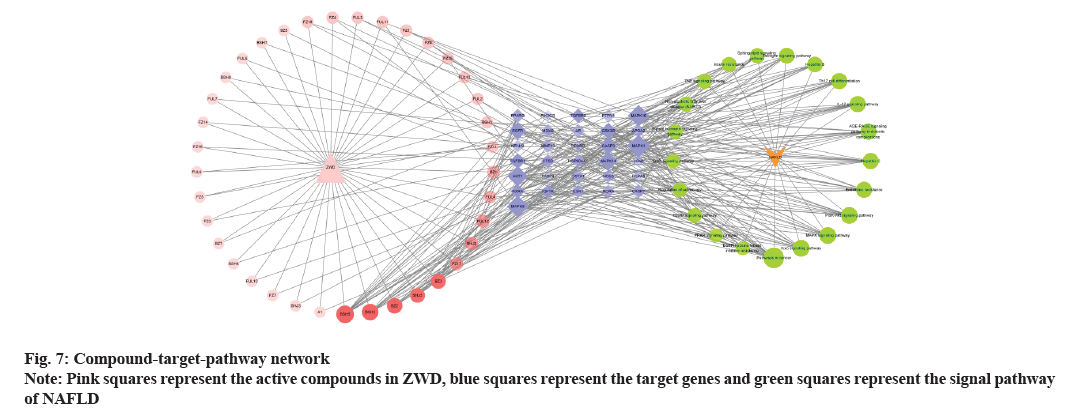
Cytoscape v3.8.2 software was used to construct a compound-target-pathway network, which is shown in fig. 7. There were 328 edges and 91 nodes in the network, which showed that ZWD could treat NAFLD through its multiple compounds and multiple target genes. Among them, paeoniflorin, paeoniflorgenone, 14-acetyl-12-senecioyl-2E, 8Z, 10E-atractylentriol, dihydrocapsaicin, etc., might play important roles in the efficacy of ZWD for the treatment of NAFLD. The specific information of the compounds is listed in Table 1.
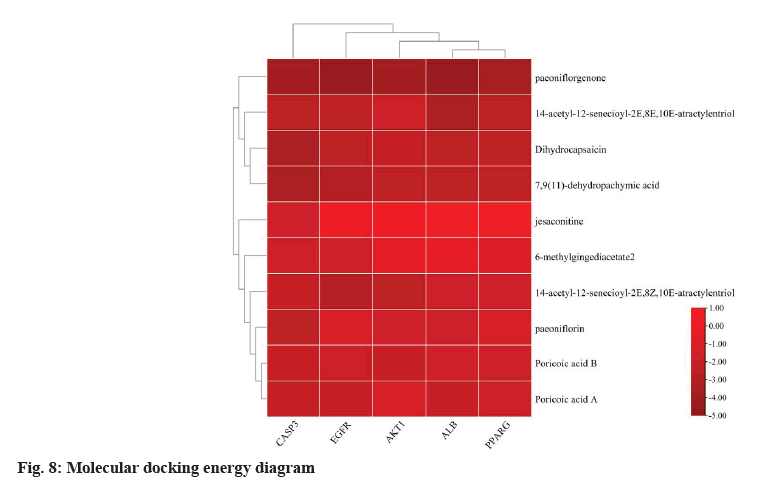
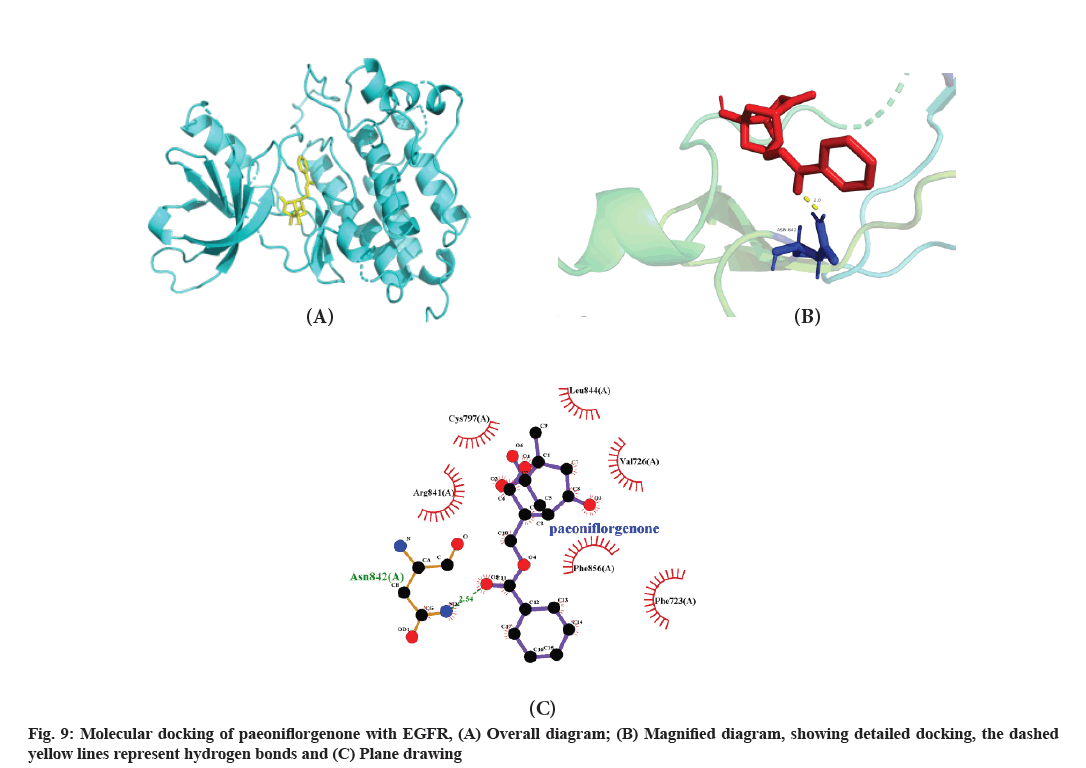
The top 10 active compounds (paeoniflorin, paeoniflorgenone, 14-acetyl-12-senecioyl-2E, 8Z, 10E-atractylentriol, dihydrocapsaicin, jesaconitine, 14-acetyl-12-senecioyl-2E, 8E, 10E-atractylentriol, poricoic acid B, poricoic acid A, 7, 9 (11)-dehydropachymic acid and 6-methylgingediacetate 2) obtained from the compound-target network analysis were docked with the five potential core target genes (ALB, AKT1, EGFR, CASP3 and PPARG) obtained from the PPI network analysis. The details of these compounds and target genes are listed in Table 2 and presented in fig. 8. The binding energies of all the compounds except jesaconitine were less than 0, showing a good binding affinity of these compounds with the top five target genes. Among them, the binding energy of paeoniflorgenone with ALB, AKT1, CASP3 and PPARG proteins was lower than that of the other compounds and the paeoniflorgenone-EGFR showed the lowest binding energy (-4.55); ASN-842 was the active site, showing hydrogen bond interactions (fig. 9). Docking prediction provided a preliminary basis for the further study of these drug targets.
The docking of the compounds with target proteins and their binding free energy, which were less than 1, suggested that the 10 major core compounds had a good binding affinity for the five core target proteins. The binding energy of paeoniflorgenone was the lowest among the 10 compounds, indicating its good binding affinity with all the five target proteins. The paeoniflorgenone-EGFR showed the lowest binding energy (-4.55) and ASN-842 was the active site for hydrogen bonding.
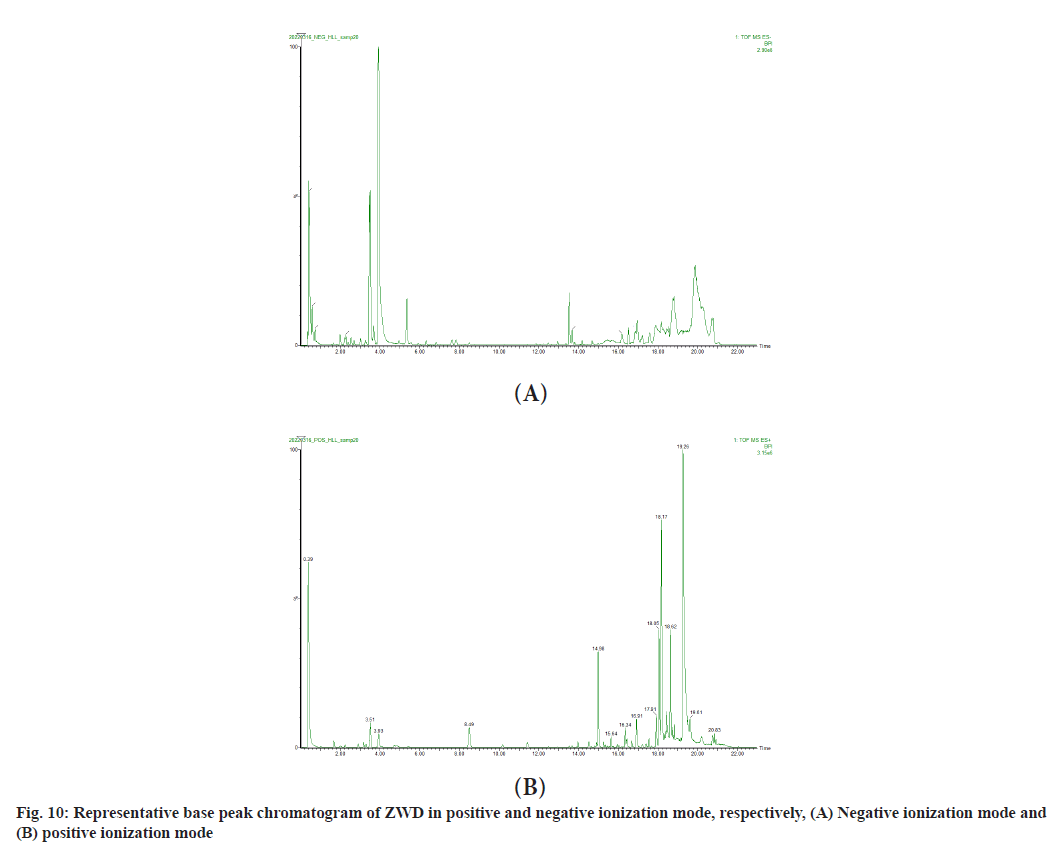
In order to verify whether the 10 components predicted by molecular docking exist in ZWD, UPLC-Q-TOFMS was performed to analyze the chemical constituents of ZWD. Full scanning was carried out in positive and negative ion mode and the analysis showed that the chromatogram of representative Base Peak Intensity (BPI) in positive and negative ion mode was as shown in fig. 10. A total of 67 compounds were detected by UniFi software (Table 3). Among them, paeoniflorin, paeoniflorgenone and poricoic acid B were proved to exist in ZWD. Therefore, it can be speculated that paeoniflorin, paeoniflorin and poricoic acid B may play a major role in ZWD in the treatment of NAFLD.
| Component name | Neutral mass (Da) | Observed neutral mass (Da) | Observed m/z | Mass error (mDa) | Mass error (ppm) | Observed RT (min) | Detector counts | Response | Adducts | Total fragments found |
|---|---|---|---|---|---|---|---|---|---|---|
| Furfuryl alcohol | 98.03678 | 98.0374 | 143.0356 | 0.6 | 4.4 | 0.43 | 430 | 430 | HCOO+ | 1 |
| Epinephrine | 416.15299 | 416.1552 | 455.1184 | 2.2 | 4.9 | 0.43 | 1149 | 1149 | K+ | 40 |
| 16-Oxoacetylpyrochloric acid | 570.39204 | 570.3945 | 609.3576 | 2.4 | 4 | 0.45 | 388 | 388 | K+ | 42 |
| Paradisamide | 421.28282 | 421.2832 | 444.2724 | 0.3 | 0.8 | 0.46 | 1283 | 400 | Na+ | 50 |
| Coryneine | 195.12593 | 195.1268 | 196.1341 | 0.9 | 4.4 | 0.46 | 619 | 619 | H+ | 12 |
| 1-Galloyl-β-D-glucose | 332.07435 | 332.0751 | 331.0678 | 0.7 | 2.2 | 0.62 | 8711 | 7759 | H- | 19 |
| Gallic acid | 170.02152 | 170.022 | 169.0148 | 0.5 | 3 | 0.74 | 42699 | 40055 | H- | 2 |
| 5-Hydroxymethyl-2-furaldehyde | 126.03169 | 126.0322 | 125.025 | 0.5 | 4.4 | 0.74 | 6362 | 6017 | H- | 3 |
| 1-O-β-D-Glucopyranosylpaeonisuffrone | 360.14203 | 360.1432 | 405.1414 | 1.1 | 2.8 | 0.81 | 17152 | 8794 | HCOO+, H- | 12 |
| Carmicheline | 377.25661 | 377.2566 | 378.2639 | 0 | 0.1 | 1.24 | 395 | 395 | H+ | 0 |
| Senbusine B | 423.26209 | 423.2619 | 424.2692 | -0.2 | -0.4 | 1.91 | 772 | 772 | H+ | 1 |
| Atractylenolide II | 232.14633 | 232.147 | 233.1542 | 0.6 | 2.7 | 2.01 | 2012 | 1710 | H+ | 5 |
| d-Catechin | 290.07904 | 290.08 | 289.0727 | 0.9 | 3.2 | 2.24 | 26247 | 22781 | H- | 8 |
| Nonyn | 357.23039 | 357.231 | 358.2383 | 0.7 | 1.8 | 2.25 | 20376 | 16960 | H+ | 1 |
| Benzoic acid | 122.03678 | 122.0372 | 123.0445 | 0.4 | 3.6 | 2.26 | 496 | 496 | H+ | 1 |
| Dictysine | 347.24604 | 347.247 | 348.2542 | 0.9 | 2.6 | 2.27 | 433 | 433 | H+ | 2 |
| Oxypaeoniflorin | 496.15808 | 496.1591 | 495.1518 | 1 | 2.1 | 2.28 | 44480 | 35623 | H-, HCOO+ | 21 |
| Paeoniflorin | 480.16316 | 480.1649 | 481.1721 | 1.7 | 3.5 | 3.04 | 2749 | 2186 | H+ | 10 |
| Fuziline | 453.27265 | 453.2739 | 454.2812 | 1.3 | 2.8 | 3.08 | 1526 | 1526 | H+ | 3 |
| Nyonin | 437.27774 | 437.2789 | 438.2862 | 1.2 | 2.7 | 3.18 | 33673 | 27072 | H+ | 6 |
| Kaempferol-3,7-di-O-β-D-glucoside | 610.15338 | 610.1551 | 609.1478 | 1.7 | 2.8 | 3.21 | 326 | 326 | H- | 1 |
| β-Eudesmol | 222.19837 | 222.1978 | 245.187 | -0.6 | -2.4 | 3.27 | 697 | 697 | Na+ | 0 |
| 6-O-β-D-Glucopyranosyllactinolide | 362.15768 | 362.1584 | 361.1511 | 0.7 | 2 | 3.31 | 1142 | 1142 | H- | 14 |
| (Z)-(1S,5R)-β-Pinen-10-yl-β-vicanoside | 446.2152 | 446.2168 | 491.215 | 1.6 | 3.3 | 3.62 | 563 | 563 | HCOO+ | 1 |
| Denudatine | 343.25113 | 343.2524 | 344.2597 | 1.3 | 3.7 | 3.72 | 4188 | 3440 | H+ | 4 |
| Ethyl salicylate | 166.06299 | 166.0637 | 165.0565 | 0.7 | 4.4 | 3.93 | 2153 | 2153 | H- | 14 |
| Poricoic acid B | 196.07356 | 196.0744 | 197.0817 | 0.8 | 4.2 | 3.94 | 9904 | 9086 | H+ | 18 |
| Benzyl alcohol | 108.05751 | 108.058 | 109.0652 | 0.5 | 4.2 | 3.94 | 2463 | 2463 | H+ | 4 |
| Delbruline | 479.2883 | 479.2903 | 480.2976 | 2 | 4.2 | 4.42 | 373 | 373 | H+ | 3 |
| Paeoniflorigenone | 318.11034 | 318.1115 | 319.1188 | 1.2 | 3.7 | 5.3 | 1695 | 1695 | H+ | 2 |
| Lactiflorin | 462.1526 | 462.1546 | 507.1528 | 2 | 3.9 | 6.27 | 1006 | 785 | HCOO+ | 0 |
| Paeonilactone C | 318.11034 | 318.1119 | 319.1192 | 1.6 | 4.9 | 7.86 | 2726 | 2355 | H+ | 7 |
| Hokbusine A | 603.30435 | 603.3056 | 604.3128 | 1.2 | 2 | 10.17 | 48550 | 36036 | H+ | 5 |
| Benzoylaconitine | 603.30435 | 603.3057 | 648.3039 | 1.3 | 2.1 | 10.18 | 5395 | 3794 | HCOO+ | 2 |
| Benzoylhypaconine | 573.29378 | 573.2947 | 574.302 | 0.9 | 1.6 | 11.42 | 72877 | 54681 | H+ | 5 |
| Atractylenolide IV | 306.14672 | 306.148 | 305.1407 | 1.2 | 4 | 12.27 | 277 | 277 | H- | 7 |
| Beiwutine | 647.29418 | 647.2967 | 648.304 | 2.5 | 3.9 | 12.65 | 1232 | 838 | H+ | 1 |
| Benzoyloxypaeoniflorin | 600.18429 | 600.1854 | 599.1781 | 1.1 | 1.8 | 13.53 | 602 | 602 | H- | 25 |
| 3-Deoxynivalenol | 629.32 | 629.3219 | 628.3146 | 1.9 | 3 | 13.84 | 417 | 417 | H- | 5 |
| Aconitine | 645.31491 | 645.3173 | 646.3246 | 2.4 | 3.7 | 14.02 | 538 | 538 | H+ | 4 |
| Benzoylpaeoniflorin | 584.18938 | 584.1922 | 585.1995 | 2.8 | 4.8 | 14.48 | 464 | 464 | H+ | 2 |
| Palbinone | 358.21441 | 358.215 | 357.2077 | 0.6 | 1.7 | 14.49 | 319 | 319 | H- | 1 |
| Poricoic acid A | 512.35017 | 512.3483 | 551.3114 | -1.9 | -3.5 | 14.77 | 856 | 856 | K+ | 3 |
| Delbruine | 465.27265 | 465.2744 | 466.2816 | 1.7 | 3.7 | 14.8 | 1321 | 901 | H+ | 3 |
| Paeonilactone | 168.11503 | 168.1156 | 169.1229 | 0.6 | 3.5 | 14.87 | 1947 | 1947 | H+ | 5 |
| 6-Gingerol | 294.18311 | 294.184 | 317.1732 | 0.9 | 2.8 | 14.99 | 5834 | 5014 | Na+ | 38 |
| Hypaconitine | 615.30435 | 615.3066 | 638.2958 | 2.3 | 3.6 | 15 | 393 | 393 | Na+ | 14 |
| Poricoic acid D | 514.32944 | 514.3301 | 513.3229 | 0.7 | 1.3 | 15.06 | 905 | 642 | H- | 2 |
| 25-Hydroxy-3-epidehydrotumulosic acid | 500.35017 | 500.3498 | 499.3425 | -0.4 | -0.8 | 15.23 | 1282 | 955 | H- | 3 |
| Poricoic acid B | 484.31887 | 484.3196 | 483.3123 | 0.7 | 1.4 | 15.25 | 217 | 217 | H- | 0 |
| Hydroxy-atractylenolide | 248.14124 | 248.1425 | 247.1352 | 1.2 | 5 | 15.26 | 222 | 222 | H- | 0 |
| 3β-hydroxy-atractylon | 232.14633 | 232.1473 | 233.1545 | 0.9 | 4 | 16.03 | 7081 | 6265 | H+ | 4 |
| 3β,16α-Dihydroxylauroster-8,24-dien-21-oic acid | 472.35526 | 472.356 | 471.3487 | 0.8 | 1.6 | 16.2 | 217 | 217 | H- | 3 |
| Poricoic acid F | 498.33452 | 498.3365 | 497.3293 | 2 | 4 | 16.24 | 309 | 309 | H- | 4 |
| 3-Epidehydrotumulosic acid | 484.35526 | 484.3564 | 483.3491 | 1.1 | 2.3 | 16.39 | 452 | 452 | H- | 3 |
| Dehydrotumulosic acid | 486.37091 | 486.3718 | 485.3645 | 0.9 | 1.8 | 16.48 | 386 | 386 | H- | 3 |
| Atractylenolide A | 230.13068 | 230.1315 | 231.1387 | 0.8 | 3.4 | 16.53 | 438 | 438 | H+ | 2 |
| 3β-Hydroxy-11α,12α-epoxyolean-28-13β-olide | 486.33452 | 486.3369 | 485.3296 | 2.4 | 4.8 | 16.61 | 338 | 338 | H- | 2 |
| Ginger brain | 276.17254 | 276.1732 | 277.1805 | 0.7 | 2.5 | 16.7 | 4270 | 3648 | H+ | 4 |
| Poricoic acid C | 482.33961 | 482.3401 | 481.3328 | 0.4 | 0.9 | 16.77 | 353 | 353 | H- | 1 |
| β-Red myrcene | 204.1878 | 204.1885 | 205.1958 | 0.7 | 3.6 | 17.08 | 431 | 431 | H+ | 0 |
| Pachymic acid | 528.38147 | 528.3828 | 527.3756 | 1.4 | 2.6 | 17.67 | 485 | 485 | H- | 2 |
| β-aqua-anisidine | 136.1252 | 136.1258 | 137.1331 | 0.6 | 4.4 | 18.18 | 1095 | 1095 | H+ | 14 |
| Salsolinol | 193.11028 | 193.1109 | 194.1182 | 0.7 | 3.5 | 18.55 | 2682 | 2682 | H+ | 2 |
| Eburicoic acid | 470.376 | 470.3781 | 493.3674 | 2.1 | 4.3 | 18.6 | 646 | 646 | Na+ | 4 |
| Hexadecane | 256.24023 | 256.2408 | 255.2336 | 0.6 | 2.4 | 18.82 | 15674 | 13670 | H- | 7 |
| γ-Sitosterol | 414.38617 | 414.3875 | 453.3506 | 1.3 | 2.9 | 18.88 | 1913 | 1913 | K+ | 1 |
Table 3: Identification of The Compounds of ZWD by UPLC-Q-TOF/MS Analysis
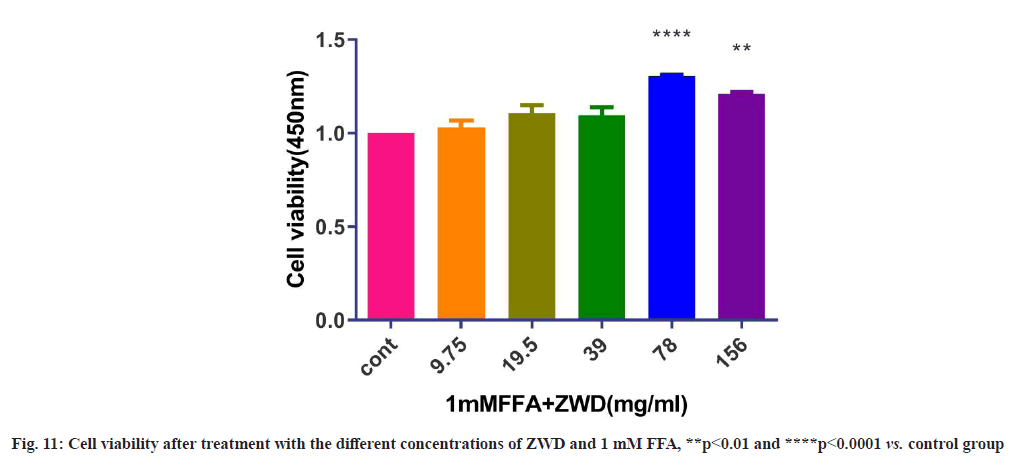
Referring to the relevant literature[24] and the experimental results in this study, 1 mM FFA was selected as model concentration. As shown in fig. 11, the results of CCK-8 assay showed that ZWD at concentrations ranging from 9.75-156 mg/ml+1 mM FFA displayed no cytotoxicity on cells.
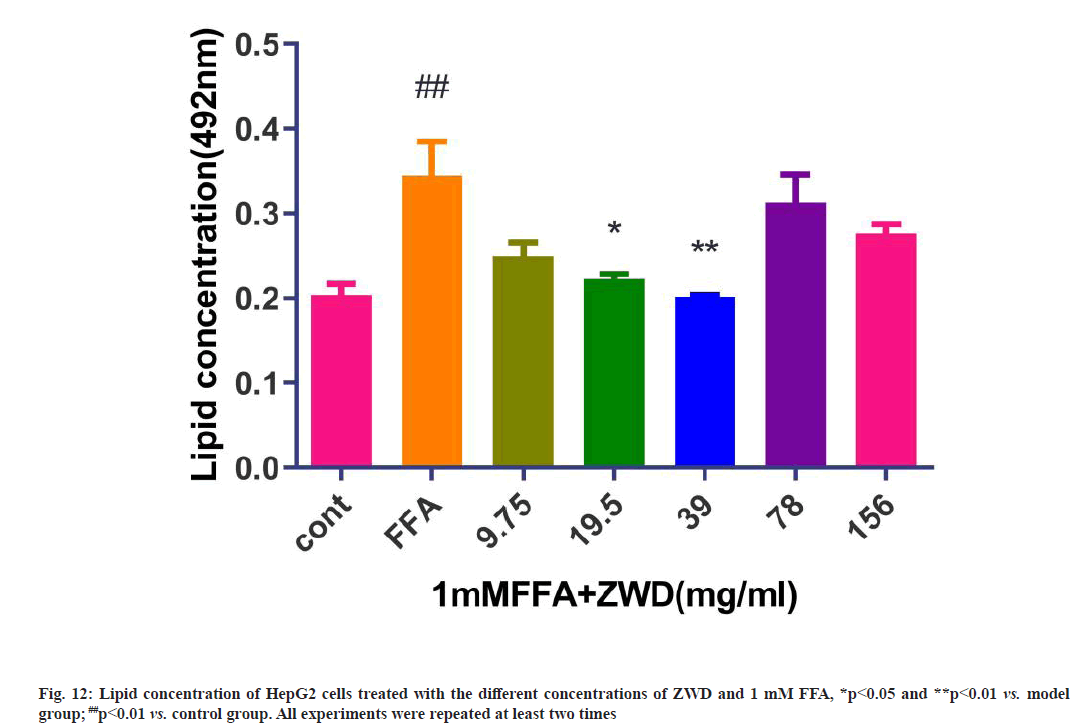
Treatment with FFA can induce significant lipid accumulation in cells. After the cells were treated with the different doses of ZWD for 24 h, the ORO staining showed a significant difference between the experimental group and model group, there were obvious differences when the cells were treated with 19.5 mg/ml and 39 mg/ml of ZWD. The TG contents in HepG2 cells in the experimental group were lower than those in the control group, as shown in fig. 12. These results suggested that 19.5 mg/ml (p≤0.05) and 39 mg/ ml (p≤0.01) of ZWD could reduce lipid accumulation in a concentration-dependent manner.
The results of cell-based assays showed that 1 mM FFA could significantly induce lipid accumulation in the HepG2 cells. After the intervention of ZWD, cell viability enhanced, which might be due to the effect of Fuzi because it significantly promote cell viability even in low concentration[25]. The pathogenesis of NAFLD is a ‘three-hit’ process, including steatosis, lipotoxicity and inflammation, which impair the hepatocyte viability, eventually leading to death and exacerbation of NAFLD[26]. Therefore, it was speculated that the improvement of cell vitality by ZWD might be the mechanism of its therapeutic effect. The intracellular lipid contents were decreased by ZWD in a concentration-dependent manner, where 19.5 mg/ml (p≤0.05) and 39 mg/ml (p≤0.01) of ZWD could reduce lipid accumulation. For validation purposes, RTqPCR was performed to test the expression of EGFR, PPARG, HSP90AA1, MAPK1 and PI3K. RT-qPCR experiments proved ZWD induced down-regulation of EGFR mRNA levels and up-regulation of HSP90AA1, MAPK1 and PI3K mRNA levels.
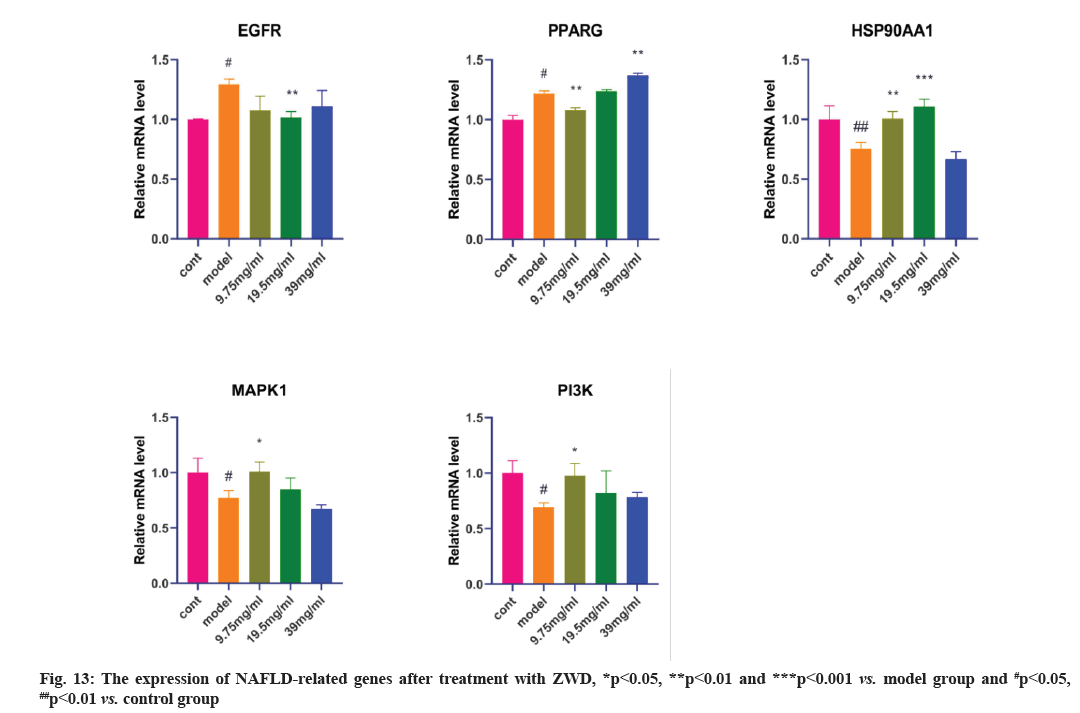
The network pharmacology results identified potential key targets and pathways of ZWD active against NAFLD. In order to verify the reliability of network pharmacological prediction results, we detected the mRNA level of these key genes by qRT-PCR. As shown in fig. 13, RT-qPCR results showed that mRNA expression of EGFR and PPARG was increased in the model group compared with that in the control group, and after three concentration gradient of ZWD treated for 24 h, the expression of EGFR mRNA was decreased, the expression of PPARG mRNA showed that there was an inconsistent change trend between high concentration and low concentration. The mRNA expression of HSP90AA1, MAPK1 and PI3K was decreased in the model group compared with that in the control group, and after treatment, the mRNA expression of HSP90AA1, MAPK1 and PI3K was increased.
The results of our compound-target-pathway suggested that the active ingredients in ZWD could interact with the PI3K-AKT signaling pathway through EGFR, HSP90AA1 and MAPK1. The activated PI3K/AKT signaling pathway can exert anti-inflammatory, anti-oxidative stress, anti-apoptotic and autophagic regulatory effects through downstream pathways and related proteins[27], these processes are closely related to the pathogenesis of NAFLD. MAPK1 has been shown to be involved in cellular processes of autophagy, lipid metabolism, proliferation, migration[28] and the activation of MAPK 1/3 ameliorates liver steatosis in leptin receptor-deficient mice[29]. The function of HSP90AA1 is to promote the maturation, structural maintenance and proper regulation of specific target proteins involved in cell cycle control and signal transduction[30]. Thus, our findings show that ZWD can positively regulate NAFLD in multiple ways through active ingredients and targets.
In summary, this study demonstrated a strategy to optimize conventional network pharmacology and explained the molecular mechanism of ZWD in NAFLD treatment was closely associated with 5 core genes (including AKT1, EGFR, CASP3, PPARG and IGF1R), which were involved in important signal pathways (including FOXO signaling pathway and PI3K-AKT signaling pathway). RT-qPCR analyses revealed that the therapeutic effect of ZWD might be achieved via down-regulation of EGFR mRNA levels and up-regulation of HSP90AA1, MAPK1 and PI3K mRNA levels.
Author’s contributions:
Chun-Jiang Zhang and Zhi-Yan Fang contributed equally to this work. Tian-e Zhang, Yan-Qiu Wang and Chun-Jiang Zhang: Conceptualization; Chun-Jiang Zhang, Zhi-Yan Fang, Zheng Luo, Hai-Yan Zhu and Lili Huang: Methodology and visualization; Wen-Ying Huai and Yan-Qiu Wang: Software; Lili Huang and Yan-Qiu Wang: in vitro cells-based experiment; Chun- Jiang Zhang and Tian-e Zhang: Writing-original draft; Tian-e Zhang and Yu-Qin Tang: Writing-review and editing; Tian-e Zhang and Qiao-Zhi Yin: Supervision and project administration. All authors have read and agreed to the published version of the manuscript.
Acknowledgements:
This study was supported by the Key Research and Development Program of Science and Technology Department of Sichuan Province (grant number 2020JDZH0018), The fourth National TCM Doctor (Clinical And Basic) Excellent Talents Research and Study Program (grant number CMM No. 24 [2017]), the Sichuan Provincial Famous TCM Doctor Studio Construction Project (Sichuan TCM 003113005003), the Special Research Project of Sichuan Provincial Administration of Traditional Chinese Medicine (grant number 2021MS099), Technological innovation research and development projects of Chengdu Science and Technology Bureau (Grant number 2021-YF05- 02379-SN) and the “Double First-class” construction project of Chengdu University of Traditional Chinese Medicine (030041043).
Conflict of interests:
The authors declared that there are no conflicts of interest regarding the publication of this paper.
References
- Chalasani N, Younossi Z, Lavine JE, Charlton M, Cusi K, Rinella M, et al. The diagnosis and management of nonalcoholic fatty liver disease: Practice guidance from the American association for the study of liver diseases. Hepatology 2018;67(1):328-57.
[Crossref] [Google scholar] [PubMed]
- Younossi Z, Anstee QM, Marietti M, Hardy T, Henry L, Eslam M, et al. Global burden of NAFLD and NASH: Trends, predictions, risk factors and prevention. Nat Rev Gastroenterol Hepatol 2018;15(1):11-20.
[Crossref] [Google scholar] [PubMed]
- Musso G, Gambino R, Cassader M, Pagano G. Meta-analysis: Natural history of non-alcoholic fatty liver disease (NAFLD) and diagnostic accuracy of non-invasive tests for liver disease severity. Ann Med 2011;43(8):617-49.
[Crossref] [Google scholar] [PubMed]
- Musso G, Gambino R, Tabibian JH, Ekstedt M, Kechagias S, Hamaguchi M, et al. Association of non-alcoholic fatty liver disease with chronic kidney disease: A systematic review and meta-analysis. PLoS Med 2014;11(7):e1001680.
[Crossref] [Google scholar] [PubMed]
- Jiansheng L. Study on the effect of Zhenwu decoction extract on experimental obesity and blood lipid metabolism and mechanism. Sichuan J Physiol Sci 2004;26(2):49-51.
- Li H. The clinical research of Sini powder combined with Zhenwu decoction to patients with type 2 diabetes mellitus overlapped nonalcoholic fatty liver disease (T2DM-NAFLD). World Latest Med Inf 2019;19(93):254-58.
- Li S, Xiao X, Han L, Wang Y, Luo G. Renoprotective effect of Zhenwu decoction against renal fibrosis by regulation of oxidative damage and energy metabolism disorder. Sci Rep 2018;8(1):1-11.
[Crossref] [Google scholar] [PubMed]
- Zhang LJ, Yang B, Yu BP. Paeoniflorin protects against nonalcoholic fatty liver disease induced by a high-fat diet in mice. Biol Pharm Bull 2015;38(7):1005-11.
[Crossref] [Google scholar] [PubMed]
- Li YC, Qiao JY, Wang BY, Bai M, Shen JD, Cheng YX. Paeoniflorin ameliorates fructose-induced insulin resistance and hepatic steatosis by activating LKB1/AMPK and AKT pathways. Nutrients 2018;10(8):1024.
[Crossref] [Google scholar] [PubMed]
- Chen Z, Liu Y, Wang H, Chen Z, Liu J, Liu H. Effects of sanshoamides and capsaicinoids on plasma and liver lipid metabolism in hyperlipidemic rats. Food Sci Biotechnol 2019;28(2):519-28.
- Kim JH, Sim HA, Jung DY, Lim EY, Kim YT, Kim BJ, et al. Poria cocus wolf extract ameliorates hepatic steatosis through regulation of lipid metabolism, inhibition of ER stress, and activation of autophagy via AMPK activation. Int J Mol Sci 2019;20(19):4801.
[Crossref] [Google scholar] [PubMed]
- Lee JT, Pao LH, Lee MS, Liao JW, Shih CM, Hsiong CH, et al. A new approach to facilitate diagnosis of nonalcoholic fatty liver disease through a galactose single point method in rats with fatty liver. Dig Liver Dis 2013;45(2):134-41.
[Crossref] [Google scholar] [PubMed]
- Sun L, Wang Q, Liu M, Xu G, Yin H, Wang D, et al. Albumin binding function is a novel biomarker for early liver damage and disease progression in non-alcoholic fatty liver disease. Endocrine 2020;69(2):294-302.
[Crossref] [Google scholar] [PubMed]
- Rametta R, Mozzi E, Dongiovanni P, Motta BM, Milano M, Roviaro G, et al. Increased insulin receptor substrate 2 expression is associated with steatohepatitis and altered lipid metabolism in obese subjects. Int J Obes 2013;37(7):986-92.
[Crossref] [Google scholar] [PubMed]
- Liang D, Chen H, Zhao L, Zhang W, Hu J, Liu Z, et al. Inhibition of EGFR attenuates fibrosis and stellate cell activation in diet-induced model of nonalcoholic fatty liver disease. Biochim Biophys Acta Mol Basis Dis 2018;1864(1):133-42.
[Crossref] [Google scholar] [PubMed]
- Thapaliya S, Wree A, Povero D, Inzaugarat ME, Berk M, Dixon L, et al. Caspase 3 inactivation protects against hepatic cell death and ameliorates fibrogenesis in a diet-induced NASH model. Dig Dis Sci 2014;59(6):1197-206.
[Crossref] [Google scholar] [PubMed]
- de Conti A, Tryndyak V, Willett RA, Borowa?Mazgaj B, Watson A, Patton R, et al. Characterization of the variability in the extent of nonalcoholic fatty liver induced by a high?fat diet in the genetically diverse collaborative cross mouse model. FASEB J 2020;34(6):7773-85.
[Crossref] [Google scholar] [PubMed]
- Loomba R, Sanyal AJ. The global NAFLD epidemic. Nat Rev Gastroenterol Hepatol 2013;10(11):686-90.
[Crossref] [Google scholar] [PubMed]
- Zhang L, Zhang Y, Jiang Y, Dou X, Li S, Chai H, et al. Upregulated SOCC and IP3R calcium channels and subsequent elevated cytoplasmic calcium signaling promote nonalcoholic fatty liver disease by inhibiting autophagy. Mol Cell Biochem 2021;476(8):3163-75.
[Crossref] [Google scholar] [PubMed]
- Matsuda S, Kobayashi M, Kitagishi Y. Roles for PI3K/AKT/PTEN pathway in cell signaling of nonalcoholic fatty liver disease. Int Sch Res Notices Endocrinol 2013;2013:1-7.
[Crossref] [Google scholar] [PubMed]
- Cai CX, Buddha H, Castelino-Prabhu S, Zhang Z, Britton RS, Bacon BR, et al. Activation of insulin-PI3K/AKT-p70S6K pathway in hepatic stellate cells contributes to fibrosis in nonalcoholic steatohepatitis. Dig Dis Sci 2017;62(4):968-78.
[Crossref] [Google scholar] [PubMed]
- Ma Z, Chu L, Liu H, Wang W, Li J, Yao W, et al. Beneficial effects of paeoniflorin on non-alcoholic fatty liver disease induced by high-fat diet in rats. Sci Repo 2017;7(1):1-10.
- Zhu XY, Xia HG, Wang ZH, Li B, Jiang HY, Li DL, et al. In vitro and in vivo approaches for identifying the role of aryl hydrocarbon receptor in the development of nonalcoholic fatty liver disease. Toxicol Lett 2020;319:85-94.
[Crossref] [Google scholar] [PubMed]
- Sukkasem N, Chatuphonprasert W, Jarukamjorn K. Cytochrome P450 expression-associated multiple-hit pathogenesis of non-alcoholic fatty liver disease (NAFLD) in HepG2 cells. Trop J Pharm Res 2020;19(4):707-14.
- Zheng W, Xue W, Jianli L, Zhishu S, Zhongxing T. Study on toxicity and heat characteristic of Aconiti Radix and Aconiti Lateralis Radix Praeparata by experimental method. Chin Arch Traditi Chin Med 2016;34(03):582-4.
- Jou J, Choi SS, Diehl AM. Mechanisms of disease progression in nonalcoholic fatty liver disease. Semin Liver Dis 2008;28(4):370-9.
[Crossref] [Google scholar] [PubMed]
- Wang M, Zhang J, Gong N. Role of the PI3K/AKT signaling pathway in liver ischemia reperfusion injury: A narrative review. Ann Palliat Med 2022;11(2):806-17.
[Crossref] [Google scholar] [PubMed]
- Yiwei T, Hua H, Hui G, Mao M, Xiang L. HOTAIR interacting with MAPK1 regulates ovarian cancer skov3 cell proliferation, migration, and invasion. Med Sci Monit 2015;21:1856-63.
[Crossref] [Google scholar] [PubMed]
- Xiao Y, Liu H, Yu J, Zhao Z, Xiao F, Xia T, et al. MAPK1/3 regulate hepatic lipid metabolism via ATG7-dependent autophagy. Autophagy 2016;12(3):592-3.
[Crossref] [Google scholar] [PubMed]
- Toraih EA, Alrefai HG, Hussein MH, Helal GM, Khashana MS, Fawzy MS. Overexpression of heat shock protein HSP90AA1 and translocase of the outer mitochondrial membrane TOM34 in HCV-induced hepatocellular carcinoma: A pilot study. Clin Biochem 2019;63:10-7.
[Crossref] [Google scholar] [PubMed]